

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
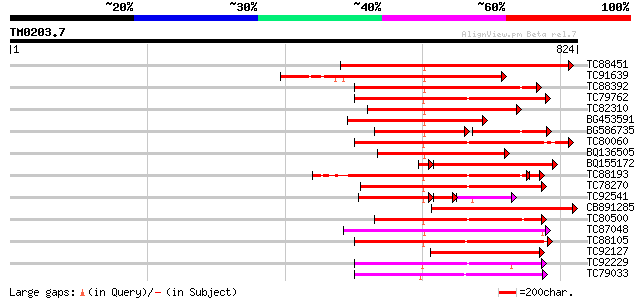
Query= TM0203.7
(824 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC88451 similar to GP|836954|gb|AAC23542.1|| receptor protein ki... 366 e-101
TC91639 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1... 327 9e-90
TC88392 similar to GP|11275527|dbj|BAB18292. putative receptor-l... 312 4e-85
TC79762 similar to PIR|H96731|H96731 hypothetical protein F5A18.... 306 2e-83
TC82310 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1... 293 2e-79
BG453591 similar to PIR|T05181|T05 S-receptor kinase (EC 2.7.1.-... 283 2e-76
BG586735 similar to GP|11275527|db putative receptor-like protei... 185 3e-75
TC80060 similar to PIR|D96574|D96574 hypothetical protein T3F20.... 278 6e-75
BQ136505 similar to PIR|T05754|T05 S-receptor kinase (EC 2.7.1.-... 270 2e-72
BQ155172 weakly similar to GP|6686398|gb F1E22.15 {Arabidopsis t... 248 2e-69
TC88193 similar to PIR|G96602|G96602 probable receptor protein k... 253 3e-69
TC78270 similar to GP|16649103|gb|AAL24403.1 Unknown protein {Ar... 247 1e-65
TC92541 similar to PIR|T02153|T02153 protein kinase homolog T1F1... 137 1e-65
CB891285 weakly similar to PIR|T14470|T14 receptor-like kinase (... 242 4e-64
TC80500 similar to PIR|G96602|G96602 probable receptor protein k... 239 3e-63
TC87048 similar to GP|8778594|gb|AAF79602.1| F5M15.3 {Arabidopsi... 226 3e-59
TC88105 similar to GP|21593085|gb|AAM65034.1 Putative protein ki... 220 2e-57
TC92127 weakly similar to GP|16040952|dbj|BAB69683. receptor kin... 218 7e-57
TC92229 similar to PIR|B86369|B86369 hypothetical protein AAC980... 218 1e-56
TC79033 similar to GP|21554229|gb|AAM63304.1 somatic embryogenes... 212 5e-55
>TC88451 similar to GP|836954|gb|AAC23542.1|| receptor protein kinase
{Ipomoea trifida}, partial (28%)
Length = 1276
Score = 366 bits (940), Expect = e-101
Identities = 187/348 (53%), Positives = 251/348 (71%), Gaps = 10/348 (2%)
Frame = +3
Query: 482 DLIESGRLQ--EDDAKAIDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEI 539
DL+ES ++ EDD K DI F+ SIL+AT F+ NKLGQGG+GPVYKG GQEI
Sbjct: 126 DLVESYDIKDLEDDFKGHDIKVFNFTSILEATMEFSPENKLGQGGYGPVYKGILATGQEI 305
Query: 540 AVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFI 599
AVKRLS SGQG+ EFKNE++LI LQH+NLV+LLG C+ +E++L+YEYMPN+SLD ++
Sbjct: 306 AVKRLSKTSGQGIVEFKNELLLICELQHKNLVQLLGCCIHEEERILIYEYMPNKSLDFYL 485
Query: 600 FD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFG 651
FD W RF II GIA+GLLYLH+ SRL+IIHRDLKASNILLDE NPKI+DFG
Sbjct: 486 FDCTKKKLLDWKKRFNIIEGIAQGLLYLHKYSRLKIIHRDLKASNILLDENMNPKIADFG 665
Query: 652 LARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFY 711
+AR+F +++V NT+R+VGTYGYMSPEYA++G S KSDV+SFGV++LEI+ G +N FY
Sbjct: 666 MARMFTQQESVVNTNRIVGTYGYMSPEYAMEGVCSTKSDVYSFGVLLLEIVCGIKNNSFY 845
Query: 712 QPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMS 771
+ L+L+G+AW LW G L +D TL+ T +E +C++VGLLC+++ +RPTMS
Sbjct: 846 DVDRPLNLIGHAWELWNDGEYLKLMDPTLNDTFVPDEVKRCIHVGLLCVEQYANDRPTMS 1025
Query: 772 NVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSKMETFSRNDLTVT 819
V+ +L ++ PR+PAF +RR +TS +T + + T++
Sbjct: 1026EVISVLTNKYVLTNLPRKPAFYVRREIFEGETTSKGQDTDTYSTTTIS 1169
>TC91639 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1.-)
T6K22.120 precursor - Arabidopsis thaliana, partial
(28%)
Length = 1313
Score = 327 bits (839), Expect = 9e-90
Identities = 185/350 (52%), Positives = 238/350 (67%), Gaps = 21/350 (6%)
Frame = +3
Query: 394 CWIWSQDLNNLEEEYEG-GCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLII 452
C +W DL ++ G G LH+R+A+SD+ GD + E+ + IIL + ++ + I
Sbjct: 27 CMVWYGDLVDILHFQHGEGNALHIRLAYSDL-GDGGKNEKIMM--VIILTSLAGLICIGI 197
Query: 453 LSSTCIYLRRRRQAKIRGIK---------LYGSERYVRDL---IESGRLQEDDAKAIDIP 500
+ + R +RQ K K + S ++ +E G L+ + +++P
Sbjct: 198 I--VLLVWRYKRQLKASCSKNSDVLPVFDAHKSREMSAEIPGSVELG-LEGNQLSKVELP 368
Query: 501 HFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVV 560
F+ + ATNNF+ NKLGQGGFGPVYKGK P G+EIAVKRLS SGQGL+EFKNE+
Sbjct: 369 FFNFSCMSSATNNFSEENKLGQGGFGPVYKGKLPSGEEIAVKRLSRRSGQGLDEFKNEMR 548
Query: 561 LIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGI 612
L A+LQHRNLV+L+G +EGDEK+LVYE+M N+SLD F+FD W R++II GI
Sbjct: 549 LFAQLQHRNLVKLMGCSIEGDEKLLVYEFMLNKSLDRFLFDPIKKTQLDWARRYEIIEGI 728
Query: 613 ARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTY 672
ARGLLYLH DSRLRIIHRDLKASNILLDE NPKISDFGLARIFGG N +VVGTY
Sbjct: 729 ARGLLYLHRDSRLRIIHRDLKASNILLDENMNPKISDFGLARIFGGNQNEENATKVVGTY 908
Query: 673 GYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGY 722
GYMSPEYA++G SVKSDV+SFGV++LEI+SG+RNT F + + SL+GY
Sbjct: 909 GYMSPEYAMEGLVSVKSDVYSFGVLLLEIVSGRRNTSFRHSD-DSSLIGY 1055
>TC88392 similar to GP|11275527|dbj|BAB18292. putative receptor-like protein
kinase {Oryza sativa (japonica cultivar-group)}, partial
(61%)
Length = 1017
Score = 312 bits (799), Expect = 4e-85
Identities = 156/278 (56%), Positives = 204/278 (73%), Gaps = 6/278 (2%)
Frame = +1
Query: 502 FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVL 561
F L ++ ATN F+ N+LG+GGFGPV+KG P G+E+A+K+LS S QG+ EF NEV L
Sbjct: 181 FELNTLQLATNFFSELNQLGRGGFGPVFKGLMPNGEEVAIKKLSMESRQGIREFTNEVRL 360
Query: 562 IARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD------WDMRFKIILGIARG 615
+ R+QH+NLV LLG C EG EKMLVYEY+PN+SLD F+FD W RF+I+ GIARG
Sbjct: 361 LLRIQHKNLVTLLGCCAEGPEKMLVYEYLPNKSLDHFLFDKKRSLDWMTRFRIVTGIARG 540
Query: 616 LLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYM 675
LLYLHE++ RIIHRD+KASNILLDE+ NPKISDFGLAR+F G+DT T R+ T+GYM
Sbjct: 541 LLYLHEEAPERIIHRDIKASNILLDEKLNPKISDFGLARLFPGEDTHVQTFRISRTHGYM 720
Query: 676 SPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDF 735
+PEYAL G+ SVK+DVFS+GV+VLEI+SG++N + LL YAW L++ + +D
Sbjct: 721 APEYALRGYLSVKTDVFSYGVLVLEIVSGRKNHDLKLDAEKADLLSYAWKLYQGRKIMDL 900
Query: 736 IDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNV 773
IDQ + + +E C+ +GLLC Q +ERP M++V
Sbjct: 901 IDQNIGK-YNGDEAAMCIQLGLLCCQASLVERPDMNSV 1011
>TC79762 similar to PIR|H96731|H96731 hypothetical protein F5A18.8
[imported] - Arabidopsis thaliana, partial (55%)
Length = 1564
Score = 306 bits (785), Expect = 2e-83
Identities = 155/293 (52%), Positives = 205/293 (69%), Gaps = 9/293 (3%)
Frame = +3
Query: 502 FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVL 561
F E++L AT NF +KLG+GGFGPVYKGK G+E+AVK+LS S QG +EF NE L
Sbjct: 348 FSYETLLSATKNFNATHKLGEGGFGPVYKGKLSDGREVAVKKLSQTSNQGKKEFMNEAKL 527
Query: 562 IARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIF---------DWDMRFKIILGI 612
+AR+QH+N+V LLGYCV G EK+LVYEY+P+ SLD F+F DW RF II G+
Sbjct: 528 LARVQHKNVVNLLGYCVHGTEKILVYEYVPHESLDKFLFKEAEKREQLDWKRRFGIITGV 707
Query: 613 ARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTY 672
A+GLLYLHEDS IIHRD+KASNILLD++ KI+DFG+AR+F + T RV GT
Sbjct: 708 AKGLLYLHEDSHNCIIHRDIKASNILLDDKWTAKIADFGMARLFPEDQSQVKT-RVAGTN 884
Query: 673 GYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRT 732
GYM+PEY + G SVK+DVFS+GV+VLE+I+G+RN+ F E +LL +A+ ++K GR+
Sbjct: 885 GYMAPEYMMHGRLSVKADVFSYGVLVLELITGQRNSSFNLXVEEHNLLDWAYKMYKKGRS 1064
Query: 733 LDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLP 785
L+ +D L+ T E+ C+ + LLC+Q DP RPTM ++ L +S T P
Sbjct: 1065LEIVDSALASTVLTEQVDMCIQLALLCIQGDPQLRPTMRRIVVKLSRKSPTKP 1223
>TC82310 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1.-)
T6K22.120 precursor - Arabidopsis thaliana, partial
(24%)
Length = 696
Score = 293 bits (749), Expect = 2e-79
Identities = 146/231 (63%), Positives = 178/231 (76%), Gaps = 8/231 (3%)
Frame = +1
Query: 521 GQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEG 580
G+GGFG VYKG +EIAVKRLS S QG+EE KNE++LIA+LQHRNLVRLL C+E
Sbjct: 1 GKGGFGTVYKGVLADEKEIAVKRLSKTSSQGVEELKNEIILIAKLQHRNLVRLLACCIEQ 180
Query: 581 DEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRDL 632
+EK+L+YEY+PN SLD +FD W R II GIA+GLLYLHEDSRLR+IHRDL
Sbjct: 181 NEKLLIYEYLPNSSLDFHLFDMVKGAQLAWRQRLNIINGIAKGLLYLHEDSRLRVIHRDL 360
Query: 633 KASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVF 692
KASNILLD+E NPKISDFGLAR FGG NT RVVGTYGYM+PEYA++G FSVKSDVF
Sbjct: 361 KASNILLDQEMNPKISDFGLARTFGGDQDEANTIRVVGTYGYMAPEYAMEGLFSVKSDVF 540
Query: 693 SFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQT 743
SFGV++LEIISG++N+ FY EH SL +AW+LW + + +D ++ ++
Sbjct: 541 SFGVLLLEIISGRKNSKFYLSEHGQSLPIFAWNLWCKRKGFELMDPSIEKS 693
>BG453591 similar to PIR|T05181|T05 S-receptor kinase (EC 2.7.1.-) T6K22.120
precursor - Arabidopsis thaliana, partial (24%)
Length = 640
Score = 283 bits (723), Expect = 2e-76
Identities = 139/212 (65%), Positives = 170/212 (79%), Gaps = 9/212 (4%)
Frame = +3
Query: 492 DDAKAIDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFP-GGQEIAVKRLSSCSGQ 550
+D + ++P F+L +I+DATN+F+ NKLG+GGFGPVYKG +EIAVKRLS S Q
Sbjct: 3 EDEQDFELPFFNLSTIIDATNDFSNDNKLGEGGFGPVYKGTLVLDRREIAVKRLSGSSKQ 182
Query: 551 GLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------W 602
G EFKNEV+L ++LQHRNLV++LG C++G+EKML+YEYMPNRSLD+F+FD W
Sbjct: 183 GTREFKNEVILCSKLQHRNLVKVLGCCIQGEEKMLIYEYMPNRSLDSFLFDQAQKKLLDW 362
Query: 603 DMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTV 662
RF II GIARGL+YLH+DSRLRIIHRDLK SNILLD + NPKISDFGLA+I G
Sbjct: 363 SKRFNIICGIARGLIYLHQDSRLRIIHRDLKPSNILLDNDMNPKISDFGLAKICGDDQVE 542
Query: 663 GNTDRVVGTYGYMSPEYALDGHFSVKSDVFSF 694
GNT+RVVGT+GYM+PEYA+DG FS+KSDVFSF
Sbjct: 543 GNTNRVVGTHGYMAPEYAIDGLFSIKSDVFSF 638
>BG586735 similar to GP|11275527|db putative receptor-like protein kinase
{Oryza sativa (japonica cultivar-group)}, partial (57%)
Length = 794
Score = 185 bits (469), Expect(2) = 3e-75
Identities = 92/145 (63%), Positives = 113/145 (77%), Gaps = 6/145 (4%)
Frame = +3
Query: 530 KGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEY 589
+G P G+E+A+K+LS S QG+ EF NEV L+ R+QH+NLV LLG C EG EKMLVYEY
Sbjct: 6 RGLMPNGEEVAIKKLSMESRQGIREFTNEVRLLLRIQHKNLVTLLGCCAEGPEKMLVYEY 185
Query: 590 MPNRSLDAFIFD------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEK 643
+PN+SLD F+FD W RF+I+ GIARGLLYLHE++ RIIHRD+KASNILLDE+
Sbjct: 186 LPNKSLDHFLFDKKRSLDWMTRFRIVTGIARGLLYLHEEAPERIIHRDIKASNILLDEKL 365
Query: 644 NPKISDFGLARIFGGKDTVGNTDRV 668
NPKISDFGLAR+F G+DT T R+
Sbjct: 366 NPKISDFGLARLFPGEDTHVQTFRI 440
Score = 116 bits (290), Expect(2) = 3e-75
Identities = 56/115 (48%), Positives = 78/115 (67%)
Frame = +2
Query: 673 GYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRT 732
GYM+PEYAL G+ SVK+DVFS+GV+VLEI+SG++N + LL YAW L++ G+
Sbjct: 446 GYMAPEYALRGYLSVKTDVFSYGVLVLEIVSGRKNHDLKLDAEKADLLSYAWKLYQGGKI 625
Query: 733 LDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSP 787
+D IDQ + + +E C+ +GLLC Q +ERP M++V ML S+S TLP P
Sbjct: 626 MDLIDQNIGK-YNGDEAAMCIQLGLLCCQASLVERPDMNSVNLMLSSDSFTLPKP 787
>TC80060 similar to PIR|D96574|D96574 hypothetical protein T3F20.24
[imported] - Arabidopsis thaliana, partial (32%)
Length = 1547
Score = 278 bits (711), Expect = 6e-75
Identities = 155/329 (47%), Positives = 212/329 (64%), Gaps = 10/329 (3%)
Frame = +1
Query: 501 HFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVV 560
++ L I ATNNF NK+G+GGFGPVYKG G IAVK+LSS S QG EF NE+
Sbjct: 154 YYSLRQIKVATNNFDPKNKIGEGGFGPVYKGVLSDGAVIAVKQLSSKSKQGNREFVNEIG 333
Query: 561 LIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIF---------DWDMRFKIILG 611
+I+ LQH NLV+L G C+EG++ +LVYEYM N SL +F DW R KI +G
Sbjct: 334 MISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGKPEQRLNLDWRTRMKICVG 513
Query: 612 IARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGT 671
IARGL YLHE+SRL+I+HRD+KA+N+LLD+ N KISDFGLA++ ++T +T R+ GT
Sbjct: 514 IARGLAYLHEESRLKIVHRDIKATNVLLDKNLNAKISDFGLAKLDEEENTHIST-RIAGT 690
Query: 672 YGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGR 731
GYM+PEYA+ G+ + K+DV+SFGVV LEI+SG NT + E + LL +A+ L + G
Sbjct: 691 IGYMAPEYAMRGYLTDKADVYSFGVVALEIVSGMSNTNYRPKEEFVYLLDWAYVLQEQGN 870
Query: 732 TLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNT-LPSPREP 790
L+ +D TL +EE M+ + + LLC P RP MS+V+ ML E NT + +P
Sbjct: 871 LLELVDPTLGSKYSSEEAMRMLQLALLCTNPSPTLRPPMSSVVSML--EGNTPIQAP--- 1035
Query: 791 AFVIRRCPSSRASTSSKMETFSRNDLTVT 819
+I+R S+ + E S++ T +
Sbjct: 1036--IIKRSDSTAGARFKAFELLSQDSQTTS 1116
>BQ136505 similar to PIR|T05754|T05 S-receptor kinase (EC 2.7.1.-) M4I22.110
precursor - Arabidopsis thaliana, partial (23%)
Length = 618
Score = 270 bits (689), Expect = 2e-72
Identities = 135/200 (67%), Positives = 156/200 (77%), Gaps = 8/200 (4%)
Frame = +2
Query: 535 GGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRS 594
GG+E+A+KR S S QGLEEFKNEV+LIA+LQHRNLV+LLG C+ +EK+L+YEYMPNRS
Sbjct: 17 GGKEVAIKRNSKMSDQGLEEFKNEVLLIAKLQHRNLVKLLGCCIHREEKLLIYEYMPNRS 196
Query: 595 LDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPK 646
LD FIFD W R II G+ARGLLYLH+DSRLRIIHRDLK SNILLD NPK
Sbjct: 197 LDYFIFDETRSKLLDWSKRSHIIAGVARGLLYLHQDSRLRIIHRDLKLSNILLDALMNPK 376
Query: 647 ISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKR 706
ISDFGLAR F G T ++VGTYGYM PEYA+ G +S+KSDVFSFGV+VLEIISGK+
Sbjct: 377 ISDFGLARTFCGDQVEAKTRKLVGTYGYMPPEYAVHGRYSMKSDVFSFGVIVLEIISGKK 556
Query: 707 NTGFYQPEHELSLLGYAWHL 726
FY P H L+LLG+AW L
Sbjct: 557 IKVFYDPXHSLNLLGHAWRL 616
>BQ155172 weakly similar to GP|6686398|gb F1E22.15 {Arabidopsis thaliana},
partial (10%)
Length = 637
Score = 248 bits (633), Expect(2) = 2e-69
Identities = 118/180 (65%), Positives = 145/180 (80%)
Frame = +3
Query: 616 LLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYM 675
LLYLH+DSRLR+IHRDLK SNILLDEE PKISDFGLARI GGK+T +T+RVVGTYGYM
Sbjct: 78 LLYLHQDSRLRVIHRDLKTSNILLDEEMQPKISDFGLARIVGGKETEASTERVVGTYGYM 257
Query: 676 SPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDF 735
SPEYALDG+FS KSDVFSFGVV+LEIISGK+NTGFY+ + SLLGYAW LW+ + D
Sbjct: 258 SPEYALDGYFSTKSDVFSFGVVLLEIISGKKNTGFYRSKEISSLLGYAWRLWREDKLQDL 437
Query: 736 IDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIR 795
+D TLS T + ++C +GLLC+Q++P +RP MSNV+ ML +E+ TL +P++P F R
Sbjct: 438 MDPTLSDTYNVNQFIRCSQIGLLCVQDEPDDRPHMSNVVTMLDNETTTLLTPKQPTFFTR 617
Score = 33.9 bits (76), Expect(2) = 2e-69
Identities = 15/22 (68%), Positives = 17/22 (77%)
Frame = +2
Query: 594 SLDAFIFDWDMRFKIILGIARG 615
S + I DW MRF+IILGIARG
Sbjct: 11 STKSVILDWPMRFEIILGIARG 76
>TC88193 similar to PIR|G96602|G96602 probable receptor protein kinase
F14G9.24 [imported] - Arabidopsis thaliana, partial
(20%)
Length = 1815
Score = 253 bits (646), Expect(2) = 3e-69
Identities = 147/327 (44%), Positives = 207/327 (62%), Gaps = 8/327 (2%)
Frame = +3
Query: 440 ILVTFTAVVFLIILSSTCIYLRRRRQAKIRGIKLYGSERYVRDLIESGRLQEDDAKAID- 498
++V AV FL + S+ Y+ RRR KLY + DD ID
Sbjct: 417 VVVGVGAVCFLAV--SSIFYILRRR-------KLYNDD--------------DDLVGIDT 527
Query: 499 IPH-FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKN 557
+P+ F + +AT++F NKLG+GGFGPVYKG G+ +AVK+LS S QG +F
Sbjct: 528 MPNTFSYYELKNATSDFNRDNKLGEGGFGPVYKGTLNDGRFVAVKQLSIGSHQGKSQFIA 707
Query: 558 EVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIF------DWDMRFKIILG 611
E+ I+ +QHRNLV+L G C+EG++++LVYEY+ N+SLD +F +W R+ + +G
Sbjct: 708 EIATISAVQHRNLVKLYGCCIEGNKRLLVYEYLENKSLDQALFGNVLFLNWSTRYDVCMG 887
Query: 612 IARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGT 671
+ARGL YLHE+SRLRI+HRD+KASNILLD E PK+SDFGLA+++ K T +T RV GT
Sbjct: 888 VARGLTYLHEESRLRIVHRDVKASNILLDSELVPKLSDFGLAKLYDDKKTHIST-RVAGT 1064
Query: 672 YGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGR 731
GY++PEYA+ G + K+DVFSFGVV LE++SG+ N+ E ++ LL +AW L +
Sbjct: 1065IGYLAPEYAMRGRLTEKADVFSFGVVALELVSGRPNSDSSLEEDKMYLLDWAWQLHERNC 1244
Query: 732 TLDFIDQTLSQTCKAEECMKCVNVGLL 758
D ID LS+ EE + V +G++
Sbjct: 1245INDLIDPRLSE-FNMEEVERLVGIGIV 1322
Score = 28.1 bits (61), Expect(2) = 3e-69
Identities = 13/22 (59%), Positives = 15/22 (68%)
Frame = +1
Query: 756 GLLCLQEDPIERPTMSNVLFML 777
GLLC Q P RP+MS V+ ML
Sbjct: 1315 GLLCTQTSPNLRPSMSRVVAML 1380
>TC78270 similar to GP|16649103|gb|AAL24403.1 Unknown protein {Arabidopsis
thaliana}, partial (80%)
Length = 1463
Score = 247 bits (631), Expect = 1e-65
Identities = 130/280 (46%), Positives = 184/280 (65%), Gaps = 9/280 (3%)
Frame = +3
Query: 510 ATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRN 569
A++NF+ ANK+G+GGFG VYKG GG+ A+K LS+ S QG++EF E+ +I+ ++H N
Sbjct: 297 ASDNFSPANKIGEGGFGSVYKGVLKGGKLAAIKVLSTESKQGVKEFLTEINVISEIKHEN 476
Query: 570 LVRLLGYCVEGDEKMLVYEYMPNRSLDAFI---------FDWDMRFKIILGIARGLLYLH 620
LV L G CVEGD ++LVY Y+ N SL + FDW R +I LG+ARGL +LH
Sbjct: 477 LVILYGCCVEGDHRILVYNYLENNSLSQTLLAGGHSNIYFDWQTRRRICLGVARGLAFLH 656
Query: 621 EDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYA 680
E+ I+HRD+KASNILLD++ PKISDFGLA++ T +T RV GT GY++PEYA
Sbjct: 657 EEVLPHIVHRDIKASNILLDKDLTPKISDFGLAKLIPSYMTHVST-RVAGTIGYLAPEYA 833
Query: 681 LDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTL 740
+ G + K+D++SFGV+++EI+SG+ NT P + +L W L++ +D +L
Sbjct: 834 IRGQLTRKADIYSFGVLLVEIVSGRSNTNTRLPIADQYILETTWQLYERKELAQLVDISL 1013
Query: 741 SQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSE 780
+ AEE K + + LLC Q+ P RPTMS+V+ ML E
Sbjct: 1014NGEFDAEEACKILKIALLCTQDTPKLRPTMSSVVKMLTGE 1133
>TC92541 similar to PIR|T02153|T02153 protein kinase homolog T1F15.1 -
Arabidopsis thaliana, partial (22%)
Length = 791
Score = 137 bits (344), Expect(3) = 1e-65
Identities = 69/117 (58%), Positives = 86/117 (72%), Gaps = 8/117 (6%)
Frame = +2
Query: 508 LDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQH 567
L+AT +F+ NKLGQGG+GPVYKG P GQEIAVKRLS S QG+ EFKNE+VLI LQH
Sbjct: 2 LEATIDFSPENKLGQGGYGPVYKGILPTGQEIAVKRLSKTSRQGIVEFKNELVLICELQH 181
Query: 568 RNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIF--------DWDMRFKIILGIARGL 616
NLV+LLG C+ +E++L+YEYM N+SLD ++F DW R II GI++GL
Sbjct: 182 TNLVQLLGCCIHEEERILIYEYMSNKSLDFYLFDSTRRKCLDWKKRLNIIEGISQGL 352
Score = 92.4 bits (228), Expect(3) = 1e-65
Identities = 48/112 (42%), Positives = 66/112 (58%), Gaps = 25/112 (22%)
Frame = +1
Query: 650 FGLARIFGGKDTVGNTDRVVGT-------------------------YGYMSPEYALDGH 684
FG+AR+F +++V NT+R+VGT GYMSPEYA++G
Sbjct: 454 FGMARMFTQQESVVNTNRIVGT**VL*TFLMIT*RKRF*FFK*FFLSSGYMSPEYAMEGI 633
Query: 685 FSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFI 736
S KSDV+SFGV++LEII G+RN FY + L+L+G+AW LW G L +
Sbjct: 634 CSTKSDVYSFGVLLLEIICGRRNNSFYDVDRPLNLIGHAWELWNDGEYLQLM 789
Score = 60.8 bits (146), Expect(3) = 1e-65
Identities = 29/34 (85%), Positives = 31/34 (90%)
Frame = +3
Query: 617 LYLHEDSRLRIIHRDLKASNILLDEEKNPKISDF 650
LYLH+ SRL+IIHRDLKASNILLDE NPKISDF
Sbjct: 354 LYLHKYSRLKIIHRDLKASNILLDENMNPKISDF 455
>CB891285 weakly similar to PIR|T14470|T14 receptor-like kinase (EC 2.7.1.-)
SFR2 - wild cabbage, partial (18%)
Length = 753
Score = 242 bits (618), Expect = 4e-64
Identities = 119/212 (56%), Positives = 154/212 (72%)
Frame = +2
Query: 613 ARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTY 672
ARGLLYLH DS+LRIIHRDLK SNILLDEE NPKISDFG+ARIF G++ NT RVVGTY
Sbjct: 2 ARGLLYLHRDSKLRIIHRDLKTSNILLDEELNPKISDFGMARIFEGREDTENTIRVVGTY 181
Query: 673 GYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRT 732
GY+SPEYA+ G FS KSDVFSFGV++LEIISG+RN+ FY EH L+LLG+ W WK G
Sbjct: 182 GYISPEYAMQGLFSEKSDVFSFGVLLLEIISGRRNSSFYDNEHALTLLGFVWIQWKEGNI 361
Query: 733 LDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAF 792
L FI+ + + ++C+++GLLC+QE ++RP M+ V+ ML SE+ LP P +PAF
Sbjct: 362 LSFINTEIYDPSHHKYVVRCIHIGLLCVQELAVDRPNMAAVISMLNSEAELLPPPSQPAF 541
Query: 793 VIRRCPSSRASTSSKMETFSRNDLTVTLENGR 824
++R+ S S +S N +++T +GR
Sbjct: 542 ILRQNMLSTMSHEESHRRYSINSVSITDISGR 637
>TC80500 similar to PIR|G96602|G96602 probable receptor protein kinase
F14G9.24 [imported] - Arabidopsis thaliana, partial
(13%)
Length = 1015
Score = 239 bits (610), Expect = 3e-63
Identities = 126/257 (49%), Positives = 174/257 (67%), Gaps = 6/257 (2%)
Frame = +1
Query: 530 KGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEY 589
+G G+++AVK+LS S QG +F E+ I+ +QHRNLV+L G C+EG +++LVYEY
Sbjct: 19 QGILNDGRDVAVKQLSIGSHQGKSQFVAEIATISAVQHRNLVKLYGCCIEGSKRLLVYEY 198
Query: 590 MPNRSLDAFIF------DWDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEK 643
+ N+SLD +F +W R+ I +G+ARGL YLHE+SRLRI+HRD+KASNILLD E
Sbjct: 199 LENKSLDQALFGNVLFLNWSTRYDICMGVARGLTYLHEESRLRIVHRDVKASNILLDSEL 378
Query: 644 NPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIIS 703
PKISDFGLA+++ K T +T RV GT GY++PEYA+ GH + K+DVFSFGVV LE++S
Sbjct: 379 VPKISDFGLAKLYDDKKTHIST-RVAGTIGYLAPEYAMRGHLTEKADVFSFGVVALELVS 555
Query: 704 GKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQED 763
G+ N+ ++ LL +AW L + + ID LS+ K EE + V + LLC Q
Sbjct: 556 GRPNSDSTLEGEKMYLLEWAWQLHERNTINELIDPRLSEFNK-EEVQRLVGIALLCTQTS 732
Query: 764 PIERPTMSNVLFMLGSE 780
P RP+MS V+ ML +
Sbjct: 733 PTLRPSMSRVVAMLSGD 783
>TC87048 similar to GP|8778594|gb|AAF79602.1| F5M15.3 {Arabidopsis
thaliana}, partial (77%)
Length = 1571
Score = 226 bits (576), Expect = 3e-59
Identities = 124/314 (39%), Positives = 187/314 (59%), Gaps = 13/314 (4%)
Frame = +1
Query: 485 ESGRLQEDDAKAIDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRL 544
E G+ + F + AT F AN +G+GGFG V+KG+ G+ +AVK+L
Sbjct: 379 EKGKSVRTGKSSTAAASFGFRELATATRGFKEANLIGEGGFGKVFKGRLSTGELVAVKQL 558
Query: 545 SSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--- 601
S QG +EF EV++++ L H NLV+L+GYC +GD+++LVYEYMP SL+ +FD
Sbjct: 559 SHDGRQGFQEFVTEVLMLSLLHHSNLVKLIGYCTDGDQRLLVYEYMPMGSLEDHLFDLPQ 738
Query: 602 ------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARI 655
W R KI +G ARGL YLH + +I+RDLK++NILLD + +PK+SDFGLA++
Sbjct: 739 DKEPLSWSSRMKIAVGAARGLEYLHCKADPPVIYRDLKSANILLDSDFSPKLSDFGLAKL 918
Query: 656 FGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEH 715
D + RV+GTYGY +PEYA+ G ++KSD++SFGVV+LE+I+G+R +
Sbjct: 919 GPVGDNTHVSTRVMGTYGYCAPEYAMSGKLTLKSDIYSFGVVLLELITGRRAIDASKKPG 1098
Query: 716 ELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECM-KCVNVGLLCLQEDPIERPTMSNV- 773
E +L+ ++ + R + L Q C+ + + + +CLQE P RP + ++
Sbjct: 1099EQNLVSWSRPYFSDRRKFVHMADPLLQGHFPVRCLHQAIAITAMCLQEQPKFRPLIGDIV 1278
Query: 774 --LFMLGSESNTLP 785
L L S+S +P
Sbjct: 1279VALEYLASQSQNIP 1320
>TC88105 similar to GP|21593085|gb|AAM65034.1 Putative protein kinase
{Arabidopsis thaliana}, partial (72%)
Length = 2379
Score = 220 bits (560), Expect = 2e-57
Identities = 119/298 (39%), Positives = 183/298 (60%), Gaps = 10/298 (3%)
Frame = +2
Query: 502 FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVL 561
F L + ATN F+ N +G+GG+G VY+G+ G +A+K+L + GQ +EF+ EV
Sbjct: 689 FTLRDLELATNKFSKDNIIGEGGYGVVYQGQLINGNPVAIKKLLNNLGQAEKEFRVEVEA 868
Query: 562 IARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFI---------FDWDMRFKIILGI 612
I ++H+NLVRLLG+C+EG ++L+YEY+ N +L+ ++ WD R KI+LG
Sbjct: 869 IGHVRHKNLVRLLGFCIEGTHRLLIYEYVNNGNLEQWLHGAMRQYGYLTWDARIKILLGT 1048
Query: 613 ARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFG-GKDTVGNTDRVVGT 671
A+ L YLHE +++HRD+K+SNIL+D++ N KISDFGLA++ G GK + T RV+GT
Sbjct: 1049AKALAYLHEAIEPKVVHRDIKSSNILIDDDFNAKISDFGLAKLLGAGKSHI--TTRVMGT 1222
Query: 672 YGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGR 731
+GY++PEYA G + KSDV+SFGV++LE I+G+ + + E++L+ + +
Sbjct: 1223FGYVAPEYANSGLLNEKSDVYSFGVLLLEAITGRDPVDYNRSAAEVNLVDWLKMMVGNRH 1402
Query: 732 TLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPRE 789
+ +D + + + L C+ D +RP MS V+ ML ES P PRE
Sbjct: 1403AEEVVDPNIETRPSTSALKRVLLTALRCVDPDSEKRPKMSQVVRML--ESEEYPIPRE 1570
>TC92127 weakly similar to GP|16040952|dbj|BAB69683. receptor kinase 5
{Brassica rapa}, partial (18%)
Length = 507
Score = 218 bits (555), Expect = 7e-57
Identities = 102/166 (61%), Positives = 136/166 (81%)
Frame = +3
Query: 612 IARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGT 671
IA+GLLYLH+ SRLRIIHRDLKASNILLDE NPKISDFG+AR+F ++T NT+R+VGT
Sbjct: 3 IAQGLLYLHKYSRLRIIHRDLKASNILLDENMNPKISDFGVARMFTRQETKANTNRIVGT 182
Query: 672 YGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGR 731
YGYMSPEYA++G FS KSDV+SFGV++LEII+G++N FY + L+L+G+AW LWK G
Sbjct: 183 YGYMSPEYAMEGVFSTKSDVYSFGVLLLEIINGEKNNSFYCEDRPLNLVGHAWELWKEGV 362
Query: 732 TLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFML 777
L+ +D L+++ +E ++CV+ GLLC++E+ +RPTMSNV+ ML
Sbjct: 363 VLELVDPLLNESFSEDEVLRCVHAGLLCVEENADDRPTMSNVIAML 500
>TC92229 similar to PIR|B86369|B86369 hypothetical protein AAC98010.1
[imported] - Arabidopsis thaliana, partial (44%)
Length = 1347
Score = 218 bits (554), Expect = 1e-56
Identities = 118/289 (40%), Positives = 175/289 (59%), Gaps = 11/289 (3%)
Frame = +2
Query: 502 FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVL 561
F + IL+ TN F+ N +G+GGFG VYK P G+ A+K L + SGQG EF+ EV
Sbjct: 158 FSYDQILEITNGFSSENVIGEGGFGRVYKALMPDGRVGALKLLKAGSGQGEREFRAEVDT 337
Query: 562 IARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAF-------IFDWDMRFKIILGIAR 614
I+R+ HR+LV L+GYC+ +++L+YE++PN +LD + DW R KI +G AR
Sbjct: 338 ISRVHHRHLVSLIGYCIAEQQRVLIYEFVPNGNLDQHLHESQWNVLDWPKRMKIAIGAAR 517
Query: 615 GLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGY 674
GL YLHE +IIHRD+K+SNILLD+ +++DFGLAR+ +T +T RV+GT+GY
Sbjct: 518 GLAYLHEGCNPKIIHRDIKSSNILLDDSYEAQVADFGLARLTDDTNTHVST-RVMGTFGY 694
Query: 675 MSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLW----KVG 730
M+PEYA G + +SDVFSFGVV+LE+++G++ QP + SL+ +A + + G
Sbjct: 695 MAPEYATSGKLTDRSDVFSFGVVLLELVTGRKPVDPTQPVGDESLVEWARPILLRAIETG 874
Query: 731 RTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGS 779
+ D L + E + + C++ +RP M + L S
Sbjct: 875 DFSELADPRLHRQYIDSEMFRMIEAAAACIRHSAPKRPRMVQIARALDS 1021
>TC79033 similar to GP|21554229|gb|AAM63304.1 somatic embryogenesis
receptor-like kinase putative {Arabidopsis thaliana},
partial (77%)
Length = 1762
Score = 212 bits (539), Expect = 5e-55
Identities = 118/289 (40%), Positives = 171/289 (58%), Gaps = 9/289 (3%)
Frame = +1
Query: 502 FHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVL 561
F L+ + ATNNF NKLG+GGFG VY G+ G +IAVKRL S + EF EV +
Sbjct: 427 FSLKELHSATNNFNYDNKLGEGGFGSVYWGQLWDGSQIAVKRLKVWSNKADMEFAVEVEI 606
Query: 562 IARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSL---------DAFIFDWDMRFKIILGI 612
+AR++H+NL+ L GYC EG E+++VY+YMPN SL + DW+ R I +G
Sbjct: 607 LARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSTESLLDWNRRMNIAIGS 786
Query: 613 ARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTY 672
A G++YLH + IIHRD+KASN+LLD + +++DFG A++ T T RV GT
Sbjct: 787 AEGIVYLHVQATPHIIHRDVKASNVLLDSDFQARVADFGFAKLIPDGAT-HVTTRVKGTL 963
Query: 673 GYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRT 732
GY++PEYA+ G + DV+SFG+++LE+ SGK+ + ++ +A L +
Sbjct: 964 GYLAPEYAMLGKANESCDVYSFGILLLELASGKKPLEKLSSSVKRAINDWALPLACEKKF 1143
Query: 733 LDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSES 781
+ D L+ EE + + V L+C Q P +RPTM V+ +L ES
Sbjct: 1144SELADPRLNGDYVEEELKRVILVALICAQNQPEKRPTMVEVVELLKGES 1290
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,348,051
Number of Sequences: 36976
Number of extensions: 451131
Number of successful extensions: 3700
Number of sequences better than 10.0: 730
Number of HSP's better than 10.0 without gapping: 2954
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3023
length of query: 824
length of database: 9,014,727
effective HSP length: 104
effective length of query: 720
effective length of database: 5,169,223
effective search space: 3721840560
effective search space used: 3721840560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0203.7


