

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.1
(305 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
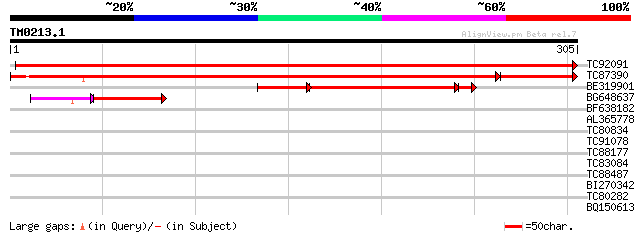
TC92091 similar to PIR|T06010|T06010 hypothetical protein T25K17... 595 e-171
TC87390 similar to GP|20197290|gb|AAC62136.2 expressed protein {... 412 e-134
BE319901 similar to GP|10176768|db gene_id:MIK19.9~pir||T06010~s... 140 4e-45
BG648637 similar to GP|20197290|gb| expressed protein {Arabidops... 46 5e-09
BF638182 weakly similar to PIR|T45594|T45 glucosidase-like prote... 29 2.6
AL365778 similar to GP|1946359|gb|A unknown protein {Arabidopsis... 28 4.4
TC80834 similar to GP|6002279|emb|CAB56741.1 cytochrome P450 mon... 28 4.4
TC91078 homologue to SP|P02828|HS83_DROME Heat shock protein 83 ... 28 5.7
TC88177 homologue to GP|10281004|dbj|BAB13742. pseudo-response r... 28 5.7
TC83084 28 5.7
TC88487 similar to PIR|T01135|T01135 probable GTP-binding protei... 27 7.5
BI270342 similar to GP|1209631|gb| GNOM gene product {Arabidopsi... 27 9.8
TC80282 similar to GP|21554593|gb|AAM63626.1 unknown {Arabidopsi... 27 9.8
BQ150613 similar to GP|10728365|gb CG18830 gene product {Drosoph... 27 9.8
>TC92091 similar to PIR|T06010|T06010 hypothetical protein T25K17.70 -
Arabidopsis thaliana, partial (86%)
Length = 1120
Score = 595 bits (1533), Expect = e-171
Identities = 277/302 (91%), Positives = 290/302 (95%)
Frame = +1
Query: 4 IIDQPDHFGSDVEGKTVPIDEKELVLDGGFVMPHTNSFGHTFRDYNAESERQEGVENFYR 63
+IDQPD+FGSDVE KTV +E ELVLDGGFV P NSFGHTFRDY AESERQEGVENFYR
Sbjct: 1 LIDQPDYFGSDVEAKTVVANETELVLDGGFVKPQANSFGHTFRDYAAESERQEGVENFYR 180
Query: 64 KNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVDESDPDLDEPQIEHLLQTAEA 123
KNHIYQS DFVKKMREEYGKLNRVEMS+WECCELLNEVVDESDPDLDEPQIEHLLQTAEA
Sbjct: 181 KNHIYQSFDFVKKMREEYGKLNRVEMSIWECCELLNEVVDESDPDLDEPQIEHLLQTAEA 360
Query: 124 IRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGDTFPVGCRFDESIVHHKHFKE 183
IRKDYPNEDWLHLTGLIHDLGKVLLLP FGGLPQWAVVGDT+PVGCRFDESIVHHK+FKE
Sbjct: 361 IRKDYPNEDWLHLTGLIHDLGKVLLLPSFGGLPQWAVVGDTYPVGCRFDESIVHHKYFKE 540
Query: 184 NPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVAKENKTTLPSAALFIIRYHSFY 243
NPDYNN AYNTRYG+YSEKCGLNNV+MSWGHDDYMYLVAKENKTTLPSAA+FIIRYHSFY
Sbjct: 541 NPDYNNSAYNTRYGIYSEKCGLNNVMMSWGHDDYMYLVAKENKTTLPSAAMFIIRYHSFY 720
Query: 244 ALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYLSLIEKYFPAKL 303
ALHREGAYKHLMN+ED ENL+WLHIFNKYDLYSKSKVR+DVEKVKPYY+SLIEKYFPAKL
Sbjct: 721 ALHREGAYKHLMNDEDVENLKWLHIFNKYDLYSKSKVRVDVEKVKPYYISLIEKYFPAKL 900
Query: 304 KW 305
KW
Sbjct: 901 KW 906
>TC87390 similar to GP|20197290|gb|AAC62136.2 expressed protein {Arabidopsis
thaliana}, partial (92%)
Length = 1249
Score = 412 bits (1058), Expect(2) = e-134
Identities = 197/276 (71%), Positives = 228/276 (82%), Gaps = 12/276 (4%)
Frame = +3
Query: 1 MTIIIDQPDHFGSDVEGKTVPIDE-KELVLDGGFVMPHT-----------NSFGHTFRDY 48
MTI+++ D FGS + K V ++ ELVLDGGF P N+FGH+FR+Y
Sbjct: 96 MTILVEHVD-FGSQFDDKNVHSEDINELVLDGGFPQPKNASQNTFFAPEINAFGHSFRNY 272
Query: 49 NAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVDESDPD 108
+ ESERQ+GVE FYR HI Q+ DFVKKMREEY KL++ EMS+WECCELLNEVVDESDPD
Sbjct: 273 DEESERQKGVEEFYRLQHINQTYDFVKKMREEYKKLDKAEMSIWECCELLNEVVDESDPD 452
Query: 109 LDEPQIEHLLQTAEAIRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGDTFPVG 168
LDEPQI+HLLQ+AEAIRKDYPNEDWLHLT LIHDLGK+LLLP FG LPQWAVVGDTFP+G
Sbjct: 453 LDEPQIQHLLQSAEAIRKDYPNEDWLHLTALIHDLGKILLLPKFGELPQWAVVGDTFPLG 632
Query: 169 CRFDESIVHHKHFKENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVAKENKTT 228
C FDES VHHK+FKENPD +PAYNT+ G+Y+E CGL+NV+MSWGHDDYM +VAKEN +T
Sbjct: 633 CAFDESNVHHKYFKENPDIKSPAYNTKNGIYNEGCGLDNVMMSWGHDDYMTMVAKENGST 812
Query: 229 LPSAALFIIRYHSFYALHREGAYKHLMNEEDFENLE 264
LP+A LFIIRYHSFY LH+EGAY HLMNEEDFENL+
Sbjct: 813 LPNAGLFIIRYHSFYPLHKEGAYTHLMNEEDFENLK 920
Score = 84.0 bits (206), Expect(2) = e-134
Identities = 36/41 (87%), Positives = 40/41 (96%)
Frame = +1
Query: 265 WLHIFNKYDLYSKSKVRIDVEKVKPYYLSLIEKYFPAKLKW 305
WLHIFNKYDLYSKSKV +DVE+V+PYYLSLIEKYFPAKL+W
Sbjct: 922 WLHIFNKYDLYSKSKVLVDVEEVRPYYLSLIEKYFPAKLRW 1044
>BE319901 similar to GP|10176768|db gene_id:MIK19.9~pir||T06010~similar to
unknown protein {Arabidopsis thaliana}, partial (35%)
Length = 462
Score = 140 bits (354), Expect(3) = 4e-45
Identities = 61/81 (75%), Positives = 72/81 (88%)
Frame = +1
Query: 162 GDTFPVGCRFDESIVHHKHFKENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLV 221
GDTFP+GC FDES VHHK+FKENPD +PAYNT+ G+Y+E CGL+NV+MSWGHDDYM +V
Sbjct: 190 GDTFPLGCAFDESNVHHKYFKENPDIKSPAYNTKNGIYNEGCGLDNVMMSWGHDDYMTMV 369
Query: 222 AKENKTTLPSAALFIIRYHSF 242
AKEN +TLP+A LFIIRYHSF
Sbjct: 370 AKENGSTLPNAGLFIIRYHSF 432
Score = 55.8 bits (133), Expect(3) = 4e-45
Identities = 25/29 (86%), Positives = 26/29 (89%)
Frame = +2
Query: 134 LHLTGLIHDLGKVLLLPVFGGLPQWAVVG 162
LHLT LIHDLGK+LLLP FG LPQWAVVG
Sbjct: 2 LHLTALIHDLGKILLLPKFGELPQWAVVG 88
Score = 23.1 bits (48), Expect(3) = 4e-45
Identities = 8/10 (80%), Positives = 9/10 (90%)
Frame = +2
Query: 242 FYALHREGAY 251
FY LH+EGAY
Sbjct: 431 FYPLHKEGAY 460
>BG648637 similar to GP|20197290|gb| expressed protein {Arabidopsis
thaliana}, partial (17%)
Length = 781
Score = 46.2 bits (108), Expect(2) = 5e-09
Identities = 20/39 (51%), Positives = 29/39 (74%)
Frame = -2
Query: 46 RDYNAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKL 84
R+Y+ ESERQ+GVE FYR HI Q+ DFV K+ + + ++
Sbjct: 264 RNYDEESERQKGVEEFYRLQHINQTYDFVSKVYDYFHQI 148
Score = 36.2 bits (82), Expect = 0.016
Identities = 15/20 (75%), Positives = 18/20 (90%)
Frame = -3
Query: 74 VKKMREEYGKLNRVEMSMWE 93
VKKMREEY KL++ EMS+WE
Sbjct: 77 VKKMREEYKKLDKAEMSIWE 18
Score = 31.2 bits (69), Expect(2) = 5e-09
Identities = 19/47 (40%), Positives = 24/47 (50%), Gaps = 12/47 (25%)
Frame = -1
Query: 12 GSDVEGKTVPIDE-KELVLDGG-----------FVMPHTNSFGHTFR 46
GS + K V ++ ELVLDGG F P N+FGH+FR
Sbjct: 493 GSQFDDKNVHSEDINELVLDGGFPQPKNASQNTFFAPEINAFGHSFR 353
>BF638182 weakly similar to PIR|T45594|T45 glucosidase-like protein -
Arabidopsis thaliana, partial (46%)
Length = 674
Score = 28.9 bits (63), Expect = 2.6
Identities = 9/27 (33%), Positives = 18/27 (66%)
Frame = +1
Query: 177 HHKHFKENPDYNNPAYNTRYGMYSEKC 203
+ +HF +NP++N+P N R+ ++ C
Sbjct: 121 NRRHFPQNPNHNHPH*NLRHKLFHPPC 201
>AL365778 similar to GP|1946359|gb|A unknown protein {Arabidopsis thaliana},
partial (12%)
Length = 467
Score = 28.1 bits (61), Expect = 4.4
Identities = 11/26 (42%), Positives = 16/26 (61%)
Frame = +1
Query: 201 EKCGLNNVLMSWGHDDYMYLVAKENK 226
+K GLNN L SW D +++V E +
Sbjct: 178 DKWGLNNFLESWDLSDMLFIVGPEER 255
>TC80834 similar to GP|6002279|emb|CAB56741.1 cytochrome P450 monooxygenase
{Cicer arietinum}, partial (62%)
Length = 1112
Score = 28.1 bits (61), Expect = 4.4
Identities = 22/84 (26%), Positives = 42/84 (49%), Gaps = 1/84 (1%)
Frame = +3
Query: 36 PHTNSFGHTFRDYNAESERQEGVENFYRKNHIYQS-VDFVKKMREEYGKLNRVEMSMWEC 94
P+ + F + ++ + R+ V + + I++S V KMRE G + ++
Sbjct: 705 PNLSDFFPMLKVFDLQGIRRRSVVSVKKVLSIFRSFVGERLKMREGTGSIGNDDV----L 872
Query: 95 CELLNEVVDESDPDLDEPQIEHLL 118
LLN +D+ ++D+ +IEHLL
Sbjct: 873 DALLNISLDDGKIEMDKDEIEHLL 944
>TC91078 homologue to SP|P02828|HS83_DROME Heat shock protein 83 (HSP 82).
[Fruit fly] {Drosophila melanogaster}, partial (2%)
Length = 682
Score = 27.7 bits (60), Expect = 5.7
Identities = 15/45 (33%), Positives = 24/45 (53%), Gaps = 1/45 (2%)
Frame = -1
Query: 79 EEYGKLNRVEMSM-WECCELLNEVVDESDPDLDEPQIEHLLQTAE 122
E +G+ + ++ M W CC + +E DE +DEP+I L E
Sbjct: 136 ERFGREDLWDLIMVWRCCGVEDEDEDERGVMMDEPRITKSLILCE 2
>TC88177 homologue to GP|10281004|dbj|BAB13742. pseudo-response regulator 7
{Arabidopsis thaliana}, partial (4%)
Length = 1310
Score = 27.7 bits (60), Expect = 5.7
Identities = 14/34 (41%), Positives = 17/34 (49%)
Frame = +2
Query: 227 TTLPSAALFIIRYHSFYALHREGAYKHLMNEEDF 260
TT+P L IIR SF LH + HL +F
Sbjct: 890 TTIPIKMLIIIRLISFQQLHLASLHVHLRRSLEF 991
>TC83084
Length = 257
Score = 27.7 bits (60), Expect = 5.7
Identities = 19/67 (28%), Positives = 30/67 (44%), Gaps = 4/67 (5%)
Frame = -1
Query: 197 GMYSEKCGLNNV----LMSWGHDDYMYLVAKENKTTLPSAALFIIRYHSFYALHREGAYK 252
G+ SE L ++ ++ W H+D V + TT+ Y LHRE YK
Sbjct: 257 GVNSEVLSLIHIHQKQIIKWNHEDLDISVGTVHATTIKC-----------YNLHRETYYK 111
Query: 253 HLMNEED 259
+ +NE +
Sbjct: 110 NGLNENE 90
>TC88487 similar to PIR|T01135|T01135 probable GTP-binding protein (extra
large) [imported] - Arabidopsis thaliana, partial (41%)
Length = 1513
Score = 27.3 bits (59), Expect = 7.5
Identities = 10/36 (27%), Positives = 22/36 (60%)
Frame = +3
Query: 251 YKHLMNEEDFENLEWLHIFNKYDLYSKSKVRIDVEK 286
++ ++ FE +E+L I NK+DL+ + ++ + K
Sbjct: 831 FETMVTHPTFEQMEFLLILNKFDLFEEKVEQVPLTK 938
>BI270342 similar to GP|1209631|gb| GNOM gene product {Arabidopsis thaliana},
partial (6%)
Length = 370
Score = 26.9 bits (58), Expect = 9.8
Identities = 10/28 (35%), Positives = 18/28 (63%)
Frame = +2
Query: 254 LMNEEDFENLEWLHIFNKYDLYSKSKVR 281
L DF L WL++ N+++++ K K+R
Sbjct: 206 LSQSTDFSKL-WLNVLNRFEIFMKVKIR 286
>TC80282 similar to GP|21554593|gb|AAM63626.1 unknown {Arabidopsis
thaliana}, partial (21%)
Length = 927
Score = 26.9 bits (58), Expect = 9.8
Identities = 11/16 (68%), Positives = 12/16 (74%)
Frame = -3
Query: 289 PYYLSLIEKYFPAKLK 304
PYYL LI KYF K+K
Sbjct: 904 PYYLHLIYKYFVTKIK 857
>BQ150613 similar to GP|10728365|gb CG18830 gene product {Drosophila
melanogaster}, partial (1%)
Length = 1151
Score = 26.9 bits (58), Expect = 9.8
Identities = 10/26 (38%), Positives = 16/26 (61%)
Frame = -1
Query: 233 ALFIIRYHSFYALHREGAYKHLMNEE 258
A+F+ RY Y+LH +YK +N +
Sbjct: 971 AMFVSRYDIIYSLHTYQSYKSCLNNQ 894
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.139 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,893,841
Number of Sequences: 36976
Number of extensions: 173422
Number of successful extensions: 726
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 717
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 724
length of query: 305
length of database: 9,014,727
effective HSP length: 96
effective length of query: 209
effective length of database: 5,465,031
effective search space: 1142191479
effective search space used: 1142191479
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0213.1


