

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0209.3
(1547 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
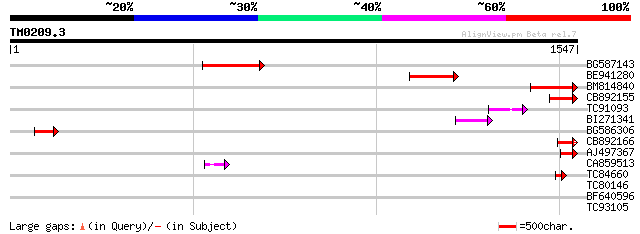
Score E
Sequences producing significant alignments: (bits) Value
BG587143 weakly similar to PIR|A96586|A9 hypothetical protein F2... 179 1e-44
BE941280 weakly similar to GP|20197614|g unknown protein {Arabid... 171 2e-42
BM814840 weakly similar to GP|15128241|db helicase-like protein ... 151 2e-36
CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292... 99 2e-20
TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein... 74 4e-13
BI271341 71 3e-12
BG586306 weakly similar to GP|15128241|db helicase-like protein ... 69 1e-11
CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis ... 61 4e-09
AJ497367 similar to GP|14140286|gb putative helicase {Oryza sati... 59 1e-08
CA859513 54 5e-07
TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG282... 44 6e-04
TC80146 40 0.009
BF640596 similar to GP|17104563|gb| putative N-hydroxycinnamoyl/... 31 4.3
TC93105 similar to GP|21655261|gb|AAM28907.1 putative TIR/NBS/LR... 30 5.6
>BG587143 weakly similar to PIR|A96586|A9 hypothetical protein F20D21.24
[imported] - Arabidopsis thaliana, partial (12%)
Length = 717
Score = 179 bits (453), Expect = 1e-44
Identities = 83/171 (48%), Positives = 117/171 (67%), Gaps = 1/171 (0%)
Frame = -3
Query: 525 FPAGMPTIEFQKRGFPHAHILLWL*PQHKIKTGDDIDKHISAELPDPKLYPKLYKAVSSY 584
+ + IEFQKRG HAHILLW + + +++D+ ISAELP+ K P+ Y V+ +
Sbjct: 601 YTTALHRIEFQKRGLRHAHILLWFGNSSRTPSSEEVDEIISAELPNKKQDPEAYNLVTKH 422
Query: 585 MIHGPCGPIDPKSVCMVDGKCSKHFPKKFQNCTTVDDDGFPIYKRRTTRIT-VTKKGVPL 643
MIHGPCG I+PKS CM + C+K +P+ + T++D G+ +Y+RR V K G L
Sbjct: 421 MIHGPCGVINPKSPCMENNVCTKKYPRPYNGSTSIDKSGYVLYRRR*NETEHVVKNGAIL 242
Query: 644 DNGFVVPYNPKLLMKYQGHINVEYCNKSNAIKYLFKYINKGPDRVNVQISK 694
+N F+VP+N KLL KY+ HIN+E+CN+++A+KYLFKYI KG DRV+V I K
Sbjct: 241 NNTFIVPHNIKLLKKYEAHINMEWCNRTSAVKYLFKYITKGVDRVSVVIEK 89
>BE941280 weakly similar to GP|20197614|g unknown protein {Arabidopsis
thaliana}, partial (8%)
Length = 403
Score = 171 bits (434), Expect = 2e-42
Identities = 88/134 (65%), Positives = 100/134 (73%)
Frame = -1
Query: 1091 LYGFGGTGKTFVWNTLSAAVRSKGLIVLNVASSGIASLLLPGGRTAHSRFSILISINEVS 1150
LY +GGT KTF+W LSAA+RS+G IVL ASSGI +LL+PGGRTAHSRF I I+E S
Sbjct: 403 LYDYGGTEKTFIWRALSAALRSEGEIVLACASSGIDALLMPGGRTAHSRFGIPFIIDETS 224
Query: 1151 TCNLRQGSPKAELLQKASLIIWDETPMMNKHCFEALDRSLNDVMKTRANYGNDRPFRGKV 1210
C + P A L+ KA LIIWDE PMM+KHCFEALDRSL DV+KT D PF GKV
Sbjct: 223 MCGVTPNIPLASLVIKAKLIIWDEAPMMHKHCFEALDRSLRDVLKTVDERNKDIPFGGKV 44
Query: 1211 VVLGGDFRQILPVI 1224
VVLGGDFRQIL V+
Sbjct: 43 VVLGGDFRQILLVM 2
>BM814840 weakly similar to GP|15128241|db helicase-like protein {Oryza sativa
(japonica cultivar-group)}, partial (6%)
Length = 733
Score = 151 bits (382), Expect = 2e-36
Identities = 76/127 (59%), Positives = 94/127 (73%)
Frame = +3
Query: 1421 NGTRLRVTHLTQYIIVATVLSGIRLGKTEYIPRITLTPSDSGLPFKFSRRQFPVTLCFAM 1480
+GTRL + L + +I A V+ G G+ YIPR+ L PS + + F R QFP+ L FAM
Sbjct: 27 HGTRLIIVSLGKNVICARVIGGTHAGEVSYIPRMNLIPSGANVSITFERCQFPLVLSFAM 206
Query: 1481 TINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKVLIVVEQGVVSTSTRNVVY 1540
TINKSQGQ+L+ VGLYLPRPVFTHGQLYVA+SRVKSR GLK+LI E G S+ST NVVY
Sbjct: 207 TINKSQGQTLTSVGLYLPRPVFTHGQLYVAVSRVKSRSGLKILITDENGSPSSSTVNVVY 386
Query: 1541 KEVFENV 1547
+EVF+ +
Sbjct: 387 QEVFQKI 407
>CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292.1 [imported]
- Arabidopsis thaliana, partial (4%)
Length = 572
Score = 98.6 bits (244), Expect = 2e-20
Identities = 49/74 (66%), Positives = 58/74 (78%)
Frame = -2
Query: 1474 VTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKVLIVVEQGVVST 1533
V + FAMTINKSQGQSL H+G+YLP VF+HGQLYVALSRV SR+GLK+LI + G
Sbjct: 370 VEVYFAMTINKSQGQSLKHIGVYLPSSVFSHGQLYVALSRVTSREGLKILISNDDGEDDC 191
Query: 1534 STRNVVYKEVFENV 1547
T NVVY+EVF N+
Sbjct: 190 VTSNVVYREVFHNL 149
>TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein T24M8.10 -
Arabidopsis thaliana, partial (11%)
Length = 701
Score = 73.9 bits (180), Expect = 4e-13
Identities = 41/106 (38%), Positives = 61/106 (56%)
Frame = +3
Query: 1307 DPLLELVNFAYPDLVANLESDSYFQERAILAPTLESVVHVNNYILSKLPGVEREYLSYDT 1366
DP+ +V YP+LV+ ++ + Q RAIL T E V +N+Y+L +PG ER S +
Sbjct: 390 DPIDAIVQSTYPNLVSQYNNEQFLQSRAILTSTDEVVDQINDYVLKLIPGEERVIYSAN- 566
Query: 1367 PCRSDEDSEVHAEWFTSEFLNDVQCSGIPNHRLILKERVPIMLLRN 1412
RS+ + + EFL ++ S +PNH+L LK PIMLLR+
Sbjct: 567 --RSEVNDVQAFDAIPPEFLQSLKTSDLPNHKLTLKVGTPIMLLRD 698
>BI271341
Length = 468
Score = 71.2 bits (173), Expect = 3e-12
Identities = 39/101 (38%), Positives = 61/101 (59%)
Frame = +1
Query: 1217 FRQILPVISKGSRADMVGSTVTSSYLWKYCKVMKLTVNMRLQSASS*TSATEIREFAQWI 1276
FR ILPVI +GSR+D++ +T+ SS + +C+V++L NM LQ ++ E +F++ I
Sbjct: 1 FR*ILPVIPRGSRSDIIHATINSSCI*DHCQVVRLKKNMWLQQNGQSSNDPEFEQFSK*I 180
Query: 1277 LKVGDETVDTIDEDETTIEIPSDLLIGQGPDPLLELVNFAY 1317
LKVGD + ++ I+IP +LLI D L +V Y
Sbjct: 181 LKVGDGKIYEPNDSYADIDIPPELLISNYDDSLQTIVQSTY 303
>BG586306 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (2%)
Length = 667
Score = 68.9 bits (167), Expect = 1e-11
Identities = 30/64 (46%), Positives = 45/64 (69%)
Frame = +2
Query: 69 FKDNIRAYNSMFAFTSMGGKVQNSINDGGGPPKFILSGQNYHRIGSLVPREGQTPKFAQL 128
F+D IR YNS+ AFTS+G K+ S+ + G + GQ +HRI SL+PR+G+ P++ Q+
Sbjct: 11 FRDTIRVYNSVLAFTSIGMKMDYSVVNAPGRYTIRIQGQTHHRIDSLIPRQGRPPEYLQI 190
Query: 129 YIYD 132
YI+D
Sbjct: 191 YIFD 202
>CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 748
Score = 60.8 bits (146), Expect = 4e-09
Identities = 31/52 (59%), Positives = 39/52 (74%)
Frame = -1
Query: 1496 YLPRPVFTHGQLYVALSRVKSRKGLKVLIVVEQGVVSTSTRNVVYKEVFENV 1547
+ R VF+HGQLYVA+SRV SR GLK+L++ E G +T NVVYK VF+NV
Sbjct: 289 FYDREVFSHGQLYVAISRVSSRNGLKILMIDENGDCIDNTTNVVYK-VFQNV 137
>AJ497367 similar to GP|14140286|gb putative helicase {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 543
Score = 58.9 bits (141), Expect = 1e-08
Identities = 28/46 (60%), Positives = 36/46 (77%)
Frame = +1
Query: 1502 FTHGQLYVALSRVKSRKGLKVLIVVEQGVVSTSTRNVVYKEVFENV 1547
F++G+LYVA+SRV SRKGLK+L+ E G +T NVVYKEVF N+
Sbjct: 19 FSNGKLYVAVSRVTSRKGLKILLAHEDGNCMNTTSNVVYKEVFRNL 156
>CA859513
Length = 363
Score = 53.9 bits (128), Expect = 5e-07
Identities = 30/68 (44%), Positives = 38/68 (55%)
Frame = +1
Query: 533 EFQKRGFPHAHILLWL*PQHKIKTGDDIDKHISAELPDPKLYPKLYKAVSSYMIHGPCGP 592
EF+K PHAH+L H+ D+ D +SAEL DP +LY+ V S MIHGP GP
Sbjct: 148 EFEKCDLPHAHMLF-----HR---ADNYDTFVSAELSDPVEQLRLYQTVVSVMIHGPYGP 303
Query: 593 IDPKSVCM 600
+ CM
Sbjct: 304 FNNNVPCM 327
>TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (1%)
Length = 1009
Score = 43.5 bits (101), Expect = 6e-04
Identities = 20/30 (66%), Positives = 25/30 (82%)
Frame = +2
Query: 1489 SLSHVGLYLPRPVFTHGQLYVALSRVKSRK 1518
S++ G+YLP+P+F HG LYVALSRV SRK
Sbjct: 68 SMATKGMYLPQPIF*HG*LYVALSRVTSRK 157
>TC80146
Length = 476
Score = 39.7 bits (91), Expect = 0.009
Identities = 16/35 (45%), Positives = 27/35 (76%)
Frame = -3
Query: 1474 VTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLY 1508
+ + + TINKS+ QSLS++ +YL RP+F+H ++Y
Sbjct: 471 IQIPYFKTINKSR*QSLSYMKIYLSRPIFSHEEMY 367
>BF640596 similar to GP|17104563|gb| putative
N-hydroxycinnamoyl/benzoyltransferase {Arabidopsis
thaliana}, partial (20%)
Length = 325
Score = 30.8 bits (68), Expect = 4.3
Identities = 21/59 (35%), Positives = 32/59 (53%), Gaps = 1/59 (1%)
Frame = +3
Query: 1002 ELMNLCLIEIEKELQCNARSLKDFPSLSYPTFAEIMNFE-NKFVADELNYNKEEMSKIH 1059
+LMN C+ E + N R + S + T +I+ F+ +F+A LN N EEM K+H
Sbjct: 36 QLMN-CM*LKENRFEFNPRKKQKRDSTFFQTLTKILQFQCAQFIA--LNQNLEEMKKLH 203
>TC93105 similar to GP|21655261|gb|AAM28907.1 putative TIR/NBS/LRR disease
resistance protein {Pinus taeda}, partial (6%)
Length = 761
Score = 30.4 bits (67), Expect = 5.6
Identities = 12/27 (44%), Positives = 18/27 (66%)
Frame = +1
Query: 773 HLPNQQKVLFKESDDFEEVVQKCSRKE 799
H PN+ K L K+ D +E+V KCS ++
Sbjct: 271 HSPNETKTLIKQMKDGKELVLKCSNED 351
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.331 0.145 0.455
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 50,466,552
Number of Sequences: 36976
Number of extensions: 752179
Number of successful extensions: 4095
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 3007
Number of HSP's successfully gapped in prelim test: 118
Number of HSP's that attempted gapping in prelim test: 1027
Number of HSP's gapped (non-prelim): 3221
length of query: 1547
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1438
effective length of database: 4,984,343
effective search space: 7167485234
effective search space used: 7167485234
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0209.3


