

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0209.4
(342 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
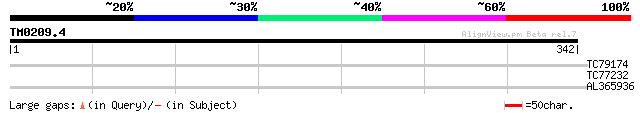
Score E
Sequences producing significant alignments: (bits) Value
TC79174 homologue to PIR|T13029|T13029 beta-adaptin homolog F8L2... 29 3.0
TC77232 similar to SP|O04130|SERA_ARATH D-3-phosphoglycerate deh... 28 5.1
AL365936 27 8.7
>TC79174 homologue to PIR|T13029|T13029 beta-adaptin homolog F8L21.170 -
Arabidopsis thaliana, partial (27%)
Length = 830
Score = 28.9 bits (63), Expect = 3.0
Identities = 17/55 (30%), Positives = 27/55 (48%)
Frame = -2
Query: 194 VDVSARTILVSTLGILAKVPSLGGRIRPLGVGGALPIWRSGLASSSFATAIRAIM 248
+ SA T L +T+G+L+ + S +PL S S SFAT + A++
Sbjct: 655 ISASAATALATTIGLLSDIKSFSDSKKPLS------STNSAFMSYSFATQMAAVL 509
>TC77232 similar to SP|O04130|SERA_ARATH D-3-phosphoglycerate dehydrogenase
chloroplast precursor (EC 1.1.1.95) (3-PGDH). [Mouse-ear
cress], partial (87%)
Length = 2474
Score = 28.1 bits (61), Expect = 5.1
Identities = 12/35 (34%), Positives = 22/35 (62%)
Frame = +3
Query: 40 PKMRNDNQSCDTTPTLTQSESRTINSLFTSMTSLL 74
P ++N S +TPTL +S T+NSL ++ +++
Sbjct: 177 PNLQNPLTSHSSTPTLLESYPTTLNSLMLTLNNVV 281
>AL365936
Length = 441
Score = 27.3 bits (59), Expect = 8.7
Identities = 25/66 (37%), Positives = 29/66 (43%), Gaps = 13/66 (19%)
Frame = +2
Query: 1 MKVSQPK--MASMRPPYVR---------QTPCDPHGREPQKKLIVGDLPCPKMRNDN--Q 47
MK PK M RPP R QTP + + P KLI+G PK RN + Q
Sbjct: 113 MKFPNPKLIMGMPRPPKSRNPSLKQSTHQTPQNQTMKFPNPKLIMGMPRPPKSRNPSLKQ 292
Query: 48 SCDTTP 53
S TP
Sbjct: 293 STHQTP 310
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,230,151
Number of Sequences: 36976
Number of extensions: 132360
Number of successful extensions: 602
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 584
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 602
length of query: 342
length of database: 9,014,727
effective HSP length: 97
effective length of query: 245
effective length of database: 5,428,055
effective search space: 1329873475
effective search space used: 1329873475
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0209.4


