

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0214.10
(74 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
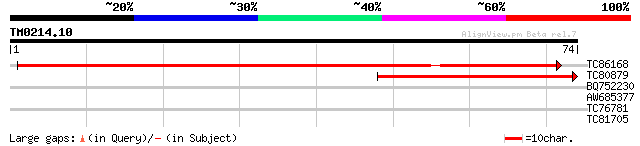
TC86168 homologue to GP|13936812|gb|AAK49947.1 TGF-beta receptor... 72 4e-14
TC80879 40 2e-04
BQ752230 26 3.3
AW685377 26 3.3
TC76781 similar to GP|12539609|gb|AAG59584.1 pathogen-inducible ... 25 5.6
TC81705 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428... 25 7.3
>TC86168 homologue to GP|13936812|gb|AAK49947.1 TGF-beta
receptor-interacting protein 1 {Phaseolus vulgaris},
complete
Length = 1322
Score = 72.0 bits (175), Expect = 4e-14
Identities = 39/71 (54%), Positives = 48/71 (66%)
Frame = +2
Query: 2 VLRGGQVASAVSTTDHHAGNFEAKFYDMILQEEFRDVKGHSGLINVMAFIMEERPALSDP 61
VL GGQ ASAV+TTDH AG FEAKF+D ILQEE VKGH G IN +AF + + + S
Sbjct: 911 VLGGGQDASAVTTTDHRAGKFEAKFFDKILQEEIGGVKGHFGPINALAFNPDGK-SFSSG 1087
Query: 62 VSDGHVLVQFY 72
DG+V + +
Sbjct: 1088GEDGYVRLHHF 1120
>TC80879
Length = 942
Score = 40.0 bits (92), Expect = 2e-04
Identities = 20/26 (76%), Positives = 22/26 (83%)
Frame = +3
Query: 49 AFIMEERPALSDPVSDGHVLVQFYGN 74
A IMEE ALSD V+DG+VLVQFYGN
Sbjct: 657 AEIMEESCALSDRVNDGNVLVQFYGN 734
>BQ752230
Length = 674
Score = 25.8 bits (55), Expect = 3.3
Identities = 13/28 (46%), Positives = 16/28 (56%)
Frame = +2
Query: 39 KGHSGLINVMAFIMEERPALSDPVSDGH 66
KGHSGLI + R L+ PV+ GH
Sbjct: 83 KGHSGLIVSSDTDTQRRACLNGPVALGH 166
>AW685377
Length = 430
Score = 25.8 bits (55), Expect = 3.3
Identities = 10/33 (30%), Positives = 16/33 (48%)
Frame = -2
Query: 35 FRDVKGHSGLINVMAFIMEERPALSDPVSDGHV 67
F+D+ GH IN+++ S SD H+
Sbjct: 159 FKDINGHCFGINILSLFFRSMAIRSQTXSDHHI 61
>TC76781 similar to GP|12539609|gb|AAG59584.1 pathogen-inducible
alpha-dioxygenase {Nicotiana attenuata}, partial (94%)
Length = 2232
Score = 25.0 bits (53), Expect = 5.6
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = +2
Query: 15 TDHHAGNFEAKFYDMILQEEFRDVKGHSGLINVMAFIMEER 55
TD G A +Y + L ++F+D GHSG + F+ +R
Sbjct: 1043 TDTLLGGMRANWYGL-LGKKFKDTFGHSGNSILSGFVGMKR 1162
>TC81705 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428(E30359)
corresponds to a region of the predicted gene.~Similar
to antifreeze-like, partial (33%)
Length = 1443
Score = 24.6 bits (52), Expect = 7.3
Identities = 10/31 (32%), Positives = 17/31 (54%)
Frame = +1
Query: 17 HHAGNFEAKFYDMILQEEFRDVKGHSGLINV 47
HH + + F M+ Q E + GH G+++V
Sbjct: 205 HHYNHAFSGFSAMLTQSEASALSGHDGVVSV 297
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.137 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,690,564
Number of Sequences: 36976
Number of extensions: 12249
Number of successful extensions: 41
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 41
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 41
length of query: 74
length of database: 9,014,727
effective HSP length: 50
effective length of query: 24
effective length of database: 7,165,927
effective search space: 171982248
effective search space used: 171982248
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0214.10


