

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0212.10
(115 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
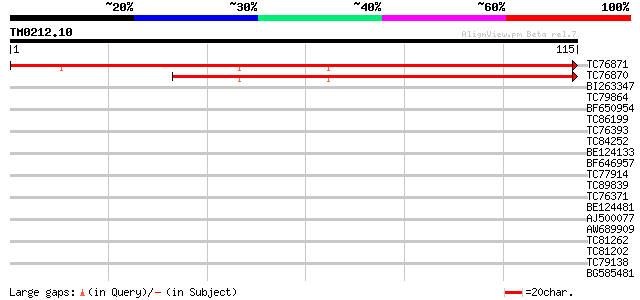
Score E
Sequences producing significant alignments: (bits) Value
TC76871 similar to PIR|H85430|H85430 hypothetical protein AT4g36... 175 2e-45
TC76870 similar to PIR|H85430|H85430 hypothetical protein AT4g36... 135 2e-33
BI263347 31 0.062
TC79864 29 0.31
BF650954 GP|14334122|gb hydroxyproline rich glycoprotein Vsp6 {C... 28 0.52
TC86199 similar to GP|14587306|dbj|BAB61217. hypothetical protei... 28 0.68
TC76393 similar to GP|12049598|emb|CAC19855. mitochondrial succi... 27 0.89
TC84252 similar to PIR|G86297|G86297 hypothetical protein AAD346... 27 0.89
BE124133 weakly similar to GP|1513240|gb|A ORFveg110 {Dictyostel... 27 1.2
BF646957 similar to GP|15215700|gb| AT3g16170/MSL1_21 {Arabidops... 27 1.5
TC77914 homologue to SP|Q9ZRA3|DAD1_PEA Defender against cell de... 26 2.0
TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC ... 26 2.6
TC76371 weakly similar to GP|23304737|emb|CAC87937. PDI-like pro... 26 2.6
BE124481 26 2.6
AJ500077 similar to PIR|T06586|T06 DNA-binding protein PD2 - gar... 26 2.6
AW689909 similar to GP|12850969|dbj DNA segment KIST 6~data sou... 25 3.4
TC81262 similar to GP|21689769|gb|AAM67528.1 putative SIR2-famil... 25 3.4
TC81202 homologue to GP|17380966|gb|AAL36295.1 putative transcri... 25 3.4
TC79138 weakly similar to GP|4115534|dbj|BAA36410.1 UDP-glycose:... 25 3.4
BG585481 weakly similar to PIR|D86358|D863 hypothetical protein ... 25 5.8
>TC76871 similar to PIR|H85430|H85430 hypothetical protein AT4g36500
[imported] - Arabidopsis thaliana, partial (28%)
Length = 664
Score = 175 bits (444), Expect = 2e-45
Identities = 90/123 (73%), Positives = 104/123 (84%), Gaps = 8/123 (6%)
Frame = +2
Query: 1 MLRSFTTRR------YERLGKETASSALLQEGFKRSTSLPSRASSSARKMA-HATFGNIN 53
M RS TTRR YERLGKE+A++ LL E FKRSTS+PSRA++++RKM +TFG IN
Sbjct: 113 MFRSMTTRRGLGNGRYERLGKESATTTLLHEEFKRSTSMPSRATNTSRKMTLGSTFGEIN 292
Query: 54 LQRNPTKKAN-NSSKSTHPLFSFLDFRRKKKTTARPEFTRYLEYLKEGGMWDLNSNKPVI 112
L+RNPTKKAN NSSK +HPL +FLDFRRKKKTTARPEF RY+EYLKEGGMWDLNSNKPVI
Sbjct: 293 LKRNPTKKANSNSSKKSHPLLNFLDFRRKKKTTARPEFARYIEYLKEGGMWDLNSNKPVI 472
Query: 113 HYK 115
+YK
Sbjct: 473 YYK 481
>TC76870 similar to PIR|H85430|H85430 hypothetical protein AT4g36500
[imported] - Arabidopsis thaliana, partial (28%)
Length = 728
Score = 135 bits (341), Expect = 2e-33
Identities = 66/84 (78%), Positives = 76/84 (89%), Gaps = 2/84 (2%)
Frame = +1
Query: 34 PSRASSSARKMA-HATFGNINLQRNPTKKAN-NSSKSTHPLFSFLDFRRKKKTTARPEFT 91
PSRA++++RKM +TFG INL+RNPTKKAN NSSK +HPL +FLDFRRKKKTTARPEF
Sbjct: 307 PSRATNTSRKMTLGSTFGEINLKRNPTKKANSNSSKKSHPLLNFLDFRRKKKTTARPEFA 486
Query: 92 RYLEYLKEGGMWDLNSNKPVIHYK 115
RY+EYLKEGGMWDLNSNKPVI+YK
Sbjct: 487 RYIEYLKEGGMWDLNSNKPVIYYK 558
>BI263347
Length = 582
Score = 31.2 bits (69), Expect = 0.062
Identities = 23/84 (27%), Positives = 39/84 (46%)
Frame = +2
Query: 3 RSFTTRRYERLGKETASSALLQEGFKRSTSLPSRASSSARKMAHATFGNINLQRNPTKKA 62
R+ R + GK++ S ++GF S S+ S ++SS +K + T K
Sbjct: 206 RTENARSGTKSGKQSTPSKFTKKGFTVSKSVKSSSTSSFKKASS--------DDKRTGKR 361
Query: 63 NNSSKSTHPLFSFLDFRRKKKTTA 86
+SKST S + FR K +++
Sbjct: 362 TTASKSTGSKLSVVGFRGKNASSS 433
>TC79864
Length = 1551
Score = 28.9 bits (63), Expect = 0.31
Identities = 24/89 (26%), Positives = 35/89 (38%), Gaps = 8/89 (8%)
Frame = +1
Query: 15 KETASSALLQEGFKRSTSLPSRASSSARKMAHATFGNINLQRNPTKKAN--------NSS 66
KET S QE S+P R SS+A ++ + RNP +N +
Sbjct: 316 KETLESLANQEN-----SIPERESSAAARIDEQKEAALAALRNPNPPSNIAAALKSLSRK 480
Query: 67 KSTHPLFSFLDFRRKKKTTARPEFTRYLE 95
L F+ +RK+ RPE L+
Sbjct: 481 MDAESLLMFVISKRKESIMLRPEIAAALK 567
>BF650954 GP|14334122|gb hydroxyproline rich glycoprotein Vsp6 {Chlamydomonas
reinhardtii}, partial (1%)
Length = 662
Score = 28.1 bits (61), Expect = 0.52
Identities = 22/68 (32%), Positives = 31/68 (45%)
Frame = +1
Query: 4 SFTTRRYERLGKETASSALLQEGFKRSTSLPSRASSSARKMAHATFGNINLQRNPTKKAN 63
SFTT+R T+SS+ +T + + S R +F LQRNP +N
Sbjct: 169 SFTTQR------STSSSSTFTRTTITNTIIKPQCLSRIRSRYSRSF----LQRNPHHDSN 318
Query: 64 NSSKSTHP 71
SS + HP
Sbjct: 319 PSSTTQHP 342
>TC86199 similar to GP|14587306|dbj|BAB61217. hypothetical protein~similar
to Arabidopsis thaliana chromosome 3 MSL1.21, partial
(15%)
Length = 735
Score = 27.7 bits (60), Expect = 0.68
Identities = 16/44 (36%), Positives = 23/44 (51%)
Frame = +3
Query: 65 SSKSTHPLFSFLDFRRKKKTTARPEFTRYLEYLKEGGMWDLNSN 108
SS STHP FSF FR +++ + ++E LK D S+
Sbjct: 126 SSSSTHPTFSFSQFRSSSHSSS--PSSSFMEVLKAIAKQDFASH 251
>TC76393 similar to GP|12049598|emb|CAC19855. mitochondrial succinate
dehydrogenase iron-sulphur subunit {Arabidopsis
thaliana}, partial (37%)
Length = 637
Score = 27.3 bits (59), Expect = 0.89
Identities = 23/85 (27%), Positives = 38/85 (44%)
Frame = +3
Query: 17 TASSALLQEGFKRSTSLPSRASSSARKMAHATFGNINLQRNPTKKANNSSKSTHPLFSFL 76
T +S +L+ R +S P+ + R AHA+ ++A+ ++ T L F
Sbjct: 186 TMASTMLKRAIHRISSSPTSRLTLLR--AHASEAQ-------AQQASPKARDTTVLKKFQ 338
Query: 77 DFRRKKKTTARPEFTRYLEYLKEGG 101
+R T ++PE Y LKE G
Sbjct: 339 IYRWNPDTPSKPELKEYEINLKECG 413
>TC84252 similar to PIR|G86297|G86297 hypothetical protein AAD34681.1
[imported] - Arabidopsis thaliana, partial (32%)
Length = 724
Score = 27.3 bits (59), Expect = 0.89
Identities = 21/81 (25%), Positives = 35/81 (42%), Gaps = 14/81 (17%)
Frame = -2
Query: 29 RSTSLPSRASSSARKMAHATFG-----NINLQRNPTKKANNSSKSTHPLFSFLDF----- 78
+ TS ASS R + A G +++ NP + +NN + ++ LFS
Sbjct: 699 KPTSRTRXASSKTRNLVLAKTGAISGLSLSKTLNPPRSSNNKMRPSNQLFSLSKHIHTTN 520
Query: 79 ----RRKKKTTARPEFTRYLE 95
+ K+ T RP+ R L+
Sbjct: 519 NHRNPKPKRRTKRPKLLRQLK 457
>BE124133 weakly similar to GP|1513240|gb|A ORFveg110 {Dictyostelium
discoideum}, partial (7%)
Length = 639
Score = 26.9 bits (58), Expect = 1.2
Identities = 16/33 (48%), Positives = 18/33 (54%)
Frame = +3
Query: 18 ASSALLQEGFKRSTSLPSRASSSARKMAHATFG 50
+ SALLQ K STS SR + RK AT G
Sbjct: 492 SQSALLQTTLKNSTSSNSRKNQVHRKSLLATLG 590
>BF646957 similar to GP|15215700|gb| AT3g16170/MSL1_21 {Arabidopsis
thaliana}, partial (34%)
Length = 643
Score = 26.6 bits (57), Expect = 1.5
Identities = 17/40 (42%), Positives = 20/40 (49%)
Frame = +3
Query: 40 SARKMAHATFGNINLQRNPTKKANNSSKSTHPLFSFLDFR 79
S RK+ H+ FG + SS STHP FSF FR
Sbjct: 198 SFRKLPHS-FGYVT-----------SSSSTHPTFSFSQFR 281
>TC77914 homologue to SP|Q9ZRA3|DAD1_PEA Defender against cell death 1
(DAD-1) (Peadad). [Garden pea] {Pisum sativum}, partial
(96%)
Length = 704
Score = 26.2 bits (56), Expect = 2.0
Identities = 15/36 (41%), Positives = 22/36 (60%), Gaps = 1/36 (2%)
Frame = -1
Query: 54 LQRNPTKKANNSSKSTHPLFSFLDF-RRKKKTTARP 88
LQR + KA + ++S + LFS L + RR+ TA P
Sbjct: 449 LQRTKSAKARSGARSLNSLFSLLTWIRRQTAKTAVP 342
>TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC 4.6.1.1)
(ATP pyrophosphate-lyase) (Adenylyl cyclase). [Yeast],
partial (0%)
Length = 665
Score = 25.8 bits (55), Expect = 2.6
Identities = 16/59 (27%), Positives = 28/59 (47%), Gaps = 1/59 (1%)
Frame = +2
Query: 14 GKETASSALLQEGFKRSTSLPSRASSSARKMAHATFGNINLQRN-PTKKANNSSKSTHP 71
G+E + +G+KRST+ P++AS I + N P+K + S + +P
Sbjct: 179 GREVN*PCIFIDGYKRSTACPTKAS-------------ITISANVPSKASTTGSSAVYP 316
>TC76371 weakly similar to GP|23304737|emb|CAC87937. PDI-like protein
{Quercus suber}, partial (21%)
Length = 1710
Score = 25.8 bits (55), Expect = 2.6
Identities = 16/32 (50%), Positives = 17/32 (53%)
Frame = -1
Query: 3 RSFTTRRYERLGKETASSALLQEGFKRSTSLP 34
RS RRYE L ETASS+ ST LP
Sbjct: 219 RSLEARRYENLKLETASSS------SSSTKLP 142
>BE124481
Length = 538
Score = 25.8 bits (55), Expect = 2.6
Identities = 16/59 (27%), Positives = 28/59 (47%), Gaps = 1/59 (1%)
Frame = +2
Query: 14 GKETASSALLQEGFKRSTSLPSRASSSARKMAHATFGNINLQRN-PTKKANNSSKSTHP 71
G+E + +G+KRST+ P++AS I + N P+K + S + +P
Sbjct: 182 GREVN*PCIFIDGYKRSTACPTKAS-------------ITISANVPSKASTTGSSAVYP 319
>AJ500077 similar to PIR|T06586|T06 DNA-binding protein PD2 - garden pea,
partial (28%)
Length = 578
Score = 25.8 bits (55), Expect = 2.6
Identities = 12/44 (27%), Positives = 23/44 (52%)
Frame = -1
Query: 33 LPSRASSSARKMAHATFGNINLQRNPTKKANNSSKSTHPLFSFL 76
L + S+++ + T INL+ + N++ K + PLF+ L
Sbjct: 179 LDGQLGGSSQRRINTTVDRINLESWDGRSKNSAKKESIPLFALL 48
>AW689909 similar to GP|12850969|dbj DNA segment KIST 6~data source:MGD
source key:MGI:108435 evidence:ISS~putative {Mus
musculus}, partial (19%)
Length = 605
Score = 25.4 bits (54), Expect = 3.4
Identities = 13/31 (41%), Positives = 16/31 (50%)
Frame = +3
Query: 60 KKANNSSKSTHPLFSFLDFRRKKKTTARPEF 90
+ A K T P L FRRK TT+ P+F
Sbjct: 111 RTATTWKKITTPQPPTLQFRRKTNTTSAPKF 203
>TC81262 similar to GP|21689769|gb|AAM67528.1 putative SIR2-family protein
{Arabidopsis thaliana}, partial (28%)
Length = 680
Score = 25.4 bits (54), Expect = 3.4
Identities = 25/100 (25%), Positives = 42/100 (42%)
Frame = +3
Query: 16 ETASSALLQEGFKRSTSLPSRASSSARKMAHATFGNINLQRNPTKKANNSSKSTHPLFSF 75
E++SS ++ F S S + ARK+ +I L +PT ++ N S L +F
Sbjct: 63 ESSSSL*IEISFSMSPSSSFTSLVFARKVLGTIITDIALCPSPTTQSWNLSTKGGQLVAF 242
Query: 76 LDFRRKKKTTARPEFTRYLEYLKEGGMWDLNSNKPVIHYK 115
R +T+ R + G + +N KP + K
Sbjct: 243 KGGARFIQTSCR---------ISAPGTFPVNDGKPQLRDK 335
>TC81202 homologue to GP|17380966|gb|AAL36295.1 putative transcription
factor {Arabidopsis thaliana}, partial (40%)
Length = 1506
Score = 25.4 bits (54), Expect = 3.4
Identities = 11/20 (55%), Positives = 14/20 (70%)
Frame = +2
Query: 54 LQRNPTKKANNSSKSTHPLF 73
L+ + T K NNSS S+H LF
Sbjct: 575 LEVSTTSKINNSSSSSHELF 634
>TC79138 weakly similar to GP|4115534|dbj|BAA36410.1 UDP-glycose:flavonoid
glycosyltransferase {Vigna mungo}, partial (36%)
Length = 1575
Score = 25.4 bits (54), Expect = 3.4
Identities = 13/42 (30%), Positives = 20/42 (46%)
Frame = +1
Query: 73 FSFLDFRRKKKTTARPEFTRYLEYLKEGGMWDLNSNKPVIHY 114
+SF+ F +KKK R + L L +GG+ + K Y
Sbjct: 1180 WSFIGFMKKKKIVGRDIIEKALRRLMDGGIEAVEIRKRAQEY 1305
>BG585481 weakly similar to PIR|D86358|D863 hypothetical protein AAF18527.1
[imported] - Arabidopsis thaliana, partial (12%)
Length = 720
Score = 24.6 bits (52), Expect = 5.8
Identities = 12/37 (32%), Positives = 23/37 (61%)
Frame = +2
Query: 16 ETASSALLQEGFKRSTSLPSRASSSARKMAHATFGNI 52
ETA+++L++E +ST S++S +A FG++
Sbjct: 608 ETANASLVEEHIVKSTPDASQSSQAATSRDIEDFGSV 718
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.127 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,147,486
Number of Sequences: 36976
Number of extensions: 37276
Number of successful extensions: 283
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 277
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 279
length of query: 115
length of database: 9,014,727
effective HSP length: 91
effective length of query: 24
effective length of database: 5,649,911
effective search space: 135597864
effective search space used: 135597864
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 50 (23.9 bits)
Lotus: description of TM0212.10


