

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.5
(852 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
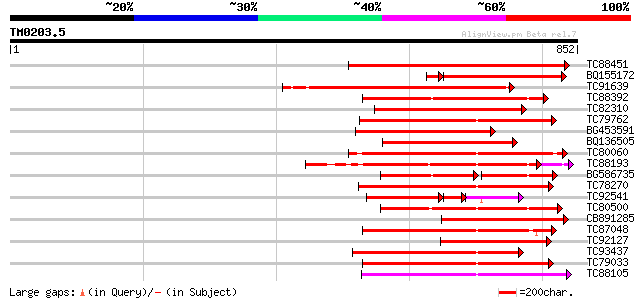
Score E
Sequences producing significant alignments: (bits) Value
TC88451 similar to GP|836954|gb|AAC23542.1|| receptor protein ki... 376 e-104
BQ155172 weakly similar to GP|6686398|gb F1E22.15 {Arabidopsis t... 313 5e-92
TC91639 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1... 333 1e-91
TC88392 similar to GP|11275527|dbj|BAB18292. putative receptor-l... 308 4e-84
TC82310 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1... 306 3e-83
TC79762 similar to PIR|H96731|H96731 hypothetical protein F5A18.... 293 1e-79
BG453591 similar to PIR|T05181|T05 S-receptor kinase (EC 2.7.1.-... 290 2e-78
BQ136505 similar to PIR|T05754|T05 S-receptor kinase (EC 2.7.1.-... 283 3e-76
TC80060 similar to PIR|D96574|D96574 hypothetical protein T3F20.... 282 3e-76
TC88193 similar to PIR|G96602|G96602 probable receptor protein k... 263 2e-74
BG586735 similar to GP|11275527|db putative receptor-like protei... 177 1e-71
TC78270 similar to GP|16649103|gb|AAL24403.1 Unknown protein {Ar... 265 7e-71
TC92541 similar to PIR|T02153|T02153 protein kinase homolog T1F1... 147 3e-68
TC80500 similar to PIR|G96602|G96602 probable receptor protein k... 249 2e-66
CB891285 weakly similar to PIR|T14470|T14 receptor-like kinase (... 234 8e-62
TC87048 similar to GP|8778594|gb|AAF79602.1| F5M15.3 {Arabidopsi... 230 2e-60
TC92127 weakly similar to GP|16040952|dbj|BAB69683. receptor kin... 225 6e-59
TC93437 similar to GP|11227578|emb|CAC16506. unnamed protein pro... 223 3e-58
TC79033 similar to GP|21554229|gb|AAM63304.1 somatic embryogenes... 220 2e-57
TC88105 similar to GP|21593085|gb|AAM65034.1 Putative protein ki... 218 8e-57
>TC88451 similar to GP|836954|gb|AAC23542.1|| receptor protein kinase
{Ipomoea trifida}, partial (28%)
Length = 1276
Score = 376 bits (966), Expect = e-104
Identities = 185/334 (55%), Positives = 248/334 (73%), Gaps = 2/334 (0%)
Frame = +3
Query: 510 DLIDKEGLE--EKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREI 567
DL++ ++ E D +G ++ F+F SIL AT FS NKLG+GGYGPVYKG L G+EI
Sbjct: 126 DLVESYDIKDLEDDFKGHDIKVFNFTSILEATMEFSPENKLGQGGYGPVYKGILATGQEI 305
Query: 568 AVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFV 627
AVKRLS S QGI EFKNE++LI +LQH+NLV+L G CI +E+ILIYEYMPNKSLD ++
Sbjct: 306 AVKRLSKTSGQGIVEFKNELLLICELQHKNLVQLLGCCIHEEERILIYEYMPNKSLDFYL 485
Query: 628 FDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFG 687
FD TK LLDW+ RF+I+ GIA+GLLYLH+ SRL++IHRDLK SNILLD M PKI+DFG
Sbjct: 486 FDCTKKKLLDWKKRFNIIEGIAQGLLYLHKYSRLKIIHRDLKASNILLDENMNPKIADFG 665
Query: 688 LARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFY 747
+AR+F +E+ NT R+VGTYGYMSPEYA++G STKSD++SFGV+LLEI+ G KN FY
Sbjct: 666 MARMFTQQESVVNTNRIVGTYGYMSPEYAMEGVCSTKSDVYSFGVLLLEIVCGIKNNSFY 845
Query: 748 QYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMS 807
L+L+G+AW+LW + + L LMD +L + + ++ RC HVGLLCV+ +DRP MS
Sbjct: 846 DVDRPLNLIGHAWELWNDGEYLKLMDPTLNDTFVPDEVKRCIHVGLLCVEQYANDRPTMS 1025
Query: 808 NVVIMLDSETATLPTPKQPTFFTRKDLSSTASSS 841
V+ +L ++ P++P F+ R+++ ++S
Sbjct: 1026EVISVLTNKYVLTNLPRKPAFYVRREIFEGETTS 1127
>BQ155172 weakly similar to GP|6686398|gb F1E22.15 {Arabidopsis thaliana},
partial (10%)
Length = 637
Score = 313 bits (802), Expect(2) = 5e-92
Identities = 156/186 (83%), Positives = 168/186 (89%), Gaps = 1/186 (0%)
Frame = +3
Query: 652 LLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYM 711
LLYLHQDSRLRVIHRDLKTSNILLD EMQPKISDFGLARI GGKETEA+T+RVVGTYGYM
Sbjct: 78 LLYLHQDSRLRVIHRDLKTSNILLDEEMQPKISDFGLARIVGGKETEASTERVVGTYGYM 257
Query: 712 SPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDL 771
SPEYALDG FSTKSD+FSFGVVLLEIISGKKNTGFY+ K SLLGYAW+LW E+KL DL
Sbjct: 258 SPEYALDGYFSTKSDVFSFGVVLLEIISGKKNTGFYRSKEISSLLGYAWRLWREDKLQDL 437
Query: 772 MDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTFFTR 831
MD +L + YN NQFIRC+ +GLLCVQDEPDDRP+MSNVV MLD+ET TL TPKQPTFFTR
Sbjct: 438 MDPTLSDTYNVNQFIRCSQIGLLCVQDEPDDRPHMSNVVTMLDNETTTLLTPKQPTFFTR 617
Query: 832 -KDLSS 836
KDLSS
Sbjct: 618 NKDLSS 635
Score = 44.3 bits (103), Expect(2) = 5e-92
Identities = 18/25 (72%), Positives = 22/25 (88%)
Frame = +2
Query: 627 VFDPTKSALLDWQMRFDILLGIARG 651
+FD TKS +LDW MRF+I+LGIARG
Sbjct: 2 IFDSTKSVILDWPMRFEIILGIARG 76
>TC91639 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1.-)
T6K22.120 precursor - Arabidopsis thaliana, partial
(28%)
Length = 1313
Score = 333 bits (855), Expect = 1e-91
Identities = 173/349 (49%), Positives = 240/349 (68%), Gaps = 1/349 (0%)
Frame = +3
Query: 411 CWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILA 470
C +W +L + + G+ L +R+A SD+ + G + + +I+ +L G++ +
Sbjct: 27 CMVWYGDLVDILH-FQHGEGNALHIRLAYSDLGDG---GKNEKIMMVIILTSLAGLICIG 194
Query: 471 CICILAYVCRRKIALKLKQESESI-LRQRGRFYDSERHVKDLIDKEGLEEKDNEGIEVPY 529
I +L + +R++ + S+ + + + + + ++ GLE +E+P+
Sbjct: 195 IIVLLVWRYKRQLKASCSKNSDVLPVFDAHKSREMSAEIPGSVEL-GLEGNQLSKVELPF 371
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVL 589
F+F + AT+ FS+ NKLG+GG+GPVYKGKL G EIAVKRLS S QG+ EFKNE+ L
Sbjct: 372 FNFSCMSSATNNFSEENKLGQGGFGPVYKGKLPSGEEIAVKRLSRRSGQGLDEFKNEMRL 551
Query: 590 IAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIA 649
A+LQHRNLV+L G I+GDEK+L+YE+M NKSLD F+FDP K LDW R++I+ GIA
Sbjct: 552 FAQLQHRNLVKLMGCSIEGDEKLLVYEFMLNKSLDRFLFDPIKKTQLDWARRYEIIEGIA 731
Query: 650 RGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYG 709
RGLLYLH+DSRLR+IHRDLK SNILLD M PKISDFGLARIFGG + E N +VVGTYG
Sbjct: 732 RGLLYLHRDSRLRIIHRDLKASNILLDENMNPKISDFGLARIFGGNQNEENATKVVGTYG 911
Query: 710 YMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGY 758
YMSPEYA++G S KSD++SFGV+LLEI+SG++NT F ++ SL+GY
Sbjct: 912 YMSPEYAMEGLVSVKSDVYSFGVLLLEIVSGRRNTSF-RHSDDSSLIGY 1055
>TC88392 similar to GP|11275527|dbj|BAB18292. putative receptor-like protein
kinase {Oryza sativa (japonica cultivar-group)}, partial
(61%)
Length = 1017
Score = 308 bits (790), Expect = 4e-84
Identities = 148/280 (52%), Positives = 208/280 (73%)
Frame = +1
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVL 589
F+ ++ +AT++FS+ N+LGRGG+GPV+KG + G E+A+K+LS S QGI+EF NEV L
Sbjct: 181 FELNTLQLATNFFSELNQLGRGGFGPVFKGLMPNGEEVAIKKLSMESRQGIREFTNEVRL 360
Query: 590 IAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIA 649
+ ++QH+NLV L G C +G EK+L+YEY+PNKSLD F+FD +S LDW RF I+ GIA
Sbjct: 361 LLRIQHKNLVTLLGCCAEGPEKMLVYEYLPNKSLDHFLFDKKRS--LDWMTRFRIVTGIA 534
Query: 650 RGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYG 709
RGLLYLH+++ R+IHRD+K SNILLD ++ PKISDFGLAR+F G++T T R+ T+G
Sbjct: 535 RGLLYLHEEAPERIIHRDIKASNILLDEKLNPKISDFGLARLFPGEDTHVQTFRISRTHG 714
Query: 710 YMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLL 769
YM+PEYAL G S K+D+FS+GV++LEI+SG+KN LL YAWKL+ K++
Sbjct: 715 YMAPEYALRGYLSVKTDVFSYGVLVLEIVSGRKNHDLKLDAEKADLLSYAWKLYQGRKIM 894
Query: 770 DLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNV 809
DL+D ++G+ YN ++ C +GLLC Q +RP+M++V
Sbjct: 895 DLIDQNIGK-YNGDEAAMCIQLGLLCCQASLVERPDMNSV 1011
>TC82310 similar to PIR|T05181|T05181 S-receptor kinase (EC 2.7.1.-)
T6K22.120 precursor - Arabidopsis thaliana, partial
(24%)
Length = 696
Score = 306 bits (783), Expect = 3e-83
Identities = 152/228 (66%), Positives = 182/228 (79%)
Frame = +1
Query: 549 GRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKG 608
G+GG+G VYKG L +EIAVKRLS SSQG++E KNE++LIAKLQHRNLVRL CI+
Sbjct: 1 GKGGFGTVYKGVLADEKEIAVKRLSKTSSQGVEELKNEIILIAKLQHRNLVRLLACCIEQ 180
Query: 609 DEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDL 668
+EK+LIYEY+PN SLD +FD K A L W+ R +I+ GIA+GLLYLH+DSRLRVIHRDL
Sbjct: 181 NEKLLIYEYLPNSSLDFHLFDMVKGAQLAWRQRLNIINGIAKGLLYLHEDSRLRVIHRDL 360
Query: 669 KTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIF 728
K SNILLD EM PKISDFGLAR FGG + EANT RVVGTYGYM+PEYA++G FS KSD+F
Sbjct: 361 KASNILLDQEMNPKISDFGLARTFGGDQDEANTIRVVGTYGYMAPEYAMEGLFSVKSDVF 540
Query: 729 SFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSL 776
SFGV+LLEIISG+KN+ FY + SL +AW LW + K +LMD S+
Sbjct: 541 SFGVLLLEIISGRKNSKFYLSEHGQSLPIFAWNLWCKRKGFELMDPSI 684
>TC79762 similar to PIR|H96731|H96731 hypothetical protein F5A18.8
[imported] - Arabidopsis thaliana, partial (55%)
Length = 1564
Score = 293 bits (751), Expect = 1e-79
Identities = 141/297 (47%), Positives = 208/297 (69%), Gaps = 1/297 (0%)
Frame = +3
Query: 526 EVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKN 585
E F +E++L AT F+ +KLG GG+GPVYKGKL GRE+AVK+LS S+QG +EF N
Sbjct: 336 EQKIFSYETLLSATKNFNATHKLGEGGFGPVYKGKLSDGREVAVKKLSQTSNQGKKEFMN 515
Query: 586 EVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVF-DPTKSALLDWQMRFDI 644
E L+A++QH+N+V L GYC+ G EKIL+YEY+P++SLD F+F + K LDW+ RF I
Sbjct: 516 EAKLLARVQHKNVVNLLGYCVHGTEKILVYEYVPHESLDKFLFKEAEKREQLDWKRRFGI 695
Query: 645 LLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRV 704
+ G+A+GLLYLH+DS +IHRD+K SNILLD + KI+DFG+AR+F +++ T RV
Sbjct: 696 ITGVAKGLLYLHEDSHNCIIHRDIKASNILLDDKWTAKIADFGMARLFPEDQSQVKT-RV 872
Query: 705 VGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWT 764
GT GYM+PEY + G+ S K+D+FS+GV++LE+I+G++N+ F +LL +A+K++
Sbjct: 873 AGTNGYMAPEYMMHGRLSVKADVFSYGVLVLELITGQRNSSFNLXVEEHNLLDWAYKMYK 1052
Query: 765 ENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLP 821
+ + L+++D +L Q C + LLC+Q +P RP M +V+ L ++ T P
Sbjct: 1053KGRSLEIVDSALASTVLTEQVDMCIQLALLCIQGDPQLRPTMRRIVVKLSRKSPTKP 1223
>BG453591 similar to PIR|T05181|T05 S-receptor kinase (EC 2.7.1.-) T6K22.120
precursor - Arabidopsis thaliana, partial (24%)
Length = 640
Score = 290 bits (742), Expect = 2e-78
Identities = 140/212 (66%), Positives = 173/212 (81%), Gaps = 1/212 (0%)
Frame = +3
Query: 520 KDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQ-GGREIAVKRLSSVSSQ 578
+D + E+P+F+ +I+ AT+ FS+ NKLG GG+GPVYKG L REIAVKRLS S Q
Sbjct: 3 EDEQDFELPFFNLSTIIDATNDFSNDNKLGEGGFGPVYKGTLVLDRREIAVKRLSGSSKQ 182
Query: 579 GIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDW 638
G +EFKNEV+L +KLQHRNLV++ G CI+G+EK+LIYEYMPN+SLD+F+FD + LLDW
Sbjct: 183 GTREFKNEVILCSKLQHRNLVKVLGCCIQGEEKMLIYEYMPNRSLDSFLFDQAQKKLLDW 362
Query: 639 QMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETE 698
RF+I+ GIARGL+YLHQDSRLR+IHRDLK SNILLD +M PKISDFGLA+I G + E
Sbjct: 363 SKRFNIICGIARGLIYLHQDSRLRIIHRDLKPSNILLDNDMNPKISDFGLAKICGDDQVE 542
Query: 699 ANTQRVVGTYGYMSPEYALDGQFSTKSDIFSF 730
NT RVVGT+GYM+PEYA+DG FS KSD+FSF
Sbjct: 543 GNTNRVVGTHGYMAPEYAIDGLFSIKSDVFSF 638
>BQ136505 similar to PIR|T05754|T05 S-receptor kinase (EC 2.7.1.-) M4I22.110
precursor - Arabidopsis thaliana, partial (23%)
Length = 618
Score = 283 bits (723), Expect = 3e-76
Identities = 137/202 (67%), Positives = 167/202 (81%)
Frame = +2
Query: 561 LQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPN 620
L GG+E+A+KR S +S QG++EFKNEV+LIAKLQHRNLV+L G CI +EK+LIYEYMPN
Sbjct: 11 LIGGKEVAIKRNSKMSDQGLEEFKNEVLLIAKLQHRNLVKLLGCCIHREEKLLIYEYMPN 190
Query: 621 KSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQ 680
+SLD F+FD T+S LLDW R I+ G+ARGLLYLHQDSRLR+IHRDLK SNILLD M
Sbjct: 191 RSLDYFIFDETRSKLLDWSKRSHIIAGVARGLLYLHQDSRLRIIHRDLKLSNILLDALMN 370
Query: 681 PKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISG 740
PKISDFGLAR F G + EA T+++VGTYGYM PEYA+ G++S KSD+FSFGV++LEIISG
Sbjct: 371 PKISDFGLARTFCGDQVEAKTRKLVGTYGYMPPEYAVHGRYSMKSDVFSFGVIVLEIISG 550
Query: 741 KKNTGFYQYKGTLSLLGYAWKL 762
KK FY +L+LLG+AW+L
Sbjct: 551 KKIKVFYDPXHSLNLLGHAWRL 616
>TC80060 similar to PIR|D96574|D96574 hypothetical protein T3F20.24
[imported] - Arabidopsis thaliana, partial (32%)
Length = 1547
Score = 282 bits (722), Expect = 3e-76
Identities = 153/331 (46%), Positives = 212/331 (63%), Gaps = 1/331 (0%)
Frame = +1
Query: 509 KDLIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIA 568
KD DKE LE K Y+ I VAT+ F NK+G GG+GPVYKG L G IA
Sbjct: 112 KDQTDKELLELKTG------YYSLRQIKVATNNFDPKNKIGEGGFGPVYKGVLSDGAVIA 273
Query: 569 VKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVF 628
VK+LSS S QG +EF NE+ +I+ LQH NLV+L+G CI+G++ +L+YEYM N SL +F
Sbjct: 274 VKQLSSKSKQGNREFVNEIGMISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALF 453
Query: 629 -DPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFG 687
P + LDW+ R I +GIARGL YLH++SRL+++HRD+K +N+LLD + KISDFG
Sbjct: 454 GKPEQRLNLDWRTRMKICVGIARGLAYLHEESRLKIVHRDIKATNVLLDKNLNAKISDFG 633
Query: 688 LARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFY 747
LA++ + T +T R+ GT GYM+PEYA+ G + K+D++SFGVV LEI+SG NT +
Sbjct: 634 LAKLDEEENTHIST-RIAGTIGYMAPEYAMRGYLTDKADVYSFGVVALEIVSGMSNTNYR 810
Query: 748 QYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMS 807
+ + LL +A+ L + LL+L+D +LG Y++ + +R + LLC P RP MS
Sbjct: 811 PKEEFVYLLDWAYVLQEQGNLLELVDPTLGSKYSSEEAMRMLQLALLCTNPSPTLRPPMS 990
Query: 808 NVVIMLDSETATLPTPKQPTFFTRKDLSSTA 838
+VV ML+ TP Q R D ++ A
Sbjct: 991 SVVSMLEGN-----TPIQAPIIKRSDSTAGA 1068
>TC88193 similar to PIR|G96602|G96602 probable receptor protein kinase
F14G9.24 [imported] - Arabidopsis thaliana, partial
(20%)
Length = 1815
Score = 263 bits (673), Expect(2) = 2e-74
Identities = 146/354 (41%), Positives = 216/354 (60%)
Frame = +3
Query: 445 PTRKGNPKSTLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDS 504
PT P S+ +G+ + VV + +C LA ILR+R + D
Sbjct: 357 PTVNSKPPSSKKNKVGLIIGVVVGVGAVCFLAV-----------SSIFYILRRRKLYNDD 503
Query: 505 ERHVKDLIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGG 564
+ D G++ N F + + AT F+ NKLG GG+GPVYKG L G
Sbjct: 504 D-------DLVGIDTMPNT------FSYYELKNATSDFNRDNKLGEGGFGPVYKGTLNDG 644
Query: 565 REIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLD 624
R +AVK+LS S QG +F E+ I+ +QHRNLV+L+G CI+G++++L+YEY+ NKSLD
Sbjct: 645 RFVAVKQLSIGSHQGKSQFIAEIATISAVQHRNLVKLYGCCIEGNKRLLVYEYLENKSLD 824
Query: 625 AFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKIS 684
+F L+W R+D+ +G+ARGL YLH++SRLR++HRD+K SNILLD E+ PK+S
Sbjct: 825 QALFG--NVLFLNWSTRYDVCMGVARGLTYLHEESRLRIVHRDVKASNILLDSELVPKLS 998
Query: 685 DFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNT 744
DFGLA+++ K+T +T RV GT GY++PEYA+ G+ + K+D+FSFGVV LE++SG+ N+
Sbjct: 999 DFGLAKLYDDKKTHIST-RVAGTIGYLAPEYAMRGRLTEKADVFSFGVVALELVSGRPNS 1175
Query: 745 GFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQD 798
+ + LL +AW+L N + DL+D L E +N + R +G++ D
Sbjct: 1176DSSLEEDKMYLLDWAWQLHERNCINDLIDPRLSE-FNMEEVERLVGIGIVVHSD 1334
Score = 35.0 bits (79), Expect(2) = 2e-74
Identities = 21/56 (37%), Positives = 29/56 (51%)
Frame = +1
Query: 792 GLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTFFTRKDLSSTASSSLQFDSS 847
GLLC Q P+ RP+MS VV ML + +P ++T D SS+ D+S
Sbjct: 1315 GLLCTQTSPNLRPSMSRVVAMLLGDIEVSTVTSRPEYWT--DWKFGDVSSIMTDTS 1476
>BG586735 similar to GP|11275527|db putative receptor-like protein kinase
{Oryza sativa (japonica cultivar-group)}, partial (57%)
Length = 794
Score = 177 bits (450), Expect(2) = 1e-71
Identities = 85/147 (57%), Positives = 113/147 (76%)
Frame = +3
Query: 558 KGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEY 617
+G + G E+A+K+LS S QGI+EF NEV L+ ++QH+NLV L G C +G EK+L+YEY
Sbjct: 6 RGLMPNGEEVAIKKLSMESRQGIREFTNEVRLLLRIQHKNLVTLLGCCAEGPEKMLVYEY 185
Query: 618 MPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDG 677
+PNKSLD F+FD +S LDW RF I+ GIARGLLYLH+++ R+IHRD+K SNILLD
Sbjct: 186 LPNKSLDHFLFDKKRS--LDWMTRFRIVTGIARGLLYLHEEAPERIIHRDIKASNILLDE 359
Query: 678 EMQPKISDFGLARIFGGKETEANTQRV 704
++ PKISDFGLAR+F G++T T R+
Sbjct: 360 KLNPKISDFGLARLFPGEDTHVQTFRI 440
Score = 111 bits (278), Expect(2) = 1e-71
Identities = 54/115 (46%), Positives = 79/115 (67%)
Frame = +2
Query: 709 GYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKL 768
GYM+PEYAL G S K+D+FS+GV++LEI+SG+KN LL YAWKL+ K+
Sbjct: 446 GYMAPEYALRGYLSVKTDVFSYGVLVLEIVSGRKNHDLKLDAEKADLLSYAWKLYQGGKI 625
Query: 769 LDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTP 823
+DL+D ++G+ YN ++ C +GLLC Q +RP+M++V +ML S++ TLP P
Sbjct: 626 MDLIDQNIGK-YNGDEAAMCIQLGLLCCQASLVERPDMNSVNLMLSSDSFTLPKP 787
>TC78270 similar to GP|16649103|gb|AAL24403.1 Unknown protein {Arabidopsis
thaliana}, partial (80%)
Length = 1463
Score = 265 bits (676), Expect = 7e-71
Identities = 132/294 (44%), Positives = 195/294 (65%), Gaps = 1/294 (0%)
Frame = +3
Query: 524 GIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEF 583
GI + + ++ + +A+D FS ANK+G GG+G VYKG L+GG+ A+K LS+ S QG++EF
Sbjct: 255 GIRIKVYTYKELKIASDNFSPANKIGEGGFGSVYKGVLKGGKLAAIKVLSTESKQGVKEF 434
Query: 584 KNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSAL-LDWQMRF 642
E+ +I++++H NLV L+G C++GD +IL+Y Y+ N SL + S + DWQ R
Sbjct: 435 LTEINVISEIKHENLVILYGCCVEGDHRILVYNYLENNSLSQTLLAGGHSNIYFDWQTRR 614
Query: 643 DILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQ 702
I LG+ARGL +LH++ ++HRD+K SNILLD ++ PKISDFGLA++ T +T
Sbjct: 615 RICLGVARGLAFLHEEVLPHIVHRDIKASNILLDKDLTPKISDFGLAKLIPSYMTHVST- 791
Query: 703 RVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKL 762
RV GT GY++PEYA+ GQ + K+DI+SFGV+L+EI+SG+ NT +L W+L
Sbjct: 792 RVAGTIGYLAPEYAIRGQLTRKADIYSFGVLLVEIVSGRSNTNTRLPIADQYILETTWQL 971
Query: 763 WTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSE 816
+ +L L+D+SL ++A + + + LLC QD P RP MS+VV ML E
Sbjct: 972 YERKELAQLVDISLNGEFDAEEACKILKIALLCTQDTPKLRPTMSSVVKMLTGE 1133
>TC92541 similar to PIR|T02153|T02153 protein kinase homolog T1F15.1 -
Arabidopsis thaliana, partial (22%)
Length = 791
Score = 147 bits (371), Expect(3) = 3e-68
Identities = 75/117 (64%), Positives = 90/117 (76%)
Frame = +2
Query: 536 LVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQH 595
L AT FS NKLG+GGYGPVYKG L G+EIAVKRLS S QGI EFKNE+VLI +LQH
Sbjct: 2 LEATIDFSPENKLGQGGYGPVYKGILPTGQEIAVKRLSKTSRQGIVEFKNELVLICELQH 181
Query: 596 RNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGL 652
NLV+L G CI +E+ILIYEYM NKSLD ++FD T+ LDW+ R +I+ GI++GL
Sbjct: 182 TNLVQLLGCCIHEEERILIYEYMSNKSLDFYLFDSTRRKCLDWKKRLNIIEGISQGL 352
Score = 95.9 bits (237), Expect(3) = 3e-68
Identities = 49/112 (43%), Positives = 67/112 (59%), Gaps = 25/112 (22%)
Frame = +1
Query: 686 FGLARIFGGKETEANTQRVVGT-------------------------YGYMSPEYALDGQ 720
FG+AR+F +E+ NT R+VGT GYMSPEYA++G
Sbjct: 454 FGMARMFTQQESVVNTNRIVGT**VL*TFLMIT*RKRF*FFK*FFLSSGYMSPEYAMEGI 633
Query: 721 FSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLM 772
STKSD++SFGV+LLEII G++N FY L+L+G+AW+LW + + L LM
Sbjct: 634 CSTKSDVYSFGVLLLEIICGRRNNSFYDVDRPLNLIGHAWELWNDGEYLQLM 789
Score = 56.2 bits (134), Expect(3) = 3e-68
Identities = 26/34 (76%), Positives = 29/34 (84%)
Frame = +3
Query: 653 LYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDF 686
LYLH+ SRL++IHRDLK SNILLD M PKISDF
Sbjct: 354 LYLHKYSRLKIIHRDLKASNILLDENMNPKISDF 455
>TC80500 similar to PIR|G96602|G96602 probable receptor protein kinase
F14G9.24 [imported] - Arabidopsis thaliana, partial
(13%)
Length = 1015
Score = 249 bits (637), Expect = 2e-66
Identities = 127/273 (46%), Positives = 182/273 (66%)
Frame = +1
Query: 558 KGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEY 617
+G L GR++AVK+LS S QG +F E+ I+ +QHRNLV+L+G CI+G +++L+YEY
Sbjct: 19 QGILNDGRDVAVKQLSIGSHQGKSQFVAEIATISAVQHRNLVKLYGCCIEGSKRLLVYEY 198
Query: 618 MPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDG 677
+ NKSLD +F L+W R+DI +G+ARGL YLH++SRLR++HRD+K SNILLD
Sbjct: 199 LENKSLDQALFGNV--LFLNWSTRYDICMGVARGLTYLHEESRLRIVHRDVKASNILLDS 372
Query: 678 EMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEI 737
E+ PKISDFGLA+++ K+T +T RV GT GY++PEYA+ G + K+D+FSFGVV LE+
Sbjct: 373 ELVPKISDFGLAKLYDDKKTHIST-RVAGTIGYLAPEYAMRGHLTEKADVFSFGVVALEL 549
Query: 738 ISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQ 797
+SG+ N+ + LL +AW+L N + +L+D L E +N + R + LLC Q
Sbjct: 550 VSGRPNSDSTLEGEKMYLLEWAWQLHERNTINELIDPRLSE-FNKEEVQRLVGIALLCTQ 726
Query: 798 DEPDDRPNMSNVVIMLDSETATLPTPKQPTFFT 830
P RP+MS VV ML + +P + T
Sbjct: 727 TSPTLRPSMSRVVAMLSGDIEVGTVTSRPGYLT 825
>CB891285 weakly similar to PIR|T14470|T14 receptor-like kinase (EC 2.7.1.-)
SFR2 - wild cabbage, partial (18%)
Length = 753
Score = 234 bits (598), Expect = 8e-62
Identities = 112/191 (58%), Positives = 143/191 (74%)
Frame = +2
Query: 649 ARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTY 708
ARGLLYLH+DS+LR+IHRDLKTSNILLD E+ PKISDFG+ARIF G+E NT RVVGTY
Sbjct: 2 ARGLLYLHRDSKLRIIHRDLKTSNILLDEELNPKISDFGMARIFEGREDTENTIRVVGTY 181
Query: 709 GYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKL 768
GY+SPEYA+ G FS KSD+FSFGV+LLEIISG++N+ FY + L+LLG+ W W E +
Sbjct: 182 GYISPEYAMQGLFSEKSDVFSFGVLLLEIISGRRNSSFYDNEHALTLLGFVWIQWKEGNI 361
Query: 769 LDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTF 828
L ++ + + + +RC H+GLLCVQ+ DRPNM+ V+ ML+SE LP P QP F
Sbjct: 362 LSFINTEIYDPSHHKYVVRCIHIGLLCVQELAVDRPNMAAVISMLNSEAELLPPPSQPAF 541
Query: 829 FTRKDLSSTAS 839
R+++ ST S
Sbjct: 542 ILRQNMLSTMS 574
>TC87048 similar to GP|8778594|gb|AAF79602.1| F5M15.3 {Arabidopsis
thaliana}, partial (77%)
Length = 1571
Score = 230 bits (586), Expect = 2e-60
Identities = 127/302 (42%), Positives = 191/302 (63%), Gaps = 10/302 (3%)
Frame = +1
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVL 589
F F + AT F +AN +G GG+G V+KG+L G +AVK+LS QG QEF EV++
Sbjct: 430 FGFRELATATRGFKEANLIGEGGFGKVFKGRLSTGELVAVKQLSHDGRQGFQEFVTEVLM 609
Query: 590 IAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFD-PTKSALLDWQMRFDILLGI 648
++ L H NLV+L GYC GD+++L+YEYMP SL+ +FD P L W R I +G
Sbjct: 610 LSLLHHSNLVKLIGYCTDGDQRLLVYEYMPMGSLEDHLFDLPQDKEPLSWSSRMKIAVGA 789
Query: 649 ARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFG-GKETEANTQRVVGT 707
ARGL YLH + VI+RDLK++NILLD + PK+SDFGLA++ G T +T RV+GT
Sbjct: 790 ARGLEYLHCKADPPVIYRDLKSANILLDSDFSPKLSDFGLAKLGPVGDNTHVST-RVMGT 966
Query: 708 YGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTE-N 766
YGY +PEYA+ G+ + KSDI+SFGVVLLE+I+G++ + G +L+ ++ +++
Sbjct: 967 YGYCAPEYAMSGKLTLKSDIYSFGVVLLELITGRRAIDASKKPGEQNLVSWSRPYFSDRR 1146
Query: 767 KLLDLMDLSLGEAYNANQFIRCTH----VGLLCVQDEPDDRPNMSNVVIMLD---SETAT 819
K + + D L + +RC H + +C+Q++P RP + ++V+ L+ S++
Sbjct: 1147KFVHMADPLLQGHFP----VRCLHQAIAITAMCLQEQPKFRPLIGDIVVALEYLASQSQN 1314
Query: 820 LP 821
+P
Sbjct: 1315IP 1320
>TC92127 weakly similar to GP|16040952|dbj|BAB69683. receptor kinase 5
{Brassica rapa}, partial (18%)
Length = 507
Score = 225 bits (573), Expect = 6e-59
Identities = 105/166 (63%), Positives = 136/166 (81%)
Frame = +3
Query: 648 IARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGT 707
IA+GLLYLH+ SRLR+IHRDLK SNILLD M PKISDFG+AR+F +ET+ANT R+VGT
Sbjct: 3 IAQGLLYLHKYSRLRIIHRDLKASNILLDENMNPKISDFGVARMFTRQETKANTNRIVGT 182
Query: 708 YGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENK 767
YGYMSPEYA++G FSTKSD++SFGV+LLEII+G+KN FY L+L+G+AW+LW E
Sbjct: 183 YGYMSPEYAMEGVFSTKSDVYSFGVLLLEIINGEKNNSFYCEDRPLNLVGHAWELWKEGV 362
Query: 768 LLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIML 813
+L+L+D L E+++ ++ +RC H GLLCV++ DDRP MSNV+ ML
Sbjct: 363 VLELVDPLLNESFSEDEVLRCVHAGLLCVEENADDRPTMSNVIAML 500
>TC93437 similar to GP|11227578|emb|CAC16506. unnamed protein product
{Arabidopsis thaliana}, partial (62%)
Length = 978
Score = 223 bits (567), Expect = 3e-58
Identities = 115/258 (44%), Positives = 167/258 (64%), Gaps = 1/258 (0%)
Frame = +3
Query: 515 EGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSS 574
E E+ + ++ F + S+ AT F + K+G GGYG VYKG L+ G ++A+K LS
Sbjct: 168 EPAEQLHEQVMKTKSFSYNSLRSATGDFHPSCKIGGGGYGVVYKGVLRDGTQVAIKSLSV 347
Query: 575 VSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSL-DAFVFDPTKS 633
S QG EF E+ +I+ +QH NLV+L G+CI+G+ +IL+YE++ N SL + + +K
Sbjct: 348 ESKQGTHEFMTEIAMISNIQHPNLVKLIGFCIEGNHRILVYEFLENNSLTSSLLGSKSKC 527
Query: 634 ALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFG 693
LDWQ R I G A GL +LH++++ ++HRD+K SNILLD PKI DFGLA++F
Sbjct: 528 VPLDWQKRAIICRGTASGLSFLHEEAQPNIVHRDIKASNILLDENFHPKIGDFGLAKLFP 707
Query: 694 GKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTL 753
T +T RV GT GY++PEYAL Q + K+D++SFG+++LEIISGK + L
Sbjct: 708 DNVTHVST-RVAGTMGYLAPEYALLRQLTKKADVYSFGILMLEIISGKSSXKAAFGDNIL 884
Query: 754 SLLGYAWKLWTENKLLDL 771
L+ +AWKL N+LL+L
Sbjct: 885 VLVEWAWKLKEXNRLLEL 938
>TC79033 similar to GP|21554229|gb|AAM63304.1 somatic embryogenesis
receptor-like kinase putative {Arabidopsis thaliana},
partial (77%)
Length = 1762
Score = 220 bits (561), Expect = 2e-57
Identities = 116/289 (40%), Positives = 177/289 (61%), Gaps = 1/289 (0%)
Frame = +1
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVL 589
F + + AT+ F+ NKLG GG+G VY G+L G +IAVKRL S++ EF EV +
Sbjct: 427 FSLKELHSATNNFNYDNKLGEGGFGSVYWGQLWDGSQIAVKRLKVWSNKADMEFAVEVEI 606
Query: 590 IAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVF-DPTKSALLDWQMRFDILLGI 648
+A+++H+NL+ L GYC +G E++++Y+YMPN SL + + + +LLDW R +I +G
Sbjct: 607 LARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSTESLLDWNRRMNIAIGS 786
Query: 649 ARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTY 708
A G++YLH + +IHRD+K SN+LLD + Q +++DFG A++ T T RV GT
Sbjct: 787 AEGIVYLHVQATPHIIHRDVKASNVLLDSDFQARVADFGFAKLIPDGATHVTT-RVKGTL 963
Query: 709 GYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKL 768
GY++PEYA+ G+ + D++SFG++LLE+ SGKK ++ +A L E K
Sbjct: 964 GYLAPEYAMLGKANESCDVYSFGILLLELASGKKPLEKLSSSVKRAINDWALPLACEKKF 1143
Query: 769 LDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSET 817
+L D L Y + R V L+C Q++P+ RP M VV +L E+
Sbjct: 1144SELADPRLNGDYVEEELKRVILVALICAQNQPEKRPTMVEVVELLKGES 1290
>TC88105 similar to GP|21593085|gb|AAM65034.1 Putative protein kinase
{Arabidopsis thaliana}, partial (72%)
Length = 2379
Score = 218 bits (555), Expect = 8e-57
Identities = 120/317 (37%), Positives = 186/317 (57%), Gaps = 1/317 (0%)
Frame = +2
Query: 529 YFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVV 588
+F + +AT+ FS N +G GGYG VY+G+L G +A+K+L + Q +EF+ EV
Sbjct: 686 WFTLRDLELATNKFSKDNIIGEGGYGVVYQGQLINGNPVAIKKLLNNLGQAEKEFRVEVE 865
Query: 589 LIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKS-ALLDWQMRFDILLG 647
I ++H+NLVRL G+CI+G ++LIYEY+ N +L+ ++ + L W R ILLG
Sbjct: 866 AIGHVRHKNLVRLLGFCIEGTHRLLIYEYVNNGNLEQWLHGAMRQYGYLTWDARIKILLG 1045
Query: 648 IARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGT 707
A+ L YLH+ +V+HRD+K+SNIL+D + KISDFGLA++ G ++ T RV+GT
Sbjct: 1046 TAKALAYLHEAIEPKVVHRDIKSSNILIDDDFNAKISDFGLAKLLGAGKSHITT-RVMGT 1222
Query: 708 YGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENK 767
+GY++PEYA G + KSD++SFGV+LLE I+G+ + + ++L+ + +
Sbjct: 1223 FGYVAPEYANSGLLNEKSDVYSFGVLLLEAITGRDPVDYNRSAAEVNLVDWLKMMVGNRH 1402
Query: 768 LLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPT 827
+++D ++ + + R L CV + + RP MS VV ML+SE +P
Sbjct: 1403 AEEVVDPNIETRPSTSALKRVLLTALRCVDPDSEKRPKMSQVVRMLESEEYPIPREV*LL 1582
Query: 828 FFTRKDLSSTASSSLQF 844
F K S S L F
Sbjct: 1583 FLIHKIASF*ISQQLIF 1633
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,360,247
Number of Sequences: 36976
Number of extensions: 439070
Number of successful extensions: 3952
Number of sequences better than 10.0: 722
Number of HSP's better than 10.0 without gapping: 3220
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3290
length of query: 852
length of database: 9,014,727
effective HSP length: 104
effective length of query: 748
effective length of database: 5,169,223
effective search space: 3866578804
effective search space used: 3866578804
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0203.5


