

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0214.6
(398 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
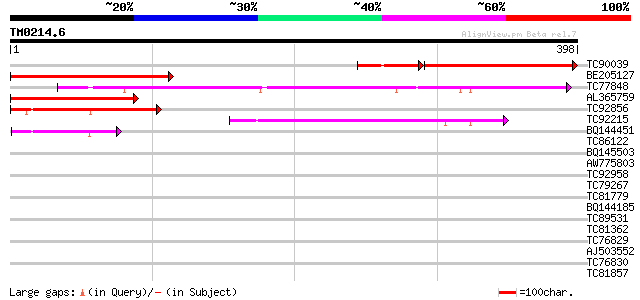
Sequences producing significant alignments: (bits) Value
TC90039 similar to GP|17381270|gb|AAL36053.1 AT3g24470/MXP5_4 {A... 205 5e-61
BE205127 similar to GP|16604677|gb| unknown protein {Arabidopsis... 195 2e-50
TC77848 similar to PIR|F86296|F86296 hypothetical protein AAF185... 193 1e-49
AL365759 similar to GP|17381270|gb| AT3g24470/MXP5_4 {Arabidopsi... 161 4e-40
TC92856 similar to GP|16604677|gb|AAL24131.1 unknown protein {Ar... 121 5e-28
TC92215 similar to GP|6862921|gb|AAF30310.1| hypothetical protei... 94 9e-20
BQ144451 weakly similar to GP|16604677|gb| unknown protein {Arab... 63 2e-10
TC86122 similar to SP|Q9SWE7|VATE_CITLI Vacuolar ATP synthase su... 33 0.15
BQ145503 33 0.25
AW775803 similar to GP|19911201|db glucosyltransferase-9 {Vigna ... 31 0.74
TC92958 31 0.96
TC79267 similar to PIR|T00782|T00782 probable anthranilate phosp... 30 2.1
TC81779 29 2.8
BQ144185 similar to GP|595768|gb|AA non-functional lacZ alpha pe... 28 4.8
TC89531 similar to GP|19034049|gb|AAL83681.1 putative extensin {... 25 6.6
TC81362 similar to GP|19911201|dbj|BAB86927. glucosyltransferase... 28 8.1
TC76829 similar to PIR|T14544|T14544 fructokinase (EC 2.7.1.4) -... 28 8.1
AJ503552 28 8.1
TC76830 homologue to SP|P37829|SCRK_SOLTU Fructokinase (EC 2.7.1... 28 8.1
TC81857 weakly similar to GP|13509841|emb|CAC35411. unnamed prot... 28 8.1
>TC90039 similar to GP|17381270|gb|AAL36053.1 AT3g24470/MXP5_4 {Arabidopsis
thaliana}, partial (24%)
Length = 809
Score = 205 bits (522), Expect(2) = 5e-61
Identities = 92/107 (85%), Positives = 99/107 (91%)
Frame = +3
Query: 292 ILAIVIATFSTGIDSKCFQFRKDDDIPAEDDVPYGYGFFHFVFATGAMYFAMLLIGWNSH 351
ILAIV+ TFSTGIDSKCFQ RK D EDDVPYGYGFFHFVFATGAMYFAMLLIGWN+H
Sbjct: 144 ILAIVMQTFSTGIDSKCFQLRKGDKPAEEDDVPYGYGFFHFVFATGAMYFAMLLIGWNTH 323
Query: 352 HSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPIIWKSRQTDST 398
HSMRKW++DVGWTS WVRIVNEWLAVCVYLWML+APIIWK+RQT+ST
Sbjct: 324 HSMRKWSLDVGWTSAWVRIVNEWLAVCVYLWMLIAPIIWKARQTEST 464
Score = 47.4 bits (111), Expect(2) = 5e-61
Identities = 25/46 (54%), Positives = 31/46 (67%)
Frame = +1
Query: 245 TPGLMGLYVVFLCWNAIRSEPDGGSCIRKSDSSTKTDWLSIISFII 290
+PGLMGLYV L SEP+G CIR S + TK DW +IISF++
Sbjct: 7 SPGLMGLYVSSLVVGN*-SEPEGDQCIRTSGTVTKPDWQNIISFVM 141
>BE205127 similar to GP|16604677|gb| unknown protein {Arabidopsis thaliana},
partial (24%)
Length = 559
Score = 195 bits (496), Expect = 2e-50
Identities = 92/115 (80%), Positives = 99/115 (86%)
Frame = +3
Query: 1 METGVSNNGNEACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAARDYGRGAL 60
METG SNNGNE C IS D+SW+ QFRNASNP MARY YA +FL+ANLLAWAARDYGR AL
Sbjct: 213 METGESNNGNERCIISNDSSWFSQFRNASNPWMARYAYAFIFLLANLLAWAARDYGRSAL 392
Query: 61 TEMERLKGCNGGKDCLGAEGVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSG 115
TEMERLKGCNGGKDCLGAE VLRVSLGCF FY IMFL+T TSKL + R+TWHSG
Sbjct: 393 TEMERLKGCNGGKDCLGAERVLRVSLGCFTFYIIMFLSTTGTSKLKQERNTWHSG 557
>TC77848 similar to PIR|F86296|F86296 hypothetical protein AAF18512.1
[imported] - Arabidopsis thaliana, complete
Length = 1592
Score = 193 bits (490), Expect = 1e-49
Identities = 120/386 (31%), Positives = 190/386 (49%), Gaps = 25/386 (6%)
Frame = +2
Query: 34 ARYVYALMFLVANLLAWAARDYGRGALTEMERLKGCNGGKDCLGAE-----GVLRVSLGC 88
AR Y +F ++ ++AW R+ A ME + N K E VLRVS G
Sbjct: 230 ARIAYCGLFALSLVVAWMLREV---AAPLMESIPWINHFKQTPSREWFETDAVLRVSFGN 400
Query: 89 FIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHF 148
F+F+ I+ + RD H G W +KI+ W + + F LP+E I Y ++ F
Sbjct: 401 FLFFTILAAMMVGVKTQKDPRDGLHHGGWMMKIICWCLLVIFMFFLPNEIISFYETISKF 580
Query: 149 GAGVFLLIQLISIISFITWLNDCCES--EKYAAKCQIHVMLFATTAYVVCLVGIILMFIW 206
G+G+FLL+Q++ ++ F+ ND E++ I + + + YV V ++F +
Sbjct: 581 GSGMFLLVQVVLLLDFVHRWNDTWVGYDEQF---WYIALFVVSLVCYVATFVFSGVLFHF 751
Query: 207 YAPK-PSCLLNIFFIAWTLVLLQLMTSVSLHPKVNAGILTPGLMGLYVVFLCWNAIRSEP 265
+ P C NIFFI+ TL+L + V+LHP VN +L ++ Y ++LC++A+ SEP
Sbjct: 752 FTPSGQDCGTNIFFISMTLMLAFVFAIVALHPAVNGSVLPASVISFYCMYLCYSALASEP 931
Query: 266 DGGSC--IRKSDSSTKTDWLSIISFIIAILAIVIATFSTGIDSKCFQFRKDD-------D 316
C + K + T L++ + +L++V + G +
Sbjct: 932 RDYECNGLHKHSKAVSTGSLTL-GLVTTVLSVVYSAVRAGSSATVLSPPSSPRAGKPLLP 1108
Query: 317 IPAEDD--------VPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWV 368
+ A+D+ V Y Y FFH +F+ +MY AMLL GW++ +DVGW S WV
Sbjct: 1109LDAKDEESNEKAKPVTYSYAFFHLIFSLASMYSAMLLTGWSTSVGESGKLVDVGWPSVWV 1288
Query: 369 RIVNEWLAVCVYLWMLVAPIIWKSRQ 394
RIV W +YLW LVAPI++ R+
Sbjct: 1289RIVTCWATALLYLWSLVAPIMFPERE 1366
>AL365759 similar to GP|17381270|gb| AT3g24470/MXP5_4 {Arabidopsis thaliana},
partial (17%)
Length = 497
Score = 161 bits (408), Expect = 4e-40
Identities = 75/90 (83%), Positives = 80/90 (88%)
Frame = -1
Query: 1 METGVSNNGNEACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAARDYGRGAL 60
METG SNNGNE C IS D+SW+ QFRNASNP MARY YA +FL+AN LAWAARDYGR AL
Sbjct: 284 METGESNNGNERCIISNDSSWFSQFRNASNPWMARYAYAFIFLLANSLAWAARDYGRSAL 105
Query: 61 TEMERLKGCNGGKDCLGAEGVLRVSLGCFI 90
TEMERLKGCNGGKDCLGAEGVLRVSLGCF+
Sbjct: 104 TEMERLKGCNGGKDCLGAEGVLRVSLGCFV 15
>TC92856 similar to GP|16604677|gb|AAL24131.1 unknown protein {Arabidopsis
thaliana}, partial (10%)
Length = 821
Score = 121 bits (303), Expect = 5e-28
Identities = 65/111 (58%), Positives = 78/111 (69%), Gaps = 5/111 (4%)
Frame = +1
Query: 1 METGVSNNGN---EACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAARDY-- 55
ME SNN N + C + KD+SW QF+NASNP MARYVY L+FL AN+LAWA RD
Sbjct: 490 MEPRESNNINSNQQGCGL-KDSSWCCQFKNASNPLMARYVYGLIFLAANMLAWATRDELS 666
Query: 56 GRGALTEMERLKGCNGGKDCLGAEGVLRVSLGCFIFYFIMFLTTARTSKLN 106
ALTE++ K C GKDCLGA GVLRVS+GCF+F+ +M +T RTS LN
Sbjct: 667 SISALTELKGFKACKVGKDCLGAHGVLRVSMGCFLFFMMMXGSTTRTSNLN 819
>TC92215 similar to GP|6862921|gb|AAF30310.1| hypothetical protein
{Arabidopsis thaliana}, partial (63%)
Length = 666
Score = 94.0 bits (232), Expect = 9e-20
Identities = 59/216 (27%), Positives = 103/216 (47%), Gaps = 20/216 (9%)
Frame = +1
Query: 155 LIQLISIISFITWLNDCCESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYAPKP-SC 213
LIQ+I ++ ND EK K I +++ + Y+ ++FIW+ P C
Sbjct: 1 LIQVIILLDCTHNWNDSWV-EKDEQKWYIALLVVSIGCYIAAFTLSGILFIWFNPGGYDC 177
Query: 214 LLNIFFIAWTLVLLQLMTSVSLHPKVNAGILTPGLMGLYVVFLCWNAIRSEPDGGSCIRK 273
LN+FF++ +++L + V+LHPKVN +L ++ LY ++C+ + SEP G C
Sbjct: 178 GLNVFFLSMSMILAFVFGVVALHPKVNGSLLPASVISLYCAYVCYTGLSSEPRGYECNGL 357
Query: 274 SDSSTKTDWLSIISFIIAILAIVIATFSTGI--------------DSKCFQFRKDDDIPA 319
+ S + ++ + +L+++ + G +SK ++
Sbjct: 358 NKSRAVSTGTLVLGMLTTVLSVLYSALRAGSSTTFLSPPSSPKAGESKPLLEEVEEGKSK 537
Query: 320 EDD-----VPYGYGFFHFVFATGAMYFAMLLIGWNS 350
+++ V Y Y FFH +FA +MY AMLL GW S
Sbjct: 538 KEEKEARPVSYSYSFFHLIFALASMYSAMLLSGWTS 645
>BQ144451 weakly similar to GP|16604677|gb| unknown protein {Arabidopsis
thaliana}, partial (9%)
Length = 758
Score = 62.8 bits (151), Expect = 2e-10
Identities = 35/79 (44%), Positives = 45/79 (56%), Gaps = 2/79 (2%)
Frame = +1
Query: 2 ETGVSNNGNEACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAARD--YGRGA 59
E+ N+ + C + KD+SW QF+NASNP MA YVY L+FL AN LAWA R+ A
Sbjct: 298 ESNSINSNQQGCGL-KDSSWCCQFKNASNPLMATYVYGLIFLTANTLAWATREELSSLVA 474
Query: 60 LTEMERLKGCNGGKDCLGA 78
L E+ K C G+
Sbjct: 475 LAELYCFKACGNWNTLSGS 531
>TC86122 similar to SP|Q9SWE7|VATE_CITLI Vacuolar ATP synthase subunit E (EC
3.6.3.14) (V-ATPase E subunit) (Vacuolar proton pump E
subunit), complete
Length = 1249
Score = 33.5 bits (75), Expect = 0.15
Identities = 19/71 (26%), Positives = 38/71 (52%)
Frame = +3
Query: 134 LPSEFIQIYGEVAHFGAGVFLLIQLISIISFITWLNDCCESEKYAAKCQIHVMLFATTAY 193
LP Q++G+VA ++ I S I FI + DCCE + A ++++L + +++
Sbjct: 792 LPHIRKQLFGQVAV*RPWLYCSI---SPIIFIIYCRDCCERKICRAIAFVNIVLLSLSSF 962
Query: 194 VVCLVGIILMF 204
+ C+ + +F
Sbjct: 963 IFCVAHLTFVF 995
>BQ145503
Length = 389
Score = 32.7 bits (73), Expect = 0.25
Identities = 13/46 (28%), Positives = 27/46 (58%)
Frame = +3
Query: 159 ISIISFITWLNDCCESEKYAAKCQIHVMLFATTAYVVCLVGIILMF 204
IS I FI + DCCE + A ++++L + ++++ C+ + +F
Sbjct: 60 ISPIIFIIYCRDCCERKICRAIAFVNIVLLSLSSFIFCVAHLTFVF 197
>AW775803 similar to GP|19911201|db glucosyltransferase-9 {Vigna angularis},
partial (34%)
Length = 716
Score = 31.2 bits (69), Expect = 0.74
Identities = 11/26 (42%), Positives = 18/26 (68%)
Frame = +2
Query: 203 MFIWYAPKPSCLLNIFFIAWTLVLLQ 228
+F PKPSC+++ FFI WT+ + +
Sbjct: 383 LFDALTPKPSCIISDFFIPWTIQIAE 460
>TC92958
Length = 652
Score = 30.8 bits (68), Expect = 0.96
Identities = 12/31 (38%), Positives = 16/31 (50%)
Frame = +2
Query: 322 DVPYGYGFFHFVFATGAMYFAMLLIGWNSHH 352
D+ Y +G FH F G+ + L I W S H
Sbjct: 302 DLLYPFGSFHLCFLIGSFHLCFLTISWGS*H 394
>TC79267 similar to PIR|T00782|T00782 probable anthranilate
phosphoribosyltransferase (EC 2.4.2.18) T22J18.21 -
Arabidopsis thaliana, partial (60%)
Length = 1612
Score = 29.6 bits (65), Expect = 2.1
Identities = 13/54 (24%), Positives = 26/54 (48%), Gaps = 4/54 (7%)
Frame = +3
Query: 157 QLISIISFIT----WLNDCCESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIW 206
+++S++S T WLND C C +HV+ Y ++ I ++++
Sbjct: 825 RIMSLLSGFTAVCKWLNDICTWRNPITTCLVHVLFLILVCYPELILPTIFLYLF 986
>TC81779
Length = 478
Score = 29.3 bits (64), Expect = 2.8
Identities = 20/77 (25%), Positives = 34/77 (43%)
Frame = +3
Query: 132 FLLPSEFIQIYGEVAHFGAGVFLLIQLISIISFITWLNDCCESEKYAAKCQIHVMLFATT 191
FLL I+ + E+A FGA F + LI +I N + + A Q+ T
Sbjct: 15 FLLE*HNIRYFVEMATFGASAFQFLMLIMVIGLFCVTNIVAQDSEIAPTSQLE----TGT 182
Query: 192 AYVVCLVGIILMFIWYA 208
+ + + G+I+ +A
Sbjct: 183 GFALPISGMIMFSSLFA 233
>BQ144185 similar to GP|595768|gb|AA non-functional lacZ alpha peptide
{unidentified cloning vector}, partial (9%)
Length = 395
Score = 28.5 bits (62), Expect = 4.8
Identities = 12/24 (50%), Positives = 16/24 (66%)
Frame = +1
Query: 131 PFLLPSEFIQIYGEVAHFGAGVFL 154
P LPS F +YGE+A F +G +L
Sbjct: 73 PIPLPSRFPNLYGEMAKFVSGYYL 144
>TC89531 similar to GP|19034049|gb|AAL83681.1 putative extensin {Oryza sativa
(japonica cultivar-group)}, partial (8%)
Length = 1184
Score = 25.0 bits (53), Expect(2) = 6.6
Identities = 12/60 (20%), Positives = 27/60 (45%), Gaps = 2/60 (3%)
Frame = -1
Query: 110 DTWHSGWWSVKIVLWVGMTVIPFLLPSE--FIQIYGEVAHFGAGVFLLIQLISIISFITW 167
++W + WW + + V T IP L+ + + ++ + + I+++ ITW
Sbjct: 1067 NSWRNSWWHMHVSTIVISTTIPVLITPVPIVMMVVRTISTLMRTTITMSE*ITLLWMITW 888
Score = 21.2 bits (43), Expect(2) = 6.6
Identities = 5/12 (41%), Positives = 9/12 (74%)
Frame = -2
Query: 161 IISFITWLNDCC 172
+ + +TW +DCC
Sbjct: 772 VAAIVTWGSDCC 737
>TC81362 similar to GP|19911201|dbj|BAB86927. glucosyltransferase-9 {Vigna
angularis}, partial (70%)
Length = 1347
Score = 27.7 bits (60), Expect = 8.1
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = +1
Query: 203 MFIWYAPKPSCLLNIFFIAWT 223
+F +PKPSC+++ F I WT
Sbjct: 421 LFDKLSPKPSCIISDFCITWT 483
>TC76829 similar to PIR|T14544|T14544 fructokinase (EC 2.7.1.4) - beet,
partial (95%)
Length = 1408
Score = 27.7 bits (60), Expect = 8.1
Identities = 18/47 (38%), Positives = 23/47 (48%)
Frame = +1
Query: 189 ATTAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSL 235
A T+ V CL G +W PK L + + W L+L M SVSL
Sbjct: 274 AMTSSVTCLPGS*RKTVWL-PKV*PLTKVHVLLWRLLLYAPMESVSL 411
>AJ503552
Length = 330
Score = 27.7 bits (60), Expect = 8.1
Identities = 12/40 (30%), Positives = 21/40 (52%)
Frame = -2
Query: 89 FIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMT 128
F+FYF + K+ E + + W VK+++W G+T
Sbjct: 167 FVFYFN*YGPLIVPEKIREKENKKGNPWSYVKMIVWFGLT 48
>TC76830 homologue to SP|P37829|SCRK_SOLTU Fructokinase (EC 2.7.1.4).
[Potato] {Solanum tuberosum}, partial (38%)
Length = 436
Score = 27.7 bits (60), Expect = 8.1
Identities = 18/47 (38%), Positives = 23/47 (48%)
Frame = +1
Query: 189 ATTAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSL 235
A T+ V CL G +W PK L + + W L+L M SVSL
Sbjct: 160 AMTSSVTCLPGS*RKTVWL-PKV*PLTKVHVLLWRLLLYAPMESVSL 297
>TC81857 weakly similar to GP|13509841|emb|CAC35411. unnamed protein product
{Zea mays}, partial (17%)
Length = 1055
Score = 27.7 bits (60), Expect = 8.1
Identities = 15/44 (34%), Positives = 19/44 (43%)
Frame = -3
Query: 175 EKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYAPKPSCLLNIF 218
EK C + +L + VVC +I M WY P S IF
Sbjct: 972 EKTNQNCMM-TLLVTSDVTVVCCAALIEMVKWYRPSTSSFFPIF 844
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.328 0.140 0.472
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,363,293
Number of Sequences: 36976
Number of extensions: 276098
Number of successful extensions: 2113
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 2079
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2107
length of query: 398
length of database: 9,014,727
effective HSP length: 98
effective length of query: 300
effective length of database: 5,391,079
effective search space: 1617323700
effective search space used: 1617323700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0214.6


