

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.8
(124 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
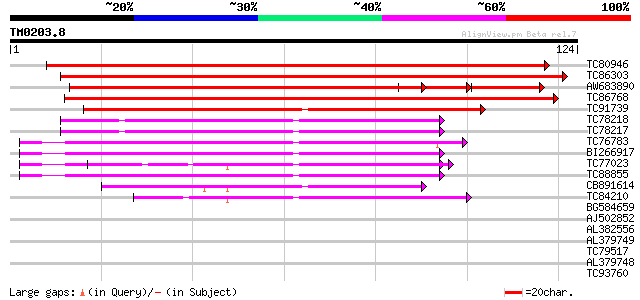
Score E
Sequences producing significant alignments: (bits) Value
TC80946 weakly similar to GP|20804873|dbj|BAB92555. contains EST... 191 5e-50
TC86303 similar to GP|16648777|gb|AAL25579.1 AT4g27280/M4I22_90 ... 157 8e-40
AW683890 similar to GP|16648777|gb| AT4g27280/M4I22_90 {Arabidop... 135 4e-39
TC86768 similar to GP|15294244|gb|AAK95299.1 At2g46600/F13A10.13... 127 8e-31
TC91739 similar to GP|21536724|gb|AAM61056.1 EF-hand Ca2+-bindin... 112 4e-26
TC78218 weakly similar to GP|12963415|gb|AAK11255.1 regulator of... 47 2e-06
TC78217 weakly similar to GP|20161870|dbj|BAB90783. hypothetical... 45 6e-06
TC76783 calmodulin 1 [Medicago truncatula] 45 7e-06
BI266917 PIR|S01173|MC calmodulin [validated] - fruit fly (Droso... 45 9e-06
TC77023 calmodulin 2 [Medicago truncatula] 44 1e-05
TC88855 PIR|A30900|MCBH calmodulin - barley, complete 44 1e-05
CB891614 similar to GP|13561063|em protein kinase {Medicago sati... 39 5e-04
TC84210 homologue to PIR|T07122|T07122 calmodulin CAM5 - soybean... 39 7e-04
BG584659 weakly similar to GP|12963415|gb| regulator of gene sil... 38 0.001
AJ502852 homologue to GP|15010602|gb| AT3g03000/F13E7_5 {Arabido... 37 0.003
AL382556 homologue to GP|169306|gb|AA calmodulin {Phytophthora i... 36 0.004
AL379749 homologue to PIR|S10995|MCUT calmodulin - Trypanosoma c... 34 0.017
TC79517 similar to PIR|T08874|T08874 calcium-dependent protein k... 34 0.017
AL379748 EGAD|125673|13 calmodulin {Sus scrofa}, partial (55%) 33 0.022
TC93760 similar to PIR|T48153|T48153 hypothetical protein T10O8.... 32 0.048
>TC80946 weakly similar to GP|20804873|dbj|BAB92555. contains ESTs
C74086(E30358) AU091313(E30358)~unknown protein, partial
(85%)
Length = 723
Score = 191 bits (486), Expect = 5e-50
Identities = 90/110 (81%), Positives = 105/110 (94%)
Frame = +1
Query: 9 GKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKE 68
GKG++FEDLLPV+ANKLGGEGL+KELCNGF +LMDK++GVITLESL+ N+A++GLQD+KE
Sbjct: 67 GKGIDFEDLLPVIANKLGGEGLMKELCNGFKLLMDKEKGVITLESLRKNSALMGLQDLKE 246
Query: 69 DELVSMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHEL 118
DELVSM+REGDLDGDGALT+MEFCVLMFRLSP+LMEESW LEEAL HEL
Sbjct: 247 DELVSMMREGDLDGDGALTEMEFCVLMFRLSPQLMEESWFLLEEALDHEL 396
>TC86303 similar to GP|16648777|gb|AAL25579.1 AT4g27280/M4I22_90
{Arabidopsis thaliana}, partial (73%)
Length = 865
Score = 157 bits (398), Expect = 8e-40
Identities = 75/111 (67%), Positives = 94/111 (84%)
Frame = +2
Query: 12 LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDEL 71
++F+D LP+MANKLGG+GLI ELCNGF++LMD +GVIT ESLK N+A+LGLQD+ + EL
Sbjct: 107 VQFQDSLPLMANKLGGDGLIDELCNGFNLLMDSTKGVITFESLKKNSALLGLQDLTDVEL 286
Query: 72 VSMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDNNY 122
SMI EGD DGDGAL QMEFCVLMFRLSPELM+ S +WLE+ LQ E+ +++
Sbjct: 287 QSMIVEGDFDGDGALNQMEFCVLMFRLSPELMDGSKMWLEQMLQQEVQDSF 439
>AW683890 similar to GP|16648777|gb| AT4g27280/M4I22_90 {Arabidopsis
thaliana}, partial (35%)
Length = 452
Score = 135 bits (339), Expect(3) = 4e-39
Identities = 65/78 (83%), Positives = 73/78 (93%)
Frame = +3
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVS 73
FEDLLPVMANKLGGEGLIKELCNGF++LMDK++GVITL+SL+ NAAVLGLQD+KEDELV
Sbjct: 45 FEDLLPVMANKLGGEGLIKELCNGFELLMDKEKGVITLDSLRQNAAVLGLQDLKEDELVG 224
Query: 74 MIREGDLDGDGALTQMEF 91
M+ EGDL DGALTQMEF
Sbjct: 225 MMNEGDLXXDGALTQMEF 278
Score = 35.8 bits (81), Expect(3) = 4e-39
Identities = 14/16 (87%), Positives = 14/16 (87%)
Frame = +2
Query: 102 LMEESWIWLEEALQHE 117
LMEESW WLEE LQHE
Sbjct: 311 LMEESWFWLEEXLQHE 358
Score = 26.2 bits (56), Expect(3) = 4e-39
Identities = 11/16 (68%), Positives = 12/16 (74%)
Frame = +1
Query: 86 LTQMEFCVLMFRLSPE 101
L + FCVLMFR SPE
Sbjct: 262 LLRWSFCVLMFRSSPE 309
>TC86768 similar to GP|15294244|gb|AAK95299.1 At2g46600/F13A10.13
{Arabidopsis thaliana}, partial (85%)
Length = 651
Score = 127 bits (320), Expect = 8e-31
Identities = 56/108 (51%), Positives = 84/108 (76%)
Frame = +2
Query: 13 EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELV 72
EFEDLLP+MA KL E + ELC GF++L D++ G+IT ESL+ N+A+LG++ M +++
Sbjct: 167 EFEDLLPIMAEKLDVETFVSELCGGFNLLADQETGLITSESLRKNSAMLGMEGMSKEDAE 346
Query: 73 SMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDN 120
+M+++GDLDGDG L + EFC+LM RLSP +ME++ WLE+A++ +L N
Sbjct: 347 AMVKQGDLDGDGKLNETEFCILMVRLSPGMMEDAETWLEKAIEEQLRN 490
>TC91739 similar to GP|21536724|gb|AAM61056.1 EF-hand Ca2+-binding protein
CCD1 {Arabidopsis thaliana}, partial (53%)
Length = 665
Score = 112 bits (280), Expect = 4e-26
Identities = 54/88 (61%), Positives = 71/88 (80%)
Frame = +1
Query: 17 LLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIR 76
+ P M +++G EG I ELCNGF +LMD ++G+IT ESLK N +LGL+ ++++ELV M+
Sbjct: 85 IFPSMISRVGAEGFIGELCNGFRLLMDVNKGLITFESLKMNCFMLGLE-VRDEELVYMLM 261
Query: 77 EGDLDGDGALTQMEFCVLMFRLSPELME 104
EGDLDGDGAL QMEFC+LMFRLSP LM+
Sbjct: 262 EGDLDGDGALNQMEFCILMFRLSPCLMD 345
>TC78218 weakly similar to GP|12963415|gb|AAK11255.1 regulator of gene
silencing {Nicotiana tabacum}, partial (58%)
Length = 793
Score = 47.0 bits (110), Expect = 2e-06
Identities = 32/84 (38%), Positives = 45/84 (53%)
Frame = +3
Query: 12 LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDEL 71
L E+L+ +M G E +K+L F+M + G IT +SLK +G + DE
Sbjct: 429 LSLEELIALMEEG-GEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMG-ESKSIDEC 602
Query: 72 VSMIREGDLDGDGALTQMEFCVLM 95
+MI+ DLDGDG LT EF +M
Sbjct: 603 KAMIKHFDLDGDGLLTFDEFITMM 674
>TC78217 weakly similar to GP|20161870|dbj|BAB90783. hypothetical protein
{Oryza sativa (japonica cultivar-group)}, partial (49%)
Length = 1027
Score = 45.4 bits (106), Expect = 6e-06
Identities = 31/84 (36%), Positives = 45/84 (52%)
Frame = +2
Query: 12 LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDEL 71
L E+L+ +M G E +K+L F+M + G IT +SLK +G + DE
Sbjct: 380 LSLEELIALMEEG-GEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMG-ESKSIDEC 553
Query: 72 VSMIREGDLDGDGALTQMEFCVLM 95
+MI+ DLDGDG L+ EF +M
Sbjct: 554 KAMIKHFDLDGDGLLSFDEFITMM 625
>TC76783 calmodulin 1 [Medicago truncatula]
Length = 957
Score = 45.1 bits (105), Expect = 7e-06
Identities = 32/99 (32%), Positives = 50/99 (50%), Gaps = 1/99 (1%)
Frame = +3
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +EL F + G I+ L+ LG
Sbjct: 297 GNGTI-----DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLG 461
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFC-VLMFRLSP 100
+ + ++E+ MIRE D+DGDG + EF V+M + SP
Sbjct: 462 -EKLTDEEVDEMIREADVDGDGQINYEEFVKVMMAK*SP 575
Score = 36.2 bits (82), Expect = 0.003
Identities = 27/58 (46%), Positives = 31/58 (52%), Gaps = 1/58 (1%)
Frame = +3
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFR 97
L DKD G IT + L T LG Q+ E EL MI E D DG+G + EF LM R
Sbjct: 174 LFDKDGDGCITTKELGTVMRSLG-QNPTEAELQDMINEVDADGNGTIDFPEFLNLMAR 344
>BI266917 PIR|S01173|MC calmodulin [validated] - fruit fly (Drosophila
melanogaster), complete
Length = 576
Score = 44.7 bits (104), Expect = 9e-06
Identities = 28/93 (30%), Positives = 46/93 (49%)
Frame = +2
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +E+ F + G I+ L+ LG
Sbjct: 188 GNGTI-----DFPEFLTMMARKMKDTDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLG 352
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MIRE D+DGDG + EF +M
Sbjct: 353 -EKLTDEEVDEMIREADIDGDGQVNYEEFVTMM 448
>TC77023 calmodulin 2 [Medicago truncatula]
Length = 999
Score = 44.3 bits (103), Expect = 1e-05
Identities = 29/93 (31%), Positives = 46/93 (49%)
Frame = +1
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +EL F + G I+ L+ LG
Sbjct: 262 GNGTI-----DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLG 426
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MIRE D+DGDG + EF +M
Sbjct: 427 -EKLTDEEVDEMIREADVDGDGQINYEEFVKVM 522
Score = 40.8 bits (94), Expect = 1e-04
Identities = 36/81 (44%), Positives = 42/81 (51%), Gaps = 1/81 (1%)
Frame = +1
Query: 18 LPVMANKLGGEGLIKELCNGFDMLMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIR 76
LP MA++L E I E F L DKD G IT + L T LG Q+ E EL MI
Sbjct: 76 LPPMADQLTDEQ-ISEFKEAFS-LFDKDGDGCITTKELGTVMRSLG-QNPTEAELQDMIN 246
Query: 77 EGDLDGDGALTQMEFCVLMFR 97
E D DG+G + EF LM R
Sbjct: 247 EVDADGNGTIDFPEFLNLMAR 309
>TC88855 PIR|A30900|MCBH calmodulin - barley, complete
Length = 889
Score = 44.3 bits (103), Expect = 1e-05
Identities = 29/93 (31%), Positives = 46/93 (49%)
Frame = +1
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +EL F + G I+ L+ LG
Sbjct: 253 GNGTI-----DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLG 417
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MIRE D+DGDG + EF +M
Sbjct: 418 -EKLTDEEVDEMIREADVDGDGQINYEEFVKVM 513
Score = 37.7 bits (86), Expect = 0.001
Identities = 34/80 (42%), Positives = 41/80 (50%), Gaps = 1/80 (1%)
Frame = +1
Query: 19 PVMANKLGGEGLIKELCNGFDMLMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIRE 77
P MA++L + I E F L DKD G IT + L T LG Q+ E EL MI E
Sbjct: 70 PPMADQLTDDQ-IAEFKEAFS-LFDKDGDGCITTKELGTVMRSLG-QNPTEAELQDMINE 240
Query: 78 GDLDGDGALTQMEFCVLMFR 97
D DG+G + EF LM R
Sbjct: 241 VDADGNGTIDFPEFLNLMAR 300
>CB891614 similar to GP|13561063|em protein kinase {Medicago sativa}, partial
(37%)
Length = 637
Score = 38.9 bits (89), Expect = 5e-04
Identities = 26/75 (34%), Positives = 41/75 (54%), Gaps = 4/75 (5%)
Frame = +3
Query: 21 MANKLGGEGLIKELCNGFDML---MDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIR 76
+A K+ E L +E G + MD D G IT E LKT A +G + + E E+ ++
Sbjct: 138 LALKVMAENLSEEEIKGLKAMFANMDTDSSGTITYEELKTGLARIGSR-LSEAEVKQLME 314
Query: 77 EGDLDGDGALTQMEF 91
D+DG+G++ +EF
Sbjct: 315 AADVDGNGSIDYLEF 359
>TC84210 homologue to PIR|T07122|T07122 calmodulin CAM5 - soybean, partial
(96%)
Length = 627
Score = 38.5 bits (88), Expect = 7e-04
Identities = 31/75 (41%), Positives = 39/75 (51%), Gaps = 1/75 (1%)
Frame = +1
Query: 28 EGLIKELCNGFDMLMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGAL 86
E I E+ F L DKD G IT+E L T L Q+ E+EL MI E D DG+G +
Sbjct: 1 EDQIVEIKEAF-CLFDKDGDGCITVEELATVIRSLD-QNPTEEELQEMINEVDADGNGTI 174
Query: 87 TQMEFCVLMFRLSPE 101
+EF LM + E
Sbjct: 175 EFVEFLNLMAKKMKE 219
Score = 35.0 bits (79), Expect = 0.007
Identities = 24/80 (30%), Positives = 39/80 (48%)
Frame = +1
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ EF + L +MA K+ ++L F + G I+ L+ LG
Sbjct: 160 GNGTI-----EFVEFLNLMAKKMKETDADEDLKEAFKVFDKDQNGYISASELRHVMINLG 324
Query: 63 LQDMKEDELVSMIREGDLDG 82
+ + ++E+ MI+E DLDG
Sbjct: 325 -EKLTDEEVDQMIKEADLDG 381
>BG584659 weakly similar to GP|12963415|gb| regulator of gene silencing
{Nicotiana tabacum}, partial (34%)
Length = 677
Score = 37.7 bits (86), Expect = 0.001
Identities = 29/73 (39%), Positives = 40/73 (54%), Gaps = 1/73 (1%)
Frame = +3
Query: 12 LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKE-DE 70
L EDL+ +M + G E +K+L F+M D+ G IT +SLK LG D K +E
Sbjct: 333 LSLEDLIALMESG-GEEEKLKDLREAFEMYDDEGCGYITPKSLKRMLKKLG--DSKSIEE 503
Query: 71 LVSMIREGDLDGD 83
MI+ DLDG+
Sbjct: 504 CKVMIKRFDLDGE 542
>AJ502852 homologue to GP|15010602|gb| AT3g03000/F13E7_5 {Arabidopsis
thaliana}, partial (36%)
Length = 642
Score = 36.6 bits (83), Expect = 0.003
Identities = 21/62 (33%), Positives = 32/62 (50%)
Frame = +2
Query: 47 GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLSPELMEES 106
G IT L + A LG + +EL MI+E D+DGDG ++ EF ++ + S
Sbjct: 14 GFITAAELAHSMAKLG-HALTAEELTGMIKEADMDGDGMISFQEFAQ---AITSAAFDNS 181
Query: 107 WI 108
W+
Sbjct: 182 WV 187
>AL382556 homologue to GP|169306|gb|AA calmodulin {Phytophthora infestans},
partial (49%)
Length = 476
Score = 35.8 bits (81), Expect = 0.004
Identities = 21/56 (37%), Positives = 32/56 (56%), Gaps = 1/56 (1%)
Frame = +2
Query: 41 LMDKD-RGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ DKD G I+ L+ LG + + ++E+ MIRE D+DGDG + EF +M
Sbjct: 50 VFDKDGNGFISAAELRHVMTNLG-EKLTDEEVDEMIREADVDGDGQINYEEFVKMM 214
>AL379749 homologue to PIR|S10995|MCUT calmodulin - Trypanosoma cruzi,
partial (34%)
Length = 550
Score = 33.9 bits (76), Expect = 0.017
Identities = 18/49 (36%), Positives = 28/49 (56%)
Frame = +3
Query: 47 GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
G I+ L+ LG + + ++E+ MIRE D+DGDG + EF +M
Sbjct: 6 GFISAAELRHVMTNLG-EKLTDEEVDEMIREADIDGDGQINYEEFVKMM 149
>TC79517 similar to PIR|T08874|T08874 calcium-dependent protein kinase (EC
2.7.1.-) gamma - soybean, partial (80%)
Length = 1930
Score = 33.9 bits (76), Expect = 0.017
Identities = 25/78 (32%), Positives = 39/78 (49%), Gaps = 4/78 (5%)
Frame = +1
Query: 18 LPVMANKLGGEGLIKELCNGFDML---MDKDR-GVITLESLKTNAAVLGLQDMKEDELVS 73
L +A K+ E L E G + MD D+ G IT E L+ LG + + E E+
Sbjct: 1492 LKKLALKVIAENLSSEEIQGLKAMFTNMDTDKSGTITYEELRAGLQRLGSK-LTEAEVRQ 1668
Query: 74 MIREGDLDGDGALTQMEF 91
++ D+DG+G + +EF
Sbjct: 1669 LMEAADVDGNGTIDYIEF 1722
>AL379748 EGAD|125673|13 calmodulin {Sus scrofa}, partial (55%)
Length = 262
Score = 33.5 bits (75), Expect = 0.022
Identities = 24/52 (46%), Positives = 28/52 (53%), Gaps = 1/52 (1%)
Frame = +1
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
L DKD G IT + L T LG Q+ E EL MI E D DG+G + EF
Sbjct: 106 LFDKDGDGTITTKELGTVMRSLG-QNPTEAELQDMINEVDADGNGTIDFPEF 258
>TC93760 similar to PIR|T48153|T48153 hypothetical protein T10O8.20 -
Arabidopsis thaliana, partial (18%)
Length = 835
Score = 32.3 bits (72), Expect = 0.048
Identities = 16/38 (42%), Positives = 21/38 (55%)
Frame = +1
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDM 40
GT T +G G E+ LPV K G+ K+ C+GF M
Sbjct: 115 GTKTCSGVGTEWRSRLPVRCPK*APFGIEKDACSGFKM 228
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.138 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,205,754
Number of Sequences: 36976
Number of extensions: 30693
Number of successful extensions: 218
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 205
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 217
length of query: 124
length of database: 9,014,727
effective HSP length: 84
effective length of query: 40
effective length of database: 5,908,743
effective search space: 236349720
effective search space used: 236349720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0203.8


