

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
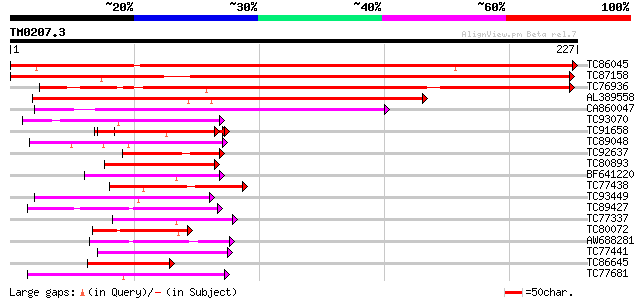
Query= TM0207.3
(227 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC86045 similar to PIR|T46225|T46225 alpha NAC-like protein - Ar... 380 e-106
TC87158 similar to GP|21426124|gb|AAM52321.1 Putative nascent po... 346 4e-96
TC76936 similar to GP|20160782|dbj|BAB89723. putative nascent po... 242 7e-65
AL389558 similar to PIR|T30826|T30 nascent polypeptide-associate... 122 1e-28
CA860047 weakly similar to GP|15081680|gb| AT3g12390/T2E22_130 {... 114 3e-26
TC93070 weakly similar to GP|17065230|gb|AAL32769.1 Unknown prot... 50 5e-07
TC91658 similar to GP|14329812|emb|CAC40753. putative nucleosome... 49 1e-06
TC89048 weakly similar to PIR|T46215|T46215 hypothetical protein... 49 2e-06
TC92637 similar to PIR|T39903|T39903 serine-rich protein - fissi... 45 2e-05
TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {... 45 2e-05
BF641220 weakly similar to PIR|G86203|G86 probable N-arginine di... 45 3e-05
TC77438 homologue to PIR|T06807|T06807 nucleosome assembly prote... 44 5e-05
TC93449 similar to GP|21328382|gb|AAM48550.1 Hypothetical protei... 44 5e-05
TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed pr... 44 7e-05
TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Atel... 43 9e-05
TC80072 similar to GP|14423520|gb|AAK62442.1 Unknown protein {Ar... 43 9e-05
AW688281 similar to PIR|T06370|T0 probable DNA (cytosine-5-)-met... 42 2e-04
TC77441 similar to GP|21592359|gb|AAM64310.1 unknown {Arabidopsi... 41 4e-04
TC86645 similar to PIR|H86321|H86321 hypothetical protein AAF271... 41 4e-04
TC77681 similar to GP|22597156|gb|AAN03465.1 nucleolar histone d... 40 6e-04
>TC86045 similar to PIR|T46225|T46225 alpha NAC-like protein - Arabidopsis
thaliana, partial (69%)
Length = 1166
Score = 380 bits (977), Expect = e-106
Identities = 194/229 (84%), Positives = 213/229 (92%), Gaps = 2/229 (0%)
Frame = +1
Query: 1 MAPGPVVDA-SSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDED 59
MAPGPV+DA ++E+E +Q+ S +ETTLKKKPQP+EDDAPI+EDVKDDDKD D+D+DED
Sbjct: 94 MAPGPVIDAPATESEVEQQIPSVDETTLKKKPQPEEDDAPIIEDVKDDDKD--DEDDDED 267
Query: 60 DDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISK 119
DDDEDDD ++ AL G++GSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISK
Sbjct: 268 DDDEDDDDKEDALAGADGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISK 447
Query: 120 PDVFKSPNSETYVIFGEAKIEDLSSQLQTQAAQQFRMPDMGSVMAKQDQGTAADGGQP-E 178
PDVFKSPNSETY+IFGEAKIEDLSSQLQTQAAQQFRMPDMGSVMAKQDQG +A G P E
Sbjct: 448 PDVFKSPNSETYIIFGEAKIEDLSSQLQTQAAQQFRMPDMGSVMAKQDQGASASGAIPEE 627
Query: 179 EEEEVDETGVEPHDIDLVMTQAGVSRGKAVKALKTHNGDIVGAIMELTT 227
EEEE+DETGVEPHDIDLVMTQAGVS+ KAVKALKTHNGDIVGAIMELTT
Sbjct: 628 EEEEIDETGVEPHDIDLVMTQAGVSKSKAVKALKTHNGDIVGAIMELTT 774
>TC87158 similar to GP|21426124|gb|AAM52321.1 Putative nascent polypeptide
associated complex alpha chain {Oryza sativa (japonica
cultivar-group)}, partial (76%)
Length = 937
Score = 346 bits (888), Expect = 4e-96
Identities = 176/227 (77%), Positives = 199/227 (87%), Gaps = 1/227 (0%)
Frame = +3
Query: 1 MAPGPVVDASSEAEALEQVLSAEETTLKKKPQPQE-DDAPIVEDVKDDDKDEADDDEDED 59
M+PGP++DA+++ A +Q+ S +ETTLK KP PQE DDAPIVED DDDKDE++D+ED++
Sbjct: 108 MSPGPIIDAAADGAADQQLPSVDETTLKNKPLPQEEDDAPIVEDDNDDDKDESEDEEDDE 287
Query: 60 DDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISK 119
GG+EGSKQSRSEKKSRKAMLKLGLKP+ GV+RVTIKRTKN++FFISK
Sbjct: 288 SPQ----------GGTEGSKQSRSEKKSRKAMLKLGLKPIDGVTRVTIKRTKNVMFFISK 437
Query: 120 PDVFKSPNSETYVIFGEAKIEDLSSQLQTQAAQQFRMPDMGSVMAKQDQGTAADGGQPEE 179
PDVFKSPNSETYVIFGEAKIEDLSSQLQTQAAQQFRMPDMGS+MAKQ+QG AADG QPEE
Sbjct: 438 PDVFKSPNSETYVIFGEAKIEDLSSQLQTQAAQQFRMPDMGSIMAKQEQGAAADGAQPEE 617
Query: 180 EEEVDETGVEPHDIDLVMTQAGVSRGKAVKALKTHNGDIVGAIMELT 226
EEEVDETGVE HDI+LVMTQAGVS+ KAVKALKTHNGDIVGAIMELT
Sbjct: 618 EEEVDETGVEAHDIELVMTQAGVSKAKAVKALKTHNGDIVGAIMELT 758
>TC76936 similar to GP|20160782|dbj|BAB89723. putative nascent polypeptide
associated complex alpha chain {Oryza sativa (japonica
cultivar-group)}, partial (86%)
Length = 914
Score = 242 bits (618), Expect = 7e-65
Identities = 135/215 (62%), Positives = 164/215 (75%), Gaps = 1/215 (0%)
Frame = +3
Query: 13 AEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGAL 72
A+ E++L+A + Q D P+VED DDD+D D++DEDDDDEDDD +G
Sbjct: 81 AQTQEELLAAH-----LEQQKIHHDEPVVED--DDDED---DEDDEDDDDEDDDNAEGFE 230
Query: 73 GGSEG-SKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISKPDVFKSPNSETY 131
G + G SKQ+RSEKKSRKAMLKLG+K VTGVSRVTIK++KNILF ISKPDVFKSP S+TY
Sbjct: 231 GDASGRSKQTRSEKKSRKAMLKLGMKAVTGVSRVTIKKSKNILFVISKPDVFKSPTSDTY 410
Query: 132 VIFGEAKIEDLSSQLQTQAAQQFRMPDMGSVMAKQDQGTAADGGQPEEEEEVDETGVEPH 191
+IFGEAKIEDLSSQLQTQAA+QF+ P++ + AK + G E+E+VDETGV+P
Sbjct: 411 IIFGEAKIEDLSSQLQTQAAEQFKAPNLTNAGAKPE-----SSGIAPEDEDVDETGVDPK 575
Query: 192 DIDLVMTQAGVSRGKAVKALKTHNGDIVGAIMELT 226
DI+LV+TQAGV R +AVKALK NGDIV AIMELT
Sbjct: 576 DIELVVTQAGVPRSRAVKALKAANGDIVAAIMELT 680
>AL389558 similar to PIR|T30826|T30 nascent polypeptide-associated complex
alpha chain muscle splice form gp220 - mouse, partial
(3%)
Length = 493
Score = 122 bits (305), Expect = 1e-28
Identities = 68/163 (41%), Positives = 99/163 (60%), Gaps = 5/163 (3%)
Frame = +2
Query: 10 SSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+SE +E++ ET + K + +E + E++ D+D + +E+
Sbjct: 5 TSEKIKIEEIEVESETKEEVKIESSSKQISEIESTSIKKTTTVAESEEDSDEDIPELEEE 184
Query: 70 G----ALGGSEGSK-QSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISKPDVFK 124
G + ++ SK QSR EKK+RK M KLGLK V G++RVTI+R KNIL +S PDV+K
Sbjct: 185 GIVPPGIAQADISKIQSRGEKKARKQMQKLGLKVVPGINRVTIRRPKNILLVVSNPDVYK 364
Query: 125 SPNSETYVIFGEAKIEDLSSQLQTQAAQQFRMPDMGSVMAKQD 167
SPNS+ Y++FGEAK+EDL+SQ Q AAQQF+ + + A QD
Sbjct: 365 SPNSDCYIVFGEAKVEDLNSQAQANAAQQFQAAETAASSAAQD 493
>CA860047 weakly similar to GP|15081680|gb| AT3g12390/T2E22_130 {Arabidopsis
thaliana}, partial (37%)
Length = 543
Score = 114 bits (285), Expect = 3e-26
Identities = 65/142 (45%), Positives = 86/142 (59%)
Frame = +3
Query: 11 SEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDG 70
SE+E +Q + +E T E+ +E + E D D+D + +E+ G
Sbjct: 141 SESEKKKQSVKIQEIT--------EESGGALEKKTTTVESEEDSDDDIPELEEETAIPSG 296
Query: 71 ALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRVTIKRTKNILFFISKPDVFKSPNSET 130
QSR EKK+RKAM KLGLK V G++RVTI+R KNILF +S PDV+KS NS+
Sbjct: 297 VAPVDITKMQSRGEKKARKAMQKLGLKHVPGINRVTIRRPKNILFVVSNPDVYKSANSDC 476
Query: 131 YVIFGEAKIEDLSSQLQTQAAQ 152
Y++FGEAKIEDL+SQ Q AAQ
Sbjct: 477 YIVFGEAKIEDLNSQAQANAAQ 542
>TC93070 weakly similar to GP|17065230|gb|AAL32769.1 Unknown protein
{Arabidopsis thaliana}, partial (19%)
Length = 679
Score = 50.4 bits (119), Expect = 5e-07
Identities = 28/82 (34%), Positives = 49/82 (59%), Gaps = 1/82 (1%)
Frame = +1
Query: 6 VVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVE-DVKDDDKDEADDDEDEDDDDED 64
V+ A S + L+ + ++T K P + D I E D +DD+ ++ D+DED+DDDD+D
Sbjct: 142 VLSAHSNPKILK---TNRKSTYGKFLSPYDSDDEIEEMDFEDDEDEDEDEDEDDDDDDDD 312
Query: 65 DDKEDGALGGSEGSKQSRSEKK 86
DD++D G +E + + +K+
Sbjct: 313 DDEDDDDDGFAEPTDLNAKDKR 378
Score = 30.4 bits (67), Expect = 0.58
Identities = 19/60 (31%), Positives = 30/60 (49%)
Frame = +1
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKL 94
ED+ +D DDD D+ DDD+D + D + +D L S+ + EK+ K + L
Sbjct: 268 EDEDEDEDDDDDDDDDDEDDDDDGFAEPTDLNAKDKRL-KSKTVPDRQQEKEKEKGVRSL 444
Score = 28.1 bits (61), Expect = 2.9
Identities = 16/64 (25%), Positives = 29/64 (45%)
Frame = +1
Query: 32 QPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAM 91
+ +++D +D DDD DE DDD+ + + + K+ + +Q EK+
Sbjct: 262 EDEDEDEDEDDDDDDDDDDEDDDDDGFAEPTDLNAKDKRLKSKTVPDRQQEKEKEKGVRS 441
Query: 92 LKLG 95
L G
Sbjct: 442 LNNG 453
>TC91658 similar to GP|14329812|emb|CAC40753. putative nucleosome assembly
protein 1 {Atropa belladonna}, partial (46%)
Length = 583
Score = 49.3 bits (116), Expect = 1e-06
Identities = 22/51 (43%), Positives = 36/51 (70%), Gaps = 2/51 (3%)
Frame = +2
Query: 36 DDAPIVEDVKDDDKDEADDDEDEDDD--DEDDDKEDGALGGSEGSKQSRSE 84
DD + ED +D D+++ DDD+D+DDD DEDD++ED G S+ + S+++
Sbjct: 71 DDIEVDEDDEDGDEEDDDDDDDDDDDEEDEDDEEEDEGKGKSKSKRGSKAK 223
Score = 45.4 bits (106), Expect = 2e-05
Identities = 19/45 (42%), Positives = 29/45 (64%)
Frame = +2
Query: 43 DVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKS 87
D +DDD D+ DDD++ED+DDE++D+ G GSK + +S
Sbjct: 107 DEEDDDDDDDDDDDEEDEDDEEEDEGKGKSKSKRGSKAKPGKDQS 241
Score = 43.1 bits (100), Expect = 9e-05
Identities = 20/54 (37%), Positives = 35/54 (64%)
Frame = +2
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSR 88
E D +E +DD+ + +DD+D+DDDD+D++ ED EG +S+S++ S+
Sbjct: 59 ESDFDDIEVDEDDEDGDEEDDDDDDDDDDDEEDEDDE-EEDEGKGKSKSKRGSK 217
Score = 38.5 bits (88), Expect = 0.002
Identities = 19/54 (35%), Positives = 30/54 (55%)
Frame = +2
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLG 95
+D++ D+ DE D DE++DDDD+DDD ++ E + K R + K G
Sbjct: 71 DDIEVDEDDE-DGDEEDDDDDDDDDDDEEDEDDEEEDEGKGKSKSKRGSKAKPG 229
Score = 35.0 bits (79), Expect = 0.024
Identities = 17/59 (28%), Positives = 30/59 (50%)
Frame = +2
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLK 93
++D +D DDD D+ ++DED++++DE K G +K + + R A K
Sbjct: 95 DEDGDEEDDDDDDDDDDDEEDEDDEEEDEGKGKSKSKRGSK--AKPGKDQSTERPADCK 265
Score = 34.7 bits (78), Expect = 0.031
Identities = 21/61 (34%), Positives = 28/61 (45%)
Frame = +2
Query: 46 DDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRV 105
D D E D+D DED D+EDDD +D E ++ K+ K G K G +
Sbjct: 65 DFDDIEVDED-DEDGDEEDDDDDDDDDDDEEDEDDEEEDEGKGKSKSKRGSKAKPGKDQS 241
Query: 106 T 106
T
Sbjct: 242 T 244
Score = 34.3 bits (77), Expect = 0.040
Identities = 17/48 (35%), Positives = 25/48 (51%)
Frame = +2
Query: 34 QEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQS 81
+EDD +D DDD DE D+D++E+D+ + K G QS
Sbjct: 110 EEDD----DDDDDDDDDEEDEDDEEEDEGKGKSKSKRGSKAKPGKDQS 241
Score = 29.3 bits (64), Expect = 1.3
Identities = 15/54 (27%), Positives = 27/54 (49%)
Frame = +2
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSR 88
E+D +D DD++DE D++EDE + A G + S + ++ K +
Sbjct: 110 EEDDDDDDDDDDDEEDEDDEEEDEGKGKSKSKRGSKAKPGKDQSTERPADCKQQ 271
>TC89048 weakly similar to PIR|T46215|T46215 hypothetical protein T8P19.220
- Arabidopsis thaliana, partial (35%)
Length = 1329
Score = 48.5 bits (114), Expect = 2e-06
Identities = 32/92 (34%), Positives = 49/92 (52%), Gaps = 13/92 (14%)
Frame = +2
Query: 9 ASSEAEALEQVLSAE-ETTLKKKPQPQED------DAPIVEDVKD------DDKDEADDD 55
+S EAL+ + E K +P+P+E+ DA + E + +D D+ D+D
Sbjct: 299 SSMATEALDDNKPKQVENEAKDEPKPKEEVEVTEQDAEVAESDEKKEIDGHEDSDKDDED 478
Query: 56 EDEDDDDEDDDKEDGALGGSEGSKQSRSEKKS 87
E D+DD DDD+E+ A +G K+ SEKKS
Sbjct: 479 EGGDEDDADDDEEEDAGEEKKGVKKESSEKKS 574
Score = 39.7 bits (91), Expect = 0.001
Identities = 25/69 (36%), Positives = 35/69 (50%), Gaps = 5/69 (7%)
Frame = +1
Query: 24 ETTLKKKPQPQEDDAPIV-EDVKDDDKDEADDDEDEDDDDED----DDKEDGALGGSEGS 78
ET KKK Q E DV++DD+ E D +DE +DED ++ D +G SEG
Sbjct: 1108 ETPAKKKKQTSESGKKRKPSDVEEDDQAELSDAKDESQEDEDVAVANNGSDNEVGKSEGE 1287
Query: 79 KQSRSEKKS 87
+ + KS
Sbjct: 1288 EDTPKAHKS 1314
>TC92637 similar to PIR|T39903|T39903 serine-rich protein - fission yeast
(Schizosaccharomyces pombe), partial (7%)
Length = 671
Score = 45.1 bits (105), Expect = 2e-05
Identities = 20/41 (48%), Positives = 28/41 (67%)
Frame = +1
Query: 46 DDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKK 86
DDD +E D+DE+ED++D DDD D GG +GS + R +K
Sbjct: 445 DDDAEEVDEDEEEDEEDYDDDDYD---GGGKGSSRKRQYRK 558
>TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {Arabidopsis
thaliana}, partial (39%)
Length = 883
Score = 45.1 bits (105), Expect = 2e-05
Identities = 19/46 (41%), Positives = 28/46 (60%)
Frame = +3
Query: 39 PIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSE 84
P ++ K D +D+ DDD+D+D DEDDD E+ G EG ++ E
Sbjct: 294 PAYKENKSDTEDDEDDDDDDDVQDEDDDGEEEDYSGDEGEEEGDPE 431
Score = 39.3 bits (90), Expect = 0.001
Identities = 20/61 (32%), Positives = 33/61 (53%)
Frame = +3
Query: 24 ETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRS 83
E L +KP +E+ + D +DD+ D+ DDD ++DDD +++ G G EG +
Sbjct: 273 EHLLGRKPAYKENKS----DTEDDEDDDDDDDVQDEDDDGEEEDYSGDEGEEEGDPEDDP 440
Query: 84 E 84
E
Sbjct: 441 E 443
Score = 37.7 bits (86), Expect = 0.004
Identities = 23/68 (33%), Positives = 40/68 (58%), Gaps = 4/68 (5%)
Frame = +3
Query: 6 VVDASSEAEALEQVLSAEETTLKKKPQ--PQEDDAPIVEDVKDDDKD--EADDDEDEDDD 61
V D + E E+ S +E + P+ P+ + A +D +DDD D E D+++ ED++
Sbjct: 357 VQDEDDDGE--EEDYSGDEGEEEGDPEDDPEANGAGGSDDGEDDDDDGDEEDEEDGEDEE 530
Query: 62 DEDDDKED 69
DE+D++ED
Sbjct: 531 DEEDEEED 554
Score = 36.2 bits (82), Expect = 0.011
Identities = 22/86 (25%), Positives = 39/86 (44%), Gaps = 6/86 (6%)
Frame = +3
Query: 5 PVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDD---- 60
P + A LE +L + + K ++D+ +D DDD + DDD +E+D
Sbjct: 237 PKLTAMDSRFPLEHLLGRKPAYKENKSDTEDDE----DDDDDDDVQDEDDDGEEEDYSGD 404
Query: 61 --DDEDDDKEDGALGGSEGSKQSRSE 84
++E D ++D G+ GS +
Sbjct: 405 EGEEEGDPEDDPEANGAGGSDDGEDD 482
Score = 34.3 bits (77), Expect = 0.040
Identities = 13/32 (40%), Positives = 25/32 (77%)
Frame = +3
Query: 35 EDDAPIVEDVKDDDKDEADDDEDEDDDDEDDD 66
EDD +D ++D+++ +D+EDE+D++EDD+
Sbjct: 474 EDDD---DDGDEEDEEDGEDEEDEEDEEEDDE 560
Score = 33.9 bits (76), Expect = 0.052
Identities = 20/62 (32%), Positives = 28/62 (44%), Gaps = 12/62 (19%)
Frame = +3
Query: 35 EDDAPIVEDVKDDDKDEADDDEDE------------DDDDEDDDKEDGALGGSEGSKQSR 82
EDD ED D+ +E D ED+ +DDD+D D+ED G E ++
Sbjct: 366 EDDDGEEEDYSGDEGEEEGDPEDDPEANGAGGSDDGEDDDDDGDEEDEEDGEDEEDEEDE 545
Query: 83 SE 84
E
Sbjct: 546 EE 551
Score = 32.3 bits (72), Expect = 0.15
Identities = 15/54 (27%), Positives = 25/54 (45%)
Frame = +3
Query: 36 DDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRK 89
+D P D E DDD+ +++D+ED + E+ E + + K RK
Sbjct: 429 EDDPEANGAGGSDDGEDDDDDGDEEDEEDGEDEEDEEDEEEDDETPQPPAKKRK 590
>BF641220 weakly similar to PIR|G86203|G86 probable N-arginine dibasic
convertase [imported] - Arabidopsis thaliana, partial
(5%)
Length = 634
Score = 44.7 bits (104), Expect = 3e-05
Identities = 20/62 (32%), Positives = 37/62 (59%), Gaps = 6/62 (9%)
Frame = +2
Query: 31 PQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDD------DKEDGALGGSEGSKQSRSE 84
P+ D I ED +++D ++ +DDE++DD+ EDD D+++ + G EG K + ++
Sbjct: 329 PEGAPKDGSIDEDDEEEDDEDEEDDEEDDDEGEDDEDEEXEDEDEXXVXGREGGKGAANQ 508
Query: 85 KK 86
K
Sbjct: 509 SK 514
Score = 37.0 bits (84), Expect = 0.006
Identities = 18/49 (36%), Positives = 30/49 (60%), Gaps = 1/49 (2%)
Frame = +2
Query: 34 QEDDAPIVEDVKDDDKDEADDDED-EDDDDEDDDKEDGALGGSEGSKQS 81
+EDD +D +DDD+ E D+DE+ ED+D+ +G G + SK++
Sbjct: 374 EEDDEDEEDDEEDDDEGEDDEDEEXEDEDEXXVXGREGGKGAANQSKKA 520
>TC77438 homologue to PIR|T06807|T06807 nucleosome assembly protein 1 - garden
pea, partial (93%)
Length = 1607
Score = 43.9 bits (102), Expect = 5e-05
Identities = 25/56 (44%), Positives = 35/56 (61%), Gaps = 1/56 (1%)
Frame = +2
Query: 41 VEDVKDDDKDEA-DDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLG 95
++D DD+ D+A DDDEDED+D+++DD ED +K+ S KKS A L G
Sbjct: 1034 LDDEDDDEDDDAEDDDEDEDEDEDEDDDEDEE---ETKTKKKSSAKKSGIAQLGEG 1192
Score = 38.9 bits (89), Expect = 0.002
Identities = 18/46 (39%), Positives = 31/46 (67%)
Frame = +2
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKS 87
E+ D D ++ D+D+D +DDDED+D ED E ++++++KKS
Sbjct: 1019 EEFGDLDDEDDDEDDDAEDDDEDED-EDEDEDDDEDEEETKTKKKS 1153
Score = 36.6 bits (83), Expect = 0.008
Identities = 15/48 (31%), Positives = 27/48 (56%)
Frame = +2
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRK 89
E + ++ + DD++D++DDD +DD ED E + E K++K
Sbjct: 1004 EAAQGEEFGDLDDEDDDEDDDAEDDDEDEDEDEDEDDDEDEEETKTKK 1147
Score = 32.3 bits (72), Expect = 0.15
Identities = 16/45 (35%), Positives = 29/45 (63%)
Frame = +2
Query: 27 LKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGA 71
L + ++DDA +D +D+D+DE D+D+DED+++ K+ A
Sbjct: 1034 LDDEDDDEDDDAE--DDDEDEDEDE-DEDDDEDEEETKTKKKSSA 1159
Score = 30.8 bits (68), Expect = 0.44
Identities = 15/51 (29%), Positives = 24/51 (46%)
Frame = +2
Query: 55 DEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKLGLKPVTGVSRV 105
D D++DDDEDDD ED E + E + K +G++++
Sbjct: 1031 DLDDEDDDEDDDAEDDDEDEDEDEDEDDDEDEEETKTKKKSSAKKSGIAQL 1183
>TC93449 similar to GP|21328382|gb|AAM48550.1 Hypothetical protein W02D3.12
{Caenorhabditis elegans}, partial (33%)
Length = 631
Score = 43.9 bits (102), Expect = 5e-05
Identities = 21/75 (28%), Positives = 44/75 (58%), Gaps = 3/75 (4%)
Frame = +3
Query: 11 SEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKD---EADDDEDEDDDDEDDDK 67
S + ++ ++L + ++ + ++ P E + DDDK E +++E+++DDD D +K
Sbjct: 213 SVSNSVTRLLFSSSSSTHHHNEKEQTQTPKPESLHDDDKQSQKEEEEEEEDNDDDIDMNK 392
Query: 68 EDGALGGSEGSKQSR 82
E G +GG +G + +R
Sbjct: 393 ETGEVGGPKGPEPTR 437
>TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed protein
{Arabidopsis thaliana}, partial (50%)
Length = 753
Score = 43.5 bits (101), Expect = 7e-05
Identities = 25/78 (32%), Positives = 40/78 (51%)
Frame = +1
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDK 67
D + E + ++ S +E + P++D P DDD DE DDD+D+D+DD +D+
Sbjct: 340 DVNDEDDDNDEDFSGDEDD--EDADPEDDPVPNGAGGSDDD-DEDDDDDDDDNDDGEDED 510
Query: 68 EDGALGGSEGSKQSRSEK 85
ED E S+ +K
Sbjct: 511 EDEEEDDDEDQPPSKKKK 564
Score = 39.7 bits (91), Expect = 0.001
Identities = 24/73 (32%), Positives = 34/73 (45%), Gaps = 4/73 (5%)
Frame = +1
Query: 16 LEQVLSAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDED----DDDEDDDKEDGA 71
L+Q+ T + K + +D +D DDD ++ DDD DED +DDED D ED
Sbjct: 250 LDQLSKGNHTCEENKDGSETEDDDDEDD--DDDVNDEDDDNDEDFSGDEDDEDADPEDDP 423
Query: 72 LGGSEGSKQSRSE 84
+ G E
Sbjct: 424 VPNGAGGSDDDDE 462
>TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Ateline
herpesvirus 3}, partial (8%)
Length = 986
Score = 43.1 bits (100), Expect = 9e-05
Identities = 21/51 (41%), Positives = 30/51 (58%), Gaps = 1/51 (1%)
Frame = +3
Query: 42 EDVKDDDKDEADDDEDEDDDDEDD-DKEDGALGGSEGSKQSRSEKKSRKAM 91
ED +D D + DDDEDEDDDDE++ +ED G E + E + +A+
Sbjct: 600 EDDEDGDDQDEDDDEDEDDDDEEEGGEEDEEEGVDEEDNEEEEEDEDEEAL 752
Score = 35.8 bits (81), Expect = 0.014
Identities = 17/55 (30%), Positives = 32/55 (57%)
Frame = +3
Query: 32 QPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKK 86
Q ++DD ED DDD++E ++++E+ DE+D++E+ E + + KK
Sbjct: 624 QDEDDD----EDEDDDDEEEGGEEDEEEGVDEEDNEEEEEDEDEEALQPPKKRKK 776
Score = 34.7 bits (78), Expect = 0.031
Identities = 16/38 (42%), Positives = 24/38 (63%)
Frame = +3
Query: 47 DDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSE 84
D + +A+ ++ DD+DE+DD DGA G EG + SE
Sbjct: 396 DKQHDAEGEDGNDDEDEEDDDGDGAFG--EGEDELSSE 503
Score = 34.7 bits (78), Expect = 0.031
Identities = 26/89 (29%), Positives = 42/89 (46%), Gaps = 4/89 (4%)
Frame = +3
Query: 1 MAPGPVVDASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDVKD--DDKDEADDDEDE 58
M +V A+ A L ++ + T K + Q D + +D DD+DE DDD D
Sbjct: 288 MLASAIVKATLLAPELYALMLTDMPTTKGDSKTQSQDKQHDAEGEDGNDDEDEEDDDGDG 467
Query: 59 DDDDEDDD--KEDGALGGSEGSKQSRSEK 85
+ +D+ EDG G+ + +S S+K
Sbjct: 468 AFGEGEDELSSEDGGGYGNNSNNKSNSKK 554
>TC80072 similar to GP|14423520|gb|AAK62442.1 Unknown protein {Arabidopsis
thaliana}, partial (17%)
Length = 746
Score = 43.1 bits (100), Expect = 9e-05
Identities = 21/45 (46%), Positives = 30/45 (66%), Gaps = 5/45 (11%)
Frame = +1
Query: 34 QEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDD-----KEDGALG 73
++D+ PI D KD D DE D+D DE++DD+D+D KE G +G
Sbjct: 442 EDDEMPIFND-KDIDNDETDEDGDEEEDDDDEDDEYDGKEKGFIG 573
Score = 26.9 bits (58), Expect = 6.4
Identities = 18/56 (32%), Positives = 25/56 (44%), Gaps = 13/56 (23%)
Frame = +1
Query: 34 QEDDAPIVEDVKDDDKDEADD-----DEDEDDD--------DEDDDKEDGALGGSE 76
Q D P+ + + DE D+ D+D D+D +EDDD ED G E
Sbjct: 391 QRDIVPLDINADSAESDEDDEMPIFNDKDIDNDETDEDGDEEEDDDDEDDEYDGKE 558
>AW688281 similar to PIR|T06370|T0 probable DNA
(cytosine-5-)-methyltransferase (EC 2.1.1.37) - garden
pea, partial (14%)
Length = 685
Score = 42.0 bits (97), Expect = 2e-04
Identities = 24/58 (41%), Positives = 34/58 (58%)
Frame = +3
Query: 33 PQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRKA 90
P++ I +VKDDD DEA++ E E++DDED + E L EG ++ S K KA
Sbjct: 423 PEDSKEVIASEVKDDD-DEAEEQEQEENDDEDAEVETVLL---EGMQKPHSVSKQTKA 584
>TC77441 similar to GP|21592359|gb|AAM64310.1 unknown {Arabidopsis
thaliana}, partial (98%)
Length = 1046
Score = 40.8 bits (94), Expect = 4e-04
Identities = 20/54 (37%), Positives = 30/54 (55%)
Frame = +3
Query: 36 DDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKEDGALGGSEGSKQSRSEKKSRK 89
+D I V DD D + +DE DDD+D+D + L E K+ R+++K RK
Sbjct: 387 EDNIIPRSVDADDSDVEVNSDDESDDDDDEDDTEALLAELEQIKKERAQEKLRK 548
>TC86645 similar to PIR|H86321|H86321 hypothetical protein AAF27100.1
[imported] - Arabidopsis thaliana, partial (76%)
Length = 1116
Score = 40.8 bits (94), Expect = 4e-04
Identities = 16/35 (45%), Positives = 25/35 (70%)
Frame = +3
Query: 32 QPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDD 66
+ +EDD + KDDD E DDD+++DDDDE+++
Sbjct: 735 EAEEDDDEAGDAGKDDDDSEDDDDQEDDDDDEEEE 839
Score = 38.1 bits (87), Expect = 0.003
Identities = 15/37 (40%), Positives = 22/37 (58%)
Frame = +3
Query: 31 PQPQEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDDK 67
P E+D D DD D DDD+ EDDDD+++++
Sbjct: 729 PDEAEEDDDEAGDAGKDDDDSEDDDDQEDDDDDEEEE 839
Score = 36.6 bits (83), Expect = 0.008
Identities = 16/32 (50%), Positives = 22/32 (68%), Gaps = 5/32 (15%)
Frame = +3
Query: 43 DVKDDDKDEA-----DDDEDEDDDDEDDDKED 69
D ++D DEA DDD+ EDDDD++DD +D
Sbjct: 732 DEAEEDDDEAGDAGKDDDDSEDDDDQEDDDDD 827
Score = 34.3 bits (77), Expect = 0.040
Identities = 11/28 (39%), Positives = 21/28 (74%)
Frame = +3
Query: 42 EDVKDDDKDEADDDEDEDDDDEDDDKED 69
++ ++DD + D +D+DD ++DDD+ED
Sbjct: 732 DEAEEDDDEAGDAGKDDDDSEDDDDQED 815
Score = 32.0 bits (71), Expect = 0.20
Identities = 12/34 (35%), Positives = 20/34 (58%)
Frame = +3
Query: 36 DDAPIVEDVKDDDKDEADDDEDEDDDDEDDDKED 69
+D E+ D+ D DD+D +DDD+ +D +D
Sbjct: 723 EDPDEAEEDDDEAGDAGKDDDDSEDDDDQEDDDD 824
Score = 30.4 bits (67), Expect = 0.58
Identities = 12/41 (29%), Positives = 25/41 (60%)
Frame = +3
Query: 21 SAEETTLKKKPQPQEDDAPIVEDVKDDDKDEADDDEDEDDD 61
++E+ ++ + DA +D +DD D+ DDD+DE+++
Sbjct: 717 NSEDPDEAEEDDDEAGDAGKDDDDSEDDDDQEDDDDDEEEE 839
>TC77681 similar to GP|22597156|gb|AAN03465.1 nucleolar histone deacetylase
HD2-P39 {Glycine max}, partial (47%)
Length = 1233
Score = 40.4 bits (93), Expect = 6e-04
Identities = 20/83 (24%), Positives = 42/83 (50%), Gaps = 2/83 (2%)
Frame = +2
Query: 8 DASSEAEALEQVLSAEETTLKKKPQPQEDDAPIVEDV--KDDDKDEADDDEDEDDDDEDD 65
DA + A A + + K + +DD +++ DD+ + AD D +++DD ++D
Sbjct: 500 DAKNAAPAKNAATAKHVKVVDPKDEDSDDDDESDDEIGSSDDEMENADSDSEDEDDSDED 679
Query: 66 DKEDGALGGSEGSKQSRSEKKSR 88
D+E+ + + K+ +E S+
Sbjct: 680 DEEETPVKKVDQGKKRPNESASK 748
Score = 36.2 bits (82), Expect = 0.011
Identities = 30/103 (29%), Positives = 46/103 (44%), Gaps = 19/103 (18%)
Frame = +2
Query: 7 VDASSEAEALEQVLSAEETTLK--KKPQPQEDDAPIVEDVKDDDKDEADDD--------- 55
V +A+A + T K K P+++D+ +D + DD+ + DD
Sbjct: 479 VSEPKKADAKNAAPAKNAATAKHVKVVDPKDEDSD--DDDESDDEIGSSDDEMENADSDS 652
Query: 56 EDEDDDDEDDDKEDGALGGSEGSKQSR--------SEKKSRKA 90
EDEDD DEDD++E +G K+ S KKS+ A
Sbjct: 653 EDEDDSDEDDEEETPVKKVDQGKKRPNESASKTPISSKKSKNA 781
Score = 34.3 bits (77), Expect = 0.040
Identities = 31/110 (28%), Positives = 49/110 (44%), Gaps = 9/110 (8%)
Frame = +2
Query: 12 EAEALEQVLSAEETTLKKKPQPQEDDAPI----VEDVKDDDKDEADDDEDEDDDDEDDDK 67
E +A E +S + K P ++ A V D KD+D D DDDE +D+ DD+
Sbjct: 455 EPKAEELKVSEPKKADAKNAAPAKNAATAKHVKVVDPKDEDSD--DDDESDDEIGSSDDE 628
Query: 68 EDGALGGSEGSKQSRSEKKSRKAMLKLG---LKPVTGVSRVTI--KRTKN 112
+ A SE S + + + K+ +P S+ I K++KN
Sbjct: 629 MENADSDSEDEDDSDEDDEEETPVKKVDQGKKRPNESASKTPISSKKSKN 778
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.305 0.127 0.336
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,280,720
Number of Sequences: 36976
Number of extensions: 51695
Number of successful extensions: 3832
Number of sequences better than 10.0: 361
Number of HSP's better than 10.0 without gapping: 962
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2053
length of query: 227
length of database: 9,014,727
effective HSP length: 93
effective length of query: 134
effective length of database: 5,575,959
effective search space: 747178506
effective search space used: 747178506
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0207.3


