

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
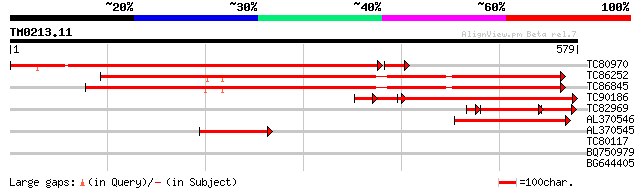
Query= TM0213.11
(579 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC80970 similar to SP|O24357|G6PC_SPIOL Glucose-6-phosphate 1-de... 630 0.0
TC86252 SP|Q42919|G6PD_MEDSA Glucose-6-phosphate 1-dehydrogenase... 456 e-128
TC86845 similar to PIR|T52611|T52611 glucose-6-phosphate 1-dehyd... 450 e-127
TC90186 homologue to PIR|T03740|T03740 glucose-6-phosphate 1-deh... 344 e-107
TC82969 similar to SP|Q43793|G6PC_TOBAC Glucose-6-phosphate 1-de... 112 2e-40
AL370546 similar to GP|15810387|gb putative glucose-6-phosphate ... 158 5e-39
AL370545 weakly similar to GP|15810387|gb| putative glucose-6-ph... 72 8e-13
TC80117 weakly similar to PIR|T48360|T48360 farnesylated protein... 29 4.4
BQ750979 similar to GP|13883467|gb PE_PGRS family protein {Mycob... 28 7.4
BG644405 weakly similar to PIR|A84564|A845 hypothetical protein ... 28 9.7
>TC80970 similar to SP|O24357|G6PC_SPIOL Glucose-6-phosphate 1-dehydrogenase
chloroplast precursor (EC 1.1.1.49) (G6PD). [Spinach],
partial (55%)
Length = 1289
Score = 630 bits (1624), Expect(2) = 0.0
Identities = 324/385 (84%), Positives = 341/385 (88%), Gaps = 5/385 (1%)
Frame = +1
Query: 1 MACVLSSSSTIVASYALRNEPQLYPVW----FSCI-RPGNLPRNHFQLKSSNGHPLNAVS 55
MA VL S S IV SY +NEPQL + SCI + GN R HFQLKSSNGHPLNAVS
Sbjct: 10 MASVLGSYSNIVTSYGFKNEPQLCTKFTTLCLSCITQHGNKGRKHFQLKSSNGHPLNAVS 189
Query: 56 SHSHDGLAGSSLTKEDSKPQPVDGPFLSSDSECTGSNLSITVVGASGDLAKKKIFPALFA 115
SH DGLA SSL KED +PQ V+G F SDSE TGSNLSITVVGASGDLAKKKIFPALFA
Sbjct: 190 SH--DGLAESSLAKEDGQPQQVEGLFSLSDSESTGSNLSITVVGASGDLAKKKIFPALFA 363
Query: 116 LFYEDCLPENFLVFGFARTKMTSEELRNMISRTLTCRIDKRANCEDKMDQFLKRCFYHSG 175
LFYED LPENF+VFG+ARTKMT EELRNMIS+TLTCRID+RANC DKMD FLKRCFYHSG
Sbjct: 364 LFYEDWLPENFIVFGYARTKMTDEELRNMISQTLTCRIDQRANCADKMDHFLKRCFYHSG 543
Query: 176 LYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPNIFVNVVRCASLKASSKKGWTRVIVE 235
LYNSE F DLD K+KEKEGG+ SNRLFYLSIPPNIFV+VVRCASLKASSK GWTRVIVE
Sbjct: 544 LYNSEEDFLDLDSKLKEKEGGRLSNRLFYLSIPPNIFVDVVRCASLKASSKNGWTRVIVE 723
Query: 236 KPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGKELVENLSVLRFSNLVFEPLWSRAYI 295
KPFGRDSESS+ELTR LKQYLTEDQIFRIDHYLGKELVENLSVLRFSNLVFEPLWSR YI
Sbjct: 724 KPFGRDSESSSELTRSLKQYLTEDQIFRIDHYLGKELVENLSVLRFSNLVFEPLWSRNYI 903
Query: 296 RNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAMEPPVSLDAEDIRNEKVK 355
RNVQ IFSEDFGTEGRGGYFDNYGIIRDIMQNHL+QILALFAMEPPVSLDAEDIRNEKVK
Sbjct: 904 RNVQLIFSEDFGTEGRGGYFDNYGIIRDIMQNHLVQILALFAMEPPVSLDAEDIRNEKVK 1083
Query: 356 VLRSMRPLQLENVVTGQYKGHSKGG 380
VLRSMRP+QLE+VV GQYKGHS+GG
Sbjct: 1084VLRSMRPIQLEDVVVGQYKGHSQGG 1158
Score = 48.9 bits (115), Expect(2) = 0.0
Identities = 22/26 (84%), Positives = 23/26 (87%)
Frame = +3
Query: 383 YPGYADDPSVPKGSLTPTFAAAALFI 408
YP Y DD +VPKGSLTPTFAAAALFI
Sbjct: 1167 YPAYIDDSTVPKGSLTPTFAAAALFI 1244
>TC86252 SP|Q42919|G6PD_MEDSA Glucose-6-phosphate 1-dehydrogenase
cytoplasmic isoform (EC 1.1.1.49) (G6PD). [Alfalfa],
complete
Length = 2081
Score = 456 bits (1172), Expect = e-128
Identities = 246/485 (50%), Positives = 329/485 (67%), Gaps = 10/485 (2%)
Frame = +1
Query: 93 LSITVVGASGDLAKKKIFPALFALFYEDCLPENFL-VFGFARTKMTSEELRNMISRTLTC 151
LSI V+GASGDLAKKK FPALF L+ ++ LP + + +FG+AR+K++ +ELRN + L
Sbjct: 253 LSIVVLGASGDLAKKKTFPALFHLYKQELLPPDEVHIFGYARSKISDDELRNKLRSYLVP 432
Query: 152 RIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSN-----RLFYLS 206
D + +FL+ Y SG Y+SE F LD ++ E E K S RLFYL+
Sbjct: 433 EKGASPKQLDDVSKFLQLVKYVSGPYDSEDGFRLLDKEISEHEYLKNSKEGSSRRLFYLA 612
Query: 207 IPPNIFVNV---VRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFR 263
+PP+++ +V ++ + S GWTRV+VEKPFGRD ES+ EL+ + + E QI+R
Sbjct: 613 LPPSVYPSVCKMIKTCCMNKSDLGGWTRVVVEKPFGRDLESAEELSTQIGELFEEPQIYR 792
Query: 264 IDHYLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRD 323
IDHYLGKELV+N+ VLRF+N F PLW+ +I NVQ +F EDFGT+GRGGYFD YGIIRD
Sbjct: 793 IDHYLGKELVQNMLVLRFANRFFLPLWNHNHIDNVQIVFREDFGTDGRGGYFDQYGIIRD 972
Query: 324 IMQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSY 383
I+QNHLLQ+L L AME PVSL E IR+EKVKVL S+ P++ + VV GQY+
Sbjct: 973 IIQNHLLQVLCLIAMEKPVSLKPEHIRDEKVKVLESVLPIRDDEVVLGQYE--------- 1125
Query: 384 PGYADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGN 443
GY DDP+VP S TPTFA L I N RW+GVPF++KAGKAL++++AEIRVQF+ VPG+
Sbjct: 1126-GYRDDPTVPDDSNTPTFATTILRIHNERWEGVPFIVKAGKALNSRKAEIRVQFKDVPGD 1302
Query: 444 LYKRNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYP-TEI 502
+++ + NE V+R+QP EAIY+K+ K PGL M +S+L+L Y RY I
Sbjct: 1303IFR-----SKKQGRNEFVIRLQPSEAIYMKLTVKQPGLEMSAVQSELDLSYGQRYQGITI 1467
Query: 503 PDAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAH 562
P+AYERL+LD I G+++ F+R DEL A+W +FTPLL +++ ++ P Y GSRGP A
Sbjct: 1468PEAYERLILDTIRGDQQHFVRRDELKASWQIFTPLLHKIDRGELKPVPYKPGSRGPAEAD 1647
Query: 563 YLAAK 567
L K
Sbjct: 1648ELLEK 1662
>TC86845 similar to PIR|T52611|T52611 glucose-6-phosphate 1-dehydrogenase
(EC 1.1.1.49) [imported] - Arabidopsis thaliana, partial
(94%)
Length = 2008
Score = 450 bits (1157), Expect = e-127
Identities = 244/500 (48%), Positives = 329/500 (65%), Gaps = 10/500 (2%)
Frame = +3
Query: 78 DGPFLSSDSECTGSNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFL-VFGFARTKM 136
D P ++ +LSI V+GASGDLAKKK FPALF L+ + L N + +FG+ARTK+
Sbjct: 198 DSPLSVDNNGPENGSLSIVVLGASGDLAKKKTFPALFNLYKQGFLLANEVCIFGYARTKI 377
Query: 137 TSEELRNMISRTLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGG 196
+ EELRN + L D + + +FL Y SG Y+SE F LD ++ + E
Sbjct: 378 SDEELRNRLRGYLVKEKDASPEKLETVSKFLHLIKYVSGSYDSENDFRLLDKEISKHEST 557
Query: 197 KR-----SNRLFYLSIPPNIFVNV---VRCASLKASSKKGWTRVIVEKPFGRDSESSNEL 248
S RLFYL++PP+++ +V ++ A + S GWTR++VEKPFG+D ES+ +L
Sbjct: 558 TNTAEGSSRRLFYLALPPSVYPSVSKMIKTACMNKSDHGGWTRIVVEKPFGKDLESAEQL 737
Query: 249 TRCLKQYLTEDQIFRIDHYLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGT 308
+ + E QI+RIDHYLGKELV+N+ VLRF+N F PLW+R I NVQ +F EDFGT
Sbjct: 738 STQIGGLFEEPQIYRIDHYLGKELVQNMLVLRFANRFFLPLWNRDNIANVQIVFKEDFGT 917
Query: 309 EGRGGYFDNYGIIRDIMQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENV 368
+GRGGYFD YGIIRDI+QNHLLQI L AME PVS+ E IR+EKVKVL S+ P++ E+V
Sbjct: 918 DGRGGYFDQYGIIRDIIQNHLLQIFCLVAMEKPVSMRPEHIRDEKVKVLESVLPIKDEDV 1097
Query: 369 VTGQYKGHSKGGKSYPGYADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHT 428
V GQY+ GY DDP+VP S TPTFA+ L + N RW+GVPF++KAGKAL +
Sbjct: 1098VLGQYE----------GYRDDPTVPDNSNTPTFASVILRVHNERWEGVPFILKAGKALDS 1247
Query: 429 KRAEIRVQFRHVPGNLYKRNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRS 488
++A+IR+QF+ VPG+++K + NE V+R+QP EA+Y+K+ K PGL M +S
Sbjct: 1248RKADIRIQFKDVPGDIFK-----CQKQGRNEFVMRLQPSEAMYMKLTVKQPGLEMSTVQS 1412
Query: 489 DLNLLYRARY-PTEIPDAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIA 547
+L+L YR RY IP+AYERL+LD I G+++ F+R DEL A+W +FTPLL ++ +
Sbjct: 1413ELDLSYRQRYHDVTIPEAYERLILDTIRGDQQHFVRRDELKASWEIFTPLLHRIDKGEFK 1592
Query: 548 PELYPHGSRGPIGAHYLAAK 567
Y GSRGP A L K
Sbjct: 1593SIPYKSGSRGPKQADELLEK 1652
>TC90186 homologue to PIR|T03740|T03740 glucose-6-phosphate 1-dehydrogenase
(EC 1.1.1.49) TPG18 - common tobacco, partial (38%)
Length = 1168
Score = 344 bits (883), Expect(3) = e-107
Identities = 167/183 (91%), Positives = 175/183 (95%)
Frame = +1
Query: 397 LTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFGADMDKA 456
L PTFAAAALFI NAR DGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFG D+DKA
Sbjct: 475 LPPTFAAAALFIGNAR*DGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFGTDLDKA 654
Query: 457 TNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLLLDAIEG 516
TNELVLRVQPDEAIYLK+NNK+PGLGMRLDRSDLNLLYR+RY EIPDAYERLLLDAIEG
Sbjct: 655 TNELVLRVQPDEAIYLKINNKVPGLGMRLDRSDLNLLYRSRYAREIPDAYERLLLDAIEG 834
Query: 517 ERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRWGDTG 576
ERRLFIRSDELDAAWALFTPLLKE+E+KKIAPELYP+GSRGP+GAHYLAAKHNVRWGD G
Sbjct: 835 ERRLFIRSDELDAAWALFTPLLKEIENKKIAPELYPYGSRGPVGAHYLAAKHNVRWGDFG 1014
Query: 577 NDD 579
DD
Sbjct: 1015GDD 1023
Score = 50.8 bits (120), Expect(3) = e-107
Identities = 21/34 (61%), Positives = 25/34 (72%)
Frame = +3
Query: 372 QYKGHSKGGKSYPGYADDPSVPKGSLTPTFAAAA 405
QYKGHSKGG+SYP Y DD +VP GSLT ++
Sbjct: 399 QYKGHSKGGRSYPAYIDDSTVPMGSLTSNICCSS 500
Score = 33.9 bits (76), Expect(3) = e-107
Identities = 16/23 (69%), Positives = 19/23 (82%)
Frame = +2
Query: 353 KVKVLRSMRPLQLENVVTGQYKG 375
+VKVLRSMRP+QLE+VV G G
Sbjct: 341 QVKVLRSMRPIQLEDVVVGSI*G 409
>TC82969 similar to SP|Q43793|G6PC_TOBAC Glucose-6-phosphate 1-dehydrogenase
chloroplast precursor (EC 1.1.1.49) (G6PD). [Common
tobacco], partial (18%)
Length = 610
Score = 112 bits (279), Expect(3) = 2e-40
Identities = 54/65 (83%), Positives = 60/65 (92%)
Frame = +2
Query: 481 LGMRLDRSDLNLLYRARYPTEIPDAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKE 540
LGM+LDRS+LNL Y ARY EIPDAYERLLLDAIEGERRLFIRSDELDAAW+LFTP+L E
Sbjct: 44 LGMKLDRSNLNLHYAARYSKEIPDAYERLLLDAIEGERRLFIRSDELDAAWSLFTPVLNE 223
Query: 541 VEDKK 545
+E+KK
Sbjct: 224 IEEKK 238
Score = 60.8 bits (146), Expect(3) = 2e-40
Identities = 25/35 (71%), Positives = 30/35 (85%)
Frame = +1
Query: 544 KKIAPELYPHGSRGPIGAHYLAAKHNVRWGDTGND 578
+KI PE YP+GSRGP+ AHYLAA++NVRWGD G D
Sbjct: 232 EKITPEYYPYGSRGPVCAHYLAARYNVRWGDLGLD 336
Score = 32.3 bits (72), Expect(3) = 2e-40
Identities = 13/14 (92%), Positives = 14/14 (99%)
Frame = +3
Query: 467 DEAIYLKVNNKIPG 480
DEAIYLK+NNKIPG
Sbjct: 3 DEAIYLKINNKIPG 44
>AL370546 similar to GP|15810387|gb putative glucose-6-phosphate
dehydrogenase {Arabidopsis thaliana}, partial (18%)
Length = 506
Score = 158 bits (400), Expect = 5e-39
Identities = 71/118 (60%), Positives = 94/118 (79%)
Frame = +3
Query: 455 KATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLLLDAI 514
+ATNEL+LR PDEAI ++VNNK+PGLG++LD S+LNLLY+ +Y E+PD+YE LLLD I
Sbjct: 6 RATNELILRDDPDEAILVRVNNKVPGLGLKLDSSELNLLYKDKYNIEVPDSYEHLLLDVI 185
Query: 515 EGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRW 572
+G+ LF+RSDEL AAW + TP+L E++ ++ ELY G RGP+GA+YL AKH VRW
Sbjct: 186 DGDNHLFMRSDELAAAWNILTPILNEIDKDNVSVELYELGGRGPVGAYYLWAKHAVRW 359
>AL370545 weakly similar to GP|15810387|gb| putative glucose-6-phosphate
dehydrogenase {Arabidopsis thaliana}, partial (11%)
Length = 412
Score = 71.6 bits (174), Expect = 8e-13
Identities = 31/75 (41%), Positives = 48/75 (63%)
Frame = +1
Query: 194 EGGKRSNRLFYLSIPPNIFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLK 253
+G ++NR+FYLS+P ++V C + A ++KGW R+I+EKPFG D+ SS LT+ L
Sbjct: 121 QGRSKTNRIFYLSVPQEALLDVASCLASSAQTQKGWNRIIIEKPFGFDALSSQRLTQYLL 300
Query: 254 QYLTEDQIFRIDHYL 268
E Q++R + L
Sbjct: 301 SKFEEKQLYRYSYLL 345
>TC80117 weakly similar to PIR|T48360|T48360 farnesylated protein-like -
Arabidopsis thaliana, partial (36%)
Length = 880
Score = 29.3 bits (64), Expect = 4.4
Identities = 14/36 (38%), Positives = 17/36 (46%)
Frame = -1
Query: 28 FSCIRPGNLPRNHFQLKSSNGHPLNAVSSHSHDGLA 63
F C P L +HF L + HPL+ V H H A
Sbjct: 751 FLCFSPLFLLTHHFDLSLYSYHPLSLVEPHPHSASA 644
>BQ750979 similar to GP|13883467|gb PE_PGRS family protein {Mycobacterium
tuberculosis CDC1551}, partial (1%)
Length = 728
Score = 28.5 bits (62), Expect = 7.4
Identities = 21/80 (26%), Positives = 34/80 (42%), Gaps = 8/80 (10%)
Frame = +2
Query: 5 LSSSSTIVASYALRNEPQLY--------PVWFSCIRPGNLPRNHFQLKSSNGHPLNAVSS 56
L +++++A+Y R+ PQ + P + +RP L R ++S +N SS
Sbjct: 425 LQRTTSLLAAYQSRSHPQNHKKPLLQPKPPRRTELRPRRLLRARSDPRTSPNLTMNTTSS 604
Query: 57 HSHDGLAGSSLTKEDSKPQP 76
HS L D P P
Sbjct: 605 HSRRNRRLPPLAPSDKLPHP 664
>BG644405 weakly similar to PIR|A84564|A845 hypothetical protein At2g18410
[imported] - Arabidopsis thaliana, partial (8%)
Length = 603
Score = 28.1 bits (61), Expect = 9.7
Identities = 24/88 (27%), Positives = 37/88 (41%), Gaps = 14/88 (15%)
Frame = +2
Query: 202 LFYLSIPPNIFVNVVRCASLKASSKKGWTRV-------------IVEKPFGRDSESSNEL 248
L LS PP+ + +++ SS W RV ++EK R+ L
Sbjct: 95 LVALSRPPSFYAELLKDKGFDVSSSSKWLRVLDCYSDPLGWKKKLMEKGTVRNPYEETSL 274
Query: 249 TRCLKQYLTE-DQIFRIDHYLGKELVEN 275
L + L E D++ LGKE+VE+
Sbjct: 275 KTSLCKNLKELDKVLSSIIELGKEIVED 358
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,746,675
Number of Sequences: 36976
Number of extensions: 250740
Number of successful extensions: 1094
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1071
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1080
length of query: 579
length of database: 9,014,727
effective HSP length: 101
effective length of query: 478
effective length of database: 5,280,151
effective search space: 2523912178
effective search space used: 2523912178
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0213.11


