

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.9
(529 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
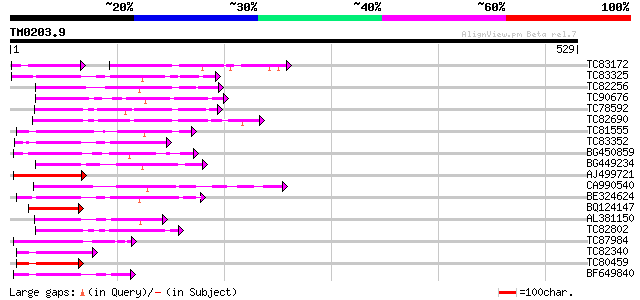
Score E
Sequences producing significant alignments: (bits) Value
TC83172 weakly similar to GP|20042907|gb|AAM08735.1 Hypothetical... 55 6e-17
TC83325 similar to PIR|D84455|D84455 hypothetical protein At2g04... 84 2e-16
TC82256 similar to GP|20042913|gb|AAM08741.1 Unknown protein {Or... 77 2e-14
TC90676 similar to GP|6503308|gb|AAF14684.1| Contains PF|00646 F... 75 5e-14
TC78592 similar to GP|9802755|gb|AAF99824.1| Unknown protein {Ar... 72 4e-13
TC82690 weakly similar to GP|9802755|gb|AAF99824.1| Unknown prot... 72 7e-13
TC81555 weakly similar to GP|20042914|gb|AAM08742.1 Unknown prot... 63 2e-10
TC83352 weakly similar to GP|20042917|gb|AAM08745.1 Unknown prot... 63 2e-10
BG450859 weakly similar to GP|21553400|gb| unknown {Arabidopsis ... 62 4e-10
BG449234 weakly similar to GP|8979942|gb|A Contains a F-box PF|0... 61 9e-10
AJ499721 weakly similar to PIR|T48291|T482 hypothetical protein ... 61 1e-09
CA990540 weakly similar to GP|10092259|gb| hypothetical protein;... 60 2e-09
BE324624 weakly similar to GP|20042914|gb| Unknown protein {Oryz... 60 2e-09
BQ124147 similar to GP|10177836|dbj emb|CAB62440.1~gene_id:MCD7.... 60 3e-09
AL381150 weakly similar to GP|14028988|gb| Unknown protein {Oryz... 60 3e-09
TC82802 similar to GP|9802755|gb|AAF99824.1| Unknown protein {Ar... 59 5e-09
TC87984 weakly similar to GP|10177368|dbj|BAB10659. contains sim... 59 6e-09
TC82340 similar to GP|10178240|dbj|BAB11672. gene_id:MDJ22.8~unk... 58 1e-08
TC80459 weakly similar to GP|22831085|dbj|BAC15947. contains EST... 58 1e-08
BF649840 weakly similar to GP|8979942|gb|A Contains a F-box PF|0... 57 2e-08
>TC83172 weakly similar to GP|20042907|gb|AAM08735.1 Hypothetical protein
{Oryza sativa (japonica cultivar-group)}, partial (31%)
Length = 1097
Score = 55.5 bits (132), Expect(2) = 6e-17
Identities = 32/69 (46%), Positives = 37/69 (53%)
Frame = +2
Query: 2 ETRSAKRKKMAQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLW 61
ET RKK N+ DR+SDLPD V+ HILS L A QT ILSKRW LW
Sbjct: 152 ETMKPGRKKY-----NDDVKEDEDRLSDLPDCVILHILSFLDTIDAVQTCILSKRWNNLW 316
Query: 62 TTFPDLDFR 70
P L+ +
Sbjct: 317 KYLPTLNIK 343
Score = 50.1 bits (118), Expect(2) = 6e-17
Identities = 56/185 (30%), Positives = 91/185 (48%), Gaps = 15/185 (8%)
Frame = +3
Query: 94 LASTMDFITQVLSTRDKDSDIRALCF-RARLSFSRL-NSLIRSAVRHNVKELDIEVTPEV 151
L+S+ F++++L+ R+ + +RAL F R + RL +I+ AV H+V+EL I ++ ++
Sbjct: 339 LSSSNKFVSRILTLRNASTPLRALHFQRHGTMYPRLLQRMIKYAVSHHVQELSINLSSDI 518
Query: 152 RTDEYFNFPRCLIASKTLRVLKLRSGF------RLPPSSIMKETFQSLHTLSLTLGPVD- 204
+ +FP CL + TL LKL L PSS+ +L S T PVD
Sbjct: 519 Q-----HFPTCLFSCHTLTSLKLVINHPTLYAGTLFPSSLNLPALATLFLESFTF-PVDD 680
Query: 205 --HQAPLSDLFTESAFPFLRNLHLQLCFGLRVLRVGCLA--LRDLSLEK--CINLHGLDV 258
H P SAF L +L ++ C L + L+ L +L+L+ N +++
Sbjct: 681 DGHTEPF------SAFSSLNSLIIRGCKVLDEQNLWILSATLANLTLDTGWAYNYGKIEL 842
Query: 259 VTPKL 263
TP L
Sbjct: 843 STPSL 857
>TC83325 similar to PIR|D84455|D84455 hypothetical protein At2g04230
[imported] - Arabidopsis thaliana, partial (4%)
Length = 755
Score = 83.6 bits (205), Expect = 2e-16
Identities = 72/197 (36%), Positives = 93/197 (46%), Gaps = 2/197 (1%)
Frame = +2
Query: 2 ETRSAKRKKMAQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLW 61
+ RS + KK + N + DRISDLPD +L HILS K A QT+ILS RW LW
Sbjct: 89 KNRSFQMKKRRKNNHNHTENK--DRISDLPDCILFHILSS*ETKHAVQTTILSTRWNNLW 262
Query: 62 TTFPDLDFRTIDPHAFSSRNLQSSDFETSGHPLASTMDFITQVLSTRDKDSDIRALCFRA 121
P L L SS F+T L S F+++VLS RD + + L F+
Sbjct: 263 KRLPTL-------------RLNSSHFKT----LKSFTKFVSRVLSLRDDSTALHTLYFQR 391
Query: 122 R--LSFSRLNSLIRSAVRHNVKELDIEVTPEVRTDEYFNFPRCLIASKTLRVLKLRSGFR 179
+ + L+ L+ A HNV+ LDI VR D + FP TL L L S
Sbjct: 392 KGCVEPPILSKLVEYAASHNVQRLDI----FVRCDGVY-FPPYFFKCNTLTSLNLSSS-- 550
Query: 180 LPPSSIMKETFQSLHTL 196
P + K+ F SL TL
Sbjct: 551 -PIPNTAKKPFPSLMTL 598
>TC82256 similar to GP|20042913|gb|AAM08741.1 Unknown protein {Oryza sativa
(japonica cultivar-group)}, partial (9%)
Length = 897
Score = 76.6 bits (187), Expect = 2e-14
Identities = 62/178 (34%), Positives = 83/178 (45%), Gaps = 3/178 (1%)
Frame = +3
Query: 25 DRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSSRNLQS 84
D ISDL D+VL HILS L A QT +LSKRW LW + + R+
Sbjct: 219 DFISDLTDSVLLHILSFLNAIQAVQTCVLSKRWIILWKSLSTITLRS------------- 359
Query: 85 SDFETSGHPLASTMDFITQVLSTRDKDSDIRALCF--RARLSFSRLNSLIRSAVRHNVKE 142
+ P +F++++ S RD + I L R + S L +I AV HNV+
Sbjct: 360 ----SYSRPRKRFDEFVSRIFSLRDGSTAIHTLDLYRRHSMKHSLLRKIIEYAVSHNVQH 527
Query: 143 LDIEVTPEVRTDEYFNFPRCLIASKTLRVLKLRSGFRLPPSSIMKETFQ-SLHTLSLT 199
L I+ T + NFP CL + TL+ L L SGF K F+ SL+ SLT
Sbjct: 528 LRIDYTCHIE-----NFPSCLFSCHTLKSLNL-SGFLYNTFVHHKPVFRNSLNLPSLT 683
>TC90676 similar to GP|6503308|gb|AAF14684.1| Contains PF|00646 F-box
domain. ESTs gb|AA586135 gb|Z26675 gb|AI993239 and
gb|AA585907 come from, partial (5%)
Length = 1119
Score = 75.5 bits (184), Expect = 5e-14
Identities = 59/184 (32%), Positives = 92/184 (49%), Gaps = 4/184 (2%)
Frame = +3
Query: 25 DRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSSRNLQS 84
D ISDLP+ +L++ILS L + A +TSIL+ +W++LWT P DFR P RN +S
Sbjct: 204 DMISDLPEGILYYILSRLSTEEAVRTSILATKWRYLWTQLPVFDFRDSSP----KRNSKS 371
Query: 85 SDFETSGHPLASTMDFITQVLSTRDKDSDIRALCFRARLSF---SRLNSLIRSAVRHNVK 141
D +D + Q+L K + I+ L ++ +++S++ SA+ H+V
Sbjct: 372 PD---------CLVDLVDQLL---HKSNRIKKLFIELPVTIVDAGKVSSMLSSALMHDVH 515
Query: 142 ELDIEVTPEVRTDEYFNFPRCLIASKTLRVLKLRSGF-RLPPSSIMKETFQSLHTLSLTL 200
+L + + E + F P AS++L L + GF P I F SL LS
Sbjct: 516 DLTLNLESE---NSRFVLPNSFSASRSLTKLDIVFGFLDSVPDGI---CFPSLKILSFAH 677
Query: 201 GPVD 204
G +D
Sbjct: 678 GGLD 689
>TC78592 similar to GP|9802755|gb|AAF99824.1| Unknown protein {Arabidopsis
thaliana}, partial (8%)
Length = 1909
Score = 72.4 bits (176), Expect = 4e-13
Identities = 58/178 (32%), Positives = 89/178 (49%), Gaps = 3/178 (1%)
Frame = +2
Query: 24 TDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSSRNLQ 83
+DRIS LPD ++ H LS L IK A +TS+LS +W+ W+T PDL F D S+
Sbjct: 458 SDRISCLPDHIIDHFLSRLSIKEAGRTSVLSNKWRKKWSTQPDLVF---DKQCVSA--AV 622
Query: 84 SSDFETSGHPLASTMDFITQVLS---TRDKDSDIRALCFRARLSFSRLNSLIRSAVRHNV 140
S D + T+D + + S + K SD F +S + ++ I R ++
Sbjct: 623 SEDPSVIENKFVRTVDHVLLLHSGPINKFKISDF-GCDFIGMISVADVDRWILHLTRRSI 799
Query: 141 KELDIEVTPEVRTDEYFNFPRCLIASKTLRVLKLRSGFRLPPSSIMKETFQSLHTLSL 198
K L++++ E R +N P CL + ++L L L PP+ M E F +L +L L
Sbjct: 800 KNLELDIWFEQR----YNIPWCLFSCQSLHHLNLFCSSLKPPA--MFEGFMNLKSLQL 955
Score = 35.4 bits (80), Expect = 0.054
Identities = 71/312 (22%), Positives = 114/312 (35%), Gaps = 40/312 (12%)
Frame = +2
Query: 221 LRNLHLQLCFGLRVLRVGCLALRDLSLEKCINLHGLDVVTPKLERLRVSECFHGYHNKSW 280
++NL L + F R CL C +LH L++ L + F G+ N
Sbjct: 797 IKNLELDIWFEQRYNIPWCLF-------SCQSLHHLNLFCSSL---KPPAMFEGFMNLKS 946
Query: 281 VRIN---------------APKLESLFWQYNAVADVTVFERSNVLHEASVDFVLVMSKND 325
+++ +P LE L + V F + N+ H S+ +V + +
Sbjct: 947 LQLEQVTVAKDAFENLISGSPLLEKL-----VLKKVDGFTQINI-HSPSIKYVDIRGNFE 1108
Query: 326 KVSMDKLQSAINFMAGLSCARSLTLDSKTIELQKRITSMCSSPWLQM---LSNSNFYIEP 382
+S D LS L+S+T + + CSS L+ L N +
Sbjct: 1109 GISFDNTSQLAKVFVNLSSY----LNSET---GQSMFHGCSSNLLKFFDHLHNIQSLLIA 1267
Query: 383 LYNLKSLE-----------------LHTCFN---KTNIQGLACLFRSSPTLHTLIVKIID 422
Y++K L L C N T I + CL +SSP L T
Sbjct: 1268 SYSIKYLAAGVVPVKLPTLCINLTFLSICLNFNDLTEISAVLCLLKSSPNLQTF------ 1429
Query: 423 HNKFDRKQWNKDLDMTSNEEEQYWES--EIPTLKPFLQHLKVVEIHGFLDCENEVTLAKF 480
F R + DL WE P + LQH+++ +I G ++E+ +F
Sbjct: 1430 -KMFARLKEETDL---LTHTSYCWEDIFSEPDMLLKLQHVEIQDISG---TKSELDFIRF 1588
Query: 481 LLKNGKALEEMV 492
LL L++M+
Sbjct: 1589 LLLYSPVLKKMI 1624
>TC82690 weakly similar to GP|9802755|gb|AAF99824.1| Unknown protein
{Arabidopsis thaliana}, partial (12%)
Length = 762
Score = 71.6 bits (174), Expect = 7e-13
Identities = 65/218 (29%), Positives = 96/218 (43%), Gaps = 2/218 (0%)
Frame = +1
Query: 22 AATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSSRN 81
A DRIS LP ++ ILS+LPIK A +TSILS +W++ W T P+L F D S
Sbjct: 106 AEPDRISSLPGHIIDQILSILPIKEAVRTSILSTKWRYKWATLPNLVF---DSQCISD-- 270
Query: 82 LQSSDFETSGHPLASTMDFITQVLSTRDKDSDIRALCFRARLSFSRLNSLIRSAVRHNVK 141
S D L+ +D + + S K L R + + ++ R VK
Sbjct: 271 -TSEDLLVIKSKLSRIIDHVLLLHSGPIKKF---KLSHRELIGVTDIDRWTLHLTRRPVK 438
Query: 142 ELDIEVTPEVRTDEYFNFPRCLIASKTLRVLKLRSGFRLPPSSIMKETFQSLHTLSLTLG 201
E +E+ R + P CL + + L L+L + + +PPS+ + F++L +L L
Sbjct: 439 EFVLEIWKGQR----YKIPSCLFSCQGLHHLELFNCWLIPPSTF--QGFRNLKSLDL--- 591
Query: 202 PVDHQAPLSDLFTE--SAFPFLRNLHLQLCFGLRVLRV 237
H D F S P L L L G L +
Sbjct: 592 --QHVTLSQDAFENLISTCPLLERLTLMNFDGFNYLNI 699
>TC81555 weakly similar to GP|20042914|gb|AAM08742.1 Unknown protein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 688
Score = 63.2 bits (152), Expect = 2e-10
Identities = 51/170 (30%), Positives = 79/170 (46%), Gaps = 2/170 (1%)
Frame = +2
Query: 7 KRKKMAQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPD 66
K+ K++ ENE DR+SDLP++V+ HILS L K A QT +LS +K LW P
Sbjct: 131 KKVKLSSECENE------DRLSDLPESVILHILSFLNTKHAVQTCVLSPIYKDLWKRLPA 292
Query: 67 LDFRTIDPHAFSSRNLQSSDFETSGHPLASTMDFITQVLSTRDKDSDIRALCFRARLS-- 124
L D F F T F++++LS RD +++L F+
Sbjct: 293 LTLHNRDFRNFKI-------FTT----------FVSKILSLRDSSISLQSLDFQRHNGRF 421
Query: 125 FSRLNSLIRSAVRHNVKELDIEVTPEVRTDEYFNFPRCLIASKTLRVLKL 174
+L ++ A+ HNV+ L + V ++ P L + + L L+L
Sbjct: 422 EPQLKKIVNYAISHNVQRLQLCVNCDIA-----QIPHSLFSFQPLTHLEL 556
>TC83352 weakly similar to GP|20042917|gb|AAM08745.1 Unknown protein {Oryza
sativa (japonica cultivar-group)}, partial (3%)
Length = 637
Score = 63.2 bits (152), Expect = 2e-10
Identities = 50/149 (33%), Positives = 73/149 (48%), Gaps = 2/149 (1%)
Frame = +3
Query: 5 SAKRKKMAQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTF 64
S KR K+++ ENE DR+SDLP++++ HILS L A QT +LS R+K LW
Sbjct: 174 SPKRIKLSE-SENE------DRLSDLPESIILHILSFLNTNHAVQTCVLSTRYKDLWKRL 332
Query: 65 PDLDFRTIDPHAFSSRNLQSSDFETSGHPLASTMDFITQVLSTRDKDSDIRALCF-RARL 123
P L L SSDF +++VL RD ++AL F R+
Sbjct: 333 PTL-------------TLHSSDFGA----YKKFTRLVSKVLFLRDSSIALQALDFKRSNG 461
Query: 124 SFS-RLNSLIRSAVRHNVKELDIEVTPEV 151
F +L ++ A+ HNVK L + ++
Sbjct: 462 RFEPKLEKIVNYALSHNVKRLGLHFNGDI 548
>BG450859 weakly similar to GP|21553400|gb| unknown {Arabidopsis thaliana},
partial (11%)
Length = 549
Score = 62.4 bits (150), Expect = 4e-10
Identities = 56/175 (32%), Positives = 76/175 (43%), Gaps = 2/175 (1%)
Frame = +2
Query: 4 RSAKRKKMAQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTT 63
R+ +R +M+ E A DRI LPD +L HILS +P K A TSILSKRW LW
Sbjct: 68 RTPRRNRMSS---PENSTATVDRIGTLPDEILIHILSFVPTKQAFTTSILSKRWIHLWRY 238
Query: 64 FPDLDFRTIDPHAFSSRNLQSSDFETSGHPLASTMDFITQVLSTRDK--DSDIRALCFRA 121
P LD F+ NL+ D + +FI V+ +R + I
Sbjct: 239 VPILD--------FTETNLEDRD------SVIRFEEFIFSVIRSRHSAGNHSINTFILDI 376
Query: 122 RLSFSRLNSLIRSAVRHNVKELDIEVTPEVRTDEYFNFPRCLIASKTLRVLKLRS 176
+ SRL++L+ + PE FP ++ S TL LKL S
Sbjct: 377 QRHSSRLHALVFGNSN--------TIAPE--------FPIPILGSTTLVXLKLAS 493
>BG449234 weakly similar to GP|8979942|gb|A Contains a F-box PF|00646 domain.
{Arabidopsis thaliana}, partial (6%)
Length = 655
Score = 61.2 bits (147), Expect = 9e-10
Identities = 54/162 (33%), Positives = 69/162 (42%), Gaps = 2/162 (1%)
Frame = +3
Query: 25 DRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSSRNLQS 84
DRIS L D +L HILS L + A QT ILSKRW LW T P L T+D + FS +
Sbjct: 156 DRISVLADCLLIHILSFLNTREAVQTCILSKRWINLWKTLPTL---TLDCYQFSDCEIYE 326
Query: 85 SDFETSGHPLASTMDFITQVLSTRDKDSDIRALCFRAR--LSFSRLNSLIRSAVRHNVKE 142
+F+ LS RD + + AL + S +I A HNV+
Sbjct: 327 --------------NFLFMFLSLRDHSTALSALLLHNNHFENISLYQMVIEYAFTHNVQH 464
Query: 143 LDIEVTPEVRTDEYFNFPRCLIASKTLRVLKLRSGFRLPPSS 184
I T +F ++S TL L L L P S
Sbjct: 465 FKINYTTAELFSPFF------LSSHTLTSLTLTGEDLLLPRS 572
>AJ499721 weakly similar to PIR|T48291|T482 hypothetical protein F9G14.10 -
Arabidopsis thaliana, partial (8%)
Length = 502
Score = 60.8 bits (146), Expect = 1e-09
Identities = 26/68 (38%), Positives = 42/68 (61%)
Frame = +1
Query: 4 RSAKRKKMAQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTT 63
RS K ++Q + N DR+ D+PD ++HHILS + + A +T +LSKRW+++W +
Sbjct: 235 RSKKTTLISQHVYNCVATGHQDRLGDMPDCLIHHILSFMETRDAIRTCVLSKRWRYIWKS 414
Query: 64 FPDLDFRT 71
P L F +
Sbjct: 415 IPCLYFNS 438
>CA990540 weakly similar to GP|10092259|gb| hypothetical protein; 40655-42260
{Arabidopsis thaliana}, partial (9%)
Length = 761
Score = 60.5 bits (145), Expect = 2e-09
Identities = 62/240 (25%), Positives = 106/240 (43%), Gaps = 3/240 (1%)
Frame = +1
Query: 23 ATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDF-RTIDPHAFSSRN 81
A +RIS+LPD +L HI+S L + A +T ILSKRWK L L + + D ++F
Sbjct: 91 AGNRISELPDCILLHIMSFLEARDAVRTCILSKRWKDLCKRLTTLTYIPSWDENSFK--- 261
Query: 82 LQSSDFETSGHPLASTMDFITQVLSTRDKDSDIRALCFRARLSFSR--LNSLIRSAVRHN 139
+F + VLS+RD+ + L + L++LI+ A+ HN
Sbjct: 262 -----------------NFKSWVLSSRDQSCSLWNLTIDTQFQEGEEDLHTLIQYALFHN 390
Query: 140 VKELDIEVTPEVRTDEYFNFPRCLIASKTLRVLKLRSGFRLPPSSIMKETFQSLHTLSLT 199
++ L+I++ P + + ++AS +L L+L +++ +++
Sbjct: 391 LQNLNIKINPSLTPKS--DLLPLILASNSLTFLEL--------------SYRRGGSIAAA 522
Query: 200 LGPVDHQAPLSDLFTESAFPFLRNLHLQLCFGLRVLRVGCLALRDLSLEKCINLHGLDVV 259
P+ L P LR LHL+ V +A RD ++ NLH L+ +
Sbjct: 523 RSPI--------LPKSLHLPALRTLHLEY--------VNFVATRDHYVDPFSNLHALNTL 654
>BE324624 weakly similar to GP|20042914|gb| Unknown protein {Oryza sativa
(japonica cultivar-group)}, partial (7%)
Length = 671
Score = 60.5 bits (145), Expect = 2e-09
Identities = 54/179 (30%), Positives = 80/179 (44%), Gaps = 3/179 (1%)
Frame = +1
Query: 7 KRKKMAQI-MENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFP 65
K KK+ Q ENE D++SDLP+ V+ HILS L K A QT +LS RWK LW P
Sbjct: 148 KMKKIRQCETENEENE---DKLSDLPECVILHILSFLDSKHAVQTCVLSTRWKHLWARIP 318
Query: 66 DLDFRTIDPHAFSSRNLQSSDFETSGHPLASTMDFITQVLSTRDKDSDIRALCF--RARL 123
L L SS+F T + F++ +L+ R+ + ++AL R +
Sbjct: 319 TL-------------ILHSSNFST----VKKFAIFVSNILTLRNSSTSLQALNLDRRGDI 447
Query: 124 SFSRLNSLIRSAVRHNVKELDIEVTPEVRTDEYFNFPRCLIASKTLRVLKLRSGFRLPP 182
L ++ HN ++ ++ + N C+ + L LKL LPP
Sbjct: 448 EPQLLKKILNYICSHNTHLHELGISLRGGSGFILN---CVSSCHALTSLKL----SLPP 603
>BQ124147 similar to GP|10177836|dbj emb|CAB62440.1~gene_id:MCD7.13~similar
to unknown protein {Arabidopsis thaliana}, partial
(11%)
Length = 677
Score = 59.7 bits (143), Expect = 3e-09
Identities = 31/52 (59%), Positives = 34/52 (64%)
Frame = +1
Query: 18 EAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDF 69
EA DRIS LPD +L HILS LP K A TS+LSKRWK LW P L+F
Sbjct: 4 EAMEEEVDRISILPDDILVHILSPLPTKQAFVTSVLSKRWKNLWCLVPALEF 159
>AL381150 weakly similar to GP|14028988|gb| Unknown protein {Oryza sativa},
partial (10%)
Length = 469
Score = 59.7 bits (143), Expect = 3e-09
Identities = 41/128 (32%), Positives = 65/128 (50%), Gaps = 4/128 (3%)
Frame = +1
Query: 24 TDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSSRNLQ 83
+DRIS+LPD VL HI+ + IK + QT +LSKRWK LW + +L L
Sbjct: 97 SDRISELPDHVLLHIIEFMNIKQSVQTCVLSKRWKSLWKSLANL-------------TLH 237
Query: 84 SSDFETSGHPLASTMDFITQVLSTRDKDSDIRALCFR----ARLSFSRLNSLIRSAVRHN 139
S+ + S F++Q+L+ RD + +L + A + + L ++ A HN
Sbjct: 238 HSEKQKS----YICKWFVSQLLARRDNSLPLHSLSYEYDNAADCNKTTLLEVMEYAASHN 405
Query: 140 VKELDIEV 147
V++L + V
Sbjct: 406 VQQLTVIV 429
>TC82802 similar to GP|9802755|gb|AAF99824.1| Unknown protein {Arabidopsis
thaliana}, partial (50%)
Length = 864
Score = 58.9 bits (141), Expect = 5e-09
Identities = 42/138 (30%), Positives = 70/138 (50%)
Frame = +3
Query: 25 DRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSSRNLQS 84
D ISDLP +++ IL LPI+ A +TSILS++W++ W+T L F D + + S
Sbjct: 132 DLISDLPQSIIETILIQLPIRDAVRTSILSRKWRYKWSTITQLVF---DEKCYPN----S 290
Query: 85 SDFETSGHPLASTMDFITQVLSTRDKDSDIRALCFRARLSFSRLNSLIRSAVRHNVKELD 144
+D E S ++FIT++L + + + +N I R+++K+LD
Sbjct: 291 TDREV---VQKSVVEFITRLLFLHQGPIHKFQIVNTSLQTCPAINQWILFLSRNDLKDLD 461
Query: 145 IEVTPEVRTDEYFNFPRC 162
+ E+ E+F P C
Sbjct: 462 L----ELGEGEFFRMPSC 503
>TC87984 weakly similar to GP|10177368|dbj|BAB10659. contains similarity to
heat shock transcription factor HSF30~gene_id:K16L22.13,
partial (5%)
Length = 696
Score = 58.5 bits (140), Expect = 6e-09
Identities = 39/116 (33%), Positives = 61/116 (51%), Gaps = 1/116 (0%)
Frame = +3
Query: 4 RSAKRKKMAQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTS-ILSKRWKFLWT 62
+ A MA+ E+ + DRI+ LP+++LHHILS LP KT QT+ ++S++W+ LW
Sbjct: 42 KEAMDTPMAEAAESISDDDGIDRINQLPNSLLHHILSFLPTKTCVQTTPLVSRKWRNLWK 221
Query: 63 TFPDLDFRTIDPHAFSSRNLQSSDFETSGHPLASTMDFITQVLSTRDKDSDIRALC 118
LDF S + Q +F+ L ++ F+ VL+ R K +R C
Sbjct: 222 NLEALDF-------CDSSSPQIYEFDNDEQFLIFSV-FVNTVLTLR-KSRVVRKFC 362
>TC82340 similar to GP|10178240|dbj|BAB11672. gene_id:MDJ22.8~unknown
protein {Arabidopsis thaliana}, partial (9%)
Length = 641
Score = 57.8 bits (138), Expect = 1e-08
Identities = 32/76 (42%), Positives = 42/76 (55%)
Frame = +3
Query: 7 KRKKMAQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPD 66
KR+ ++ E+E DR+SDLPD +L +ILS L + A QT ILS RWK LW P
Sbjct: 288 KRRSRSENEESE------DRLSDLPDCILIYILSFLKTENAVQTCILSTRWKHLWKRIPK 449
Query: 67 LDFRTIDPHAFSSRNL 82
L R+ F R +
Sbjct: 450 LLLRSSTLFLFGGREI 497
>TC80459 weakly similar to GP|22831085|dbj|BAC15947. contains EST
AU163066(S15063)~unknown protein {Oryza sativa (japonica
cultivar-group)}, partial (4%)
Length = 1116
Score = 57.8 bits (138), Expect = 1e-08
Identities = 31/63 (49%), Positives = 40/63 (63%)
Frame = +2
Query: 7 KRKKMAQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPD 66
K+KK + E E DR+S+LP +LH+ILS LP K A+TS+LSK W+ W TFP
Sbjct: 47 KKKKKMEHEEEE------DRLSNLPKIILHNILSRLPKKDGARTSVLSKAWEETWYTFPI 208
Query: 67 LDF 69
L F
Sbjct: 209 LYF 217
>BF649840 weakly similar to GP|8979942|gb|A Contains a F-box PF|00646 domain.
{Arabidopsis thaliana}, partial (6%)
Length = 658
Score = 57.0 bits (136), Expect = 2e-08
Identities = 42/114 (36%), Positives = 54/114 (46%)
Frame = +3
Query: 4 RSAKRKKMAQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTT 63
R + R + + ENE D IS+L D +L H+LS L K AAQT ILSKRW LW
Sbjct: 336 RRSLRSRKHKHNENE------DIISNLTDCILIHVLSFLNAKEAAQTCILSKRWITLWKG 497
Query: 64 FPDLDFRTIDPHAFSSRNLQSSDFETSGHPLASTMDFITQVLSTRDKDSDIRAL 117
P + L SS+F T S +F++ LS RD + I L
Sbjct: 498 LPTI-------------TLSSSNFRTE----KSLAEFLSHFLSLRDGSTTIHTL 608
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.134 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,657,684
Number of Sequences: 36976
Number of extensions: 261779
Number of successful extensions: 1544
Number of sequences better than 10.0: 106
Number of HSP's better than 10.0 without gapping: 1510
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1530
length of query: 529
length of database: 9,014,727
effective HSP length: 101
effective length of query: 428
effective length of database: 5,280,151
effective search space: 2259904628
effective search space used: 2259904628
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0203.9


