

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.7
(451 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
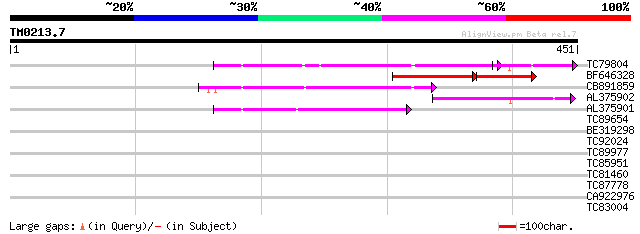
Score E
Sequences producing significant alignments: (bits) Value
TC79804 weakly similar to GP|6175165|gb|AAF04891.1| Mutator-like... 103 8e-29
BF646328 similar to PIR|F84533|F845 Mutator-like transposase [im... 59 5e-20
CB891859 similar to GP|6175165|gb| Mutator-like transposase {Ara... 74 2e-13
AL375902 similar to GP|9759134|dbj mutator-like transposase-like... 67 1e-11
AL375901 similar to GP|9759134|dbj mutator-like transposase-like... 42 6e-04
TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein... 31 1.1
BE319298 similar to GP|17385689|dbj putative Myb-related transcr... 31 1.1
TC92024 similar to PIR|T05645|T05645 hypothetical protein F20D10... 30 2.5
TC89977 similar to GP|5764395|gb|AAD51282.1| far-red impaired re... 29 3.3
TC85951 similar to GP|18000068|gb|AAL54885.1 cytochrome P450-dep... 29 4.3
TC81460 similar to PIR|B84478|B84478 probable replication protei... 29 4.3
TC87778 similar to GP|5103827|gb|AAD39657.1| ESTs gb|F20110 and ... 28 5.6
CA922976 weakly similar to GP|9502168|gb|A contains simlarity to... 28 7.3
TC83004 similar to PIR|T10985|T10985 regulator protein ROM2 - ki... 28 9.5
>TC79804 weakly similar to GP|6175165|gb|AAF04891.1| Mutator-like transposase
{Arabidopsis thaliana}, partial (57%)
Length = 2129
Score = 103 bits (256), Expect(2) = 8e-29
Identities = 68/231 (29%), Positives = 113/231 (48%), Gaps = 1/231 (0%)
Frame = +1
Query: 163 HIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIP 222
H FC+RHL N K+F +I +L+ +AA AT + EKM E++ + P A WL
Sbjct: 1021 HAFCMRHLSENIGKEFKNSRLI-HLLWSAAYATTINAFREKMAEIEEVSPNASMWLQHFH 1197
Query: 223 TRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKL 282
W F + L +N+ E FN IL ++ P++ +++ I+S + F K
Sbjct: 1198 PSQWALVYFEG-TRYGHLSSNIEE-FNKWILEAQELPIIQVIERIQSKLKTEFDDRRLKS 1371
Query: 283 EKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQF-VVNLAEHTCSCNF 341
+ + P +R+ I + + E ++EV +S D+ +VN+ H+CSC
Sbjct: 1372 SSWCSVLTPSSERRMVEAINRASTYQVLKSDEVEFEV---ISADRSDIVNIGSHSCSCRD 1542
Query: 342 WELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPINGQKL 392
W+L GIPC HAVAA+ S + + + +Y T +TP++ + L
Sbjct: 1543 WQLYGIPCSHAVAALISSRKDVYAYTAKCFTVASYRDTVCRGVTPLSLENL 1695
Score = 42.0 bits (97), Expect(2) = 8e-29
Identities = 25/73 (34%), Positives = 38/73 (51%), Gaps = 6/73 (8%)
Frame = +3
Query: 385 TPINGQKLWPTT-----NDPYIL-PPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSRCGQ 438
TP+ G+ W T ND ++ PP ++R PGRP+K +R + D T SRC Q
Sbjct: 1674 TPVPGKLEWRTDESALDNDIAVVRPPKFRRPPGRPEK-KRICVEDHNRDKHTVHCSRCNQ 1850
Query: 439 FGHNSRSCRNPVV 451
GH +C+ ++
Sbjct: 1851 TGHYKTTCKAEMI 1889
>BF646328 similar to PIR|F84533|F845 Mutator-like transposase [imported] -
Arabidopsis thaliana, partial (3%)
Length = 636
Score = 59.3 bits (142), Expect(2) = 5e-20
Identities = 27/69 (39%), Positives = 42/69 (60%), Gaps = 1/69 (1%)
Frame = -1
Query: 305 RYWVATRCGEDKYEVKHMVSQ-DQFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRP 363
R W AT G+ + ++ ++ ++++VNLA+ TC+C W L GIPC H + I H+
Sbjct: 483 RGWKATWHGDMEMNNFNVSNETNKYIVNLAQRTCACRKWNLTGIPCAHVIPCIWHNGLAD 304
Query: 364 EDFVHPYYR 372
E+FV YYR
Sbjct: 303 ENFVSSYYR 277
Score = 56.2 bits (134), Expect(2) = 5e-20
Identities = 23/48 (47%), Positives = 33/48 (67%)
Frame = -3
Query: 372 RREAYMATYGHVITPINGQKLWPTTNDPYILPPLYKRAPGRPKKLRRR 419
R+ +ATY H++ P +G KLWP TN +I P +R+ GRPKKLR++
Sbjct: 193 RKSQCLATYSHIVLPSSGPKLWPVTNTEHINPLAKRRSAGRPKKLRKK 50
>CB891859 similar to GP|6175165|gb| Mutator-like transposase {Arabidopsis
thaliana}, partial (27%)
Length = 625
Score = 73.6 bits (179), Expect = 2e-13
Identities = 51/200 (25%), Positives = 96/200 (47%), Gaps = 11/200 (5%)
Frame = +1
Query: 151 LTVFDD----LMGGVE-------HIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQL 199
LT+ D ++ GVE H FC+RHL +F+K+F +++ NL+ AA
Sbjct: 34 LTILSDRQQGIVDGVEANFPTAFHGFCMRHLSDSFRKEFNNTMLV-NLLWEAANCLTIIE 210
Query: 200 WTEKMMELQALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKP 259
+ K+ME++ + +A W+ ++P R W F + + L N+ E+ N+ IL P
Sbjct: 211 FEGKVMEIEEISQDAAYWIRRVPPRLWATAYFEGH-RFGHLTANIVEALNSWILEASGLP 387
Query: 260 VLTMVDWIRSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEV 319
++ M++ IR +M F E ++ ++P + + +E R + R E ++EV
Sbjct: 388 IIQMMECIRRQLMTWFNERRETSMQWTSILVPSAERSVAEALERARTYQVLRANEAEFEV 567
Query: 320 KHMVSQDQFVVNLAEHTCSC 339
+ + +V++ C C
Sbjct: 568 --ISHEGTNIVDIRNRCCLC 621
>AL375902 similar to GP|9759134|dbj mutator-like transposase-like protein
{Arabidopsis thaliana}, partial (20%)
Length = 542
Score = 67.0 bits (162), Expect = 1e-11
Identities = 38/125 (30%), Positives = 51/125 (40%), Gaps = 11/125 (8%)
Frame = +2
Query: 337 CSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPINGQKLWPTT 396
CSC W+L G PC HA AA+ F P + ++Y Y +I PI + W
Sbjct: 14 CSCRRWQLYGYPCAHAAAALISCGHNAHMFAEPCFTVQSYRMAYSQMIYPIPDKSQWREH 193
Query: 397 N-----------DPYILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSRCGQFGHNSRS 445
D I PP +R PGRPKK R + + RC GH+ +
Sbjct: 194 GEGAEGGGGARVDIVIHPPKIRRPPGRPKKKVLRVENFKRPKRVVQC-GRCHMLGHSQKK 370
Query: 446 CRNPV 450
C P+
Sbjct: 371 CTMPI 385
>AL375901 similar to GP|9759134|dbj mutator-like transposase-like protein
{Arabidopsis thaliana}, partial (27%)
Length = 493
Score = 41.6 bits (96), Expect = 6e-04
Identities = 33/157 (21%), Positives = 64/157 (40%)
Frame = +3
Query: 163 HIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIP 222
H FCLR++ NF+ F ++ N+ A A + K+ E+ + + +W P
Sbjct: 24 HGFCLRYVSENFRDTFKNTKLV-NIFWNAVYALTAAEFESKITEMIEVSQDVISWFQHFP 200
Query: 223 TRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKL 282
W A+ + ++E L + PV+ M+++IR + F E
Sbjct: 201 PFLWAV-AYFDGVRYGHFTLGVTELLYNWALECHELPVVQMMEYIRQQMTSWFNDRREVG 377
Query: 283 EKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEV 319
++ ++P KR+ I + R E ++E+
Sbjct: 378 MEWTSILVPSAEKRISEAIADAHCYQVLRANEVEFEI 488
>TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein F20D10.290 -
Arabidopsis thaliana, partial (65%)
Length = 1358
Score = 30.8 bits (68), Expect = 1.1
Identities = 21/65 (32%), Positives = 33/65 (50%)
Frame = +1
Query: 290 MPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEHTCSCNFWELVGIPC 349
M P ++E + Y VA + GE+ + H V+ F + + +CSC +E GI C
Sbjct: 1030 MANPATKIEDSGTITTYRVA-KFGEN--QTSHTVA---FNSSEMKASCSCQMFEYSGIVC 1191
Query: 350 RHAVA 354
RH +A
Sbjct: 1192 RHILA 1206
>BE319298 similar to GP|17385689|dbj putative Myb-related transcription
factor {Oryza sativa (japonica cultivar-group)}, partial
(20%)
Length = 425
Score = 30.8 bits (68), Expect = 1.1
Identities = 11/15 (73%), Positives = 12/15 (79%)
Frame = +2
Query: 434 SRCGQFGHNSRSCRN 448
S CG FGHNSR+C N
Sbjct: 44 SYCGNFGHNSRTCNN 88
>TC92024 similar to PIR|T05645|T05645 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (11%)
Length = 938
Score = 29.6 bits (65), Expect = 2.5
Identities = 16/42 (38%), Positives = 21/42 (49%)
Frame = +3
Query: 313 GEDKYEVKHMVSQDQFVVNLAEHTCSCNFWELVGIPCRHAVA 354
GED H +F V + TCSC +E G+ CRH +A
Sbjct: 36 GED-----HKAYNVRFNVLEMKATCSCQMFEFSGLLCRHILA 146
>TC89977 similar to GP|5764395|gb|AAD51282.1| far-red impaired response
protein {Arabidopsis thaliana}, partial (35%)
Length = 1523
Score = 29.3 bits (64), Expect = 3.3
Identities = 16/66 (24%), Positives = 32/66 (48%), Gaps = 6/66 (9%)
Frame = +2
Query: 316 KYEVKHMVSQDQFVVNL----AEHTCSCNFWELVGIPCRHAVAAISH--SSQRPEDFVHP 369
+Y V+ ++F+V +E +C C +E G CRHA++ + S P ++
Sbjct: 308 RYIVQDYEKDEEFLVTWKELSSEVSCFCRLFEYKGFLCRHALSVLQRCGCSSVPSHYIMK 487
Query: 370 YYRREA 375
+ ++A
Sbjct: 488 RWTKDA 505
>TC85951 similar to GP|18000068|gb|AAL54885.1 cytochrome P450-dependent
fatty acid hydroxylase {Vicia sativa}, partial (45%)
Length = 742
Score = 28.9 bits (63), Expect = 4.3
Identities = 12/28 (42%), Positives = 16/28 (56%)
Frame = +2
Query: 394 PTTNDPYILPPLYKRAPGRPKKLRRRDP 421
PT++ P +LPP AP P K R+ P
Sbjct: 113 PTSSKPSLLPPSSSTAPSVPAKFSRQTP 196
>TC81460 similar to PIR|B84478|B84478 probable replication protein A1
[imported] - Arabidopsis thaliana, partial (8%)
Length = 762
Score = 28.9 bits (63), Expect = 4.3
Identities = 12/20 (60%), Positives = 12/20 (60%)
Frame = +2
Query: 388 NGQKLWPTTNDPYILPPLYK 407
NGQ PT PY PPLYK
Sbjct: 602 NGQNFRPTVQPPYQPPPLYK 661
>TC87778 similar to GP|5103827|gb|AAD39657.1| ESTs gb|F20110 and gb|F20109
come from this gene. {Arabidopsis thaliana}, partial
(24%)
Length = 1350
Score = 28.5 bits (62), Expect = 5.6
Identities = 16/43 (37%), Positives = 22/43 (50%)
Frame = -2
Query: 35 LHVFGGLQEGVC*RVQAIYRVGWLPFEDTVWGNLTVCCGQGPQ 77
LH F Q+G +VQA+ + PF D W + CG+ PQ
Sbjct: 743 LHSFSLCQDG---QVQALSHLYPTPFADKWWPG*NLLCGREPQ 624
>CA922976 weakly similar to GP|9502168|gb|A contains simlarity to Arabidopsis
thaliana far-red impaired response protein
(GB:AAD51282.1), partial (11%)
Length = 827
Score = 28.1 bits (61), Expect = 7.3
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 2/40 (5%)
Frame = -3
Query: 337 CSCNFWELVGIPCRHA--VAAISHSSQRPEDFVHPYYRRE 374
CSC +E GI CRHA + + Q PE + + RE
Sbjct: 702 CSCKEFESSGILCRHALRILVTKNYFQLPEKYYLSRWHRE 583
>TC83004 similar to PIR|T10985|T10985 regulator protein ROM2 - kidney bean,
partial (78%)
Length = 1250
Score = 27.7 bits (60), Expect = 9.5
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +3
Query: 344 LVGIPCRHAVAAISHSS 360
L+G+PCRH + SHSS
Sbjct: 42 LIGLPCRHIMGKESHSS 92
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.341 0.149 0.525
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,902,973
Number of Sequences: 36976
Number of extensions: 236472
Number of successful extensions: 1718
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1698
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1713
length of query: 451
length of database: 9,014,727
effective HSP length: 99
effective length of query: 352
effective length of database: 5,354,103
effective search space: 1884644256
effective search space used: 1884644256
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0213.7


