

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.3
(475 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
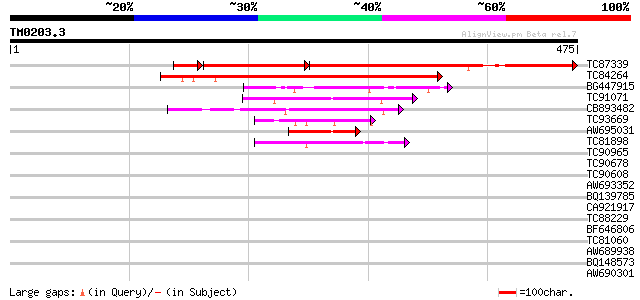
Sequences producing significant alignments: (bits) Value
TC87339 similar to PIR|D85041|D85041 hypothetical protein AT4g03... 333 e-136
TC84264 similar to PIR|G96811|G96811 unknown protein T11I11.17 [... 223 1e-58
BG447915 similar to GP|7268809|emb leucine rich repeat-like prot... 61 1e-09
TC91071 similar to GP|21536755|gb|AAM61087.1 unknown {Arabidopsi... 59 4e-09
CB893482 similar to PIR|T41744|T41 hypothetical protein F15J1.40... 57 1e-08
TC93669 weakly similar to GP|20042892|gb|AAM08720.1 Putative Cf2... 47 2e-05
AW695031 similar to GP|21536755|gb unknown {Arabidopsis thaliana... 44 1e-04
TC81898 weakly similar to GP|20260576|gb|AAM13186.1 putative rec... 42 4e-04
TC90965 similar to GP|3132477|gb|AAC16266.1| unknown protein {Ar... 40 0.003
TC90678 weakly similar to GP|9759550|dbj|BAB11152.1 receptor pro... 39 0.006
TC90608 weakly similar to GP|10716599|gb|AAG21897.1 putative dis... 37 0.013
AW693352 weakly similar to GP|20466532|g unknown protein {Arabid... 37 0.013
BQ139785 36 0.037
CA921917 similar to GP|9759078|dbj disease resistance protein-li... 35 0.063
TC88229 similar to PIR|E84471|E84471 probable beta-1 3-glucanase... 35 0.063
BF646806 weakly similar to PIR|T02565|T025 disease resistance pr... 35 0.063
TC81060 similar to GP|22136192|gb|AAM91174.1 unknown protein {Ar... 35 0.082
AW689938 weakly similar to GP|22296343|db similar to leucine-ric... 33 0.18
BQ148573 weakly similar to GP|8570048|dbj Similar to Lycopersico... 33 0.24
AW690301 weakly similar to GP|18175662|gb putative receptor prot... 33 0.24
>TC87339 similar to PIR|D85041|D85041 hypothetical protein AT4g03260
[imported] - Arabidopsis thaliana, partial (40%)
Length = 1316
Score = 333 bits (854), Expect(3) = e-136
Identities = 180/229 (78%), Positives = 189/229 (81%), Gaps = 5/229 (2%)
Frame = +2
Query: 252 ELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKI 311
ELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILD+RFNKI
Sbjct: 350 ELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDIRFNKI 529
Query: 312 STAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQLKKYLQGLLPHLVYYNRQAMKVSTLK 371
STAKCLGQLAANYNSL AINL+GNPCQKNVGDEQ+KKYLQ LLPHLVYYNRQ K + LK
Sbjct: 530 STAKCLGQLAANYNSLLAINLEGNPCQKNVGDEQIKKYLQSLLPHLVYYNRQPFKANGLK 709
Query: 372 DGADRLVRLGTN----DRSLRVDRKTTRKGSHGVAAARRPPSTSTHSHRSQTVESPKLSK 427
DG DR VRLG N DR+LRVDRK+TRKG P ST RS V SPKLSK
Sbjct: 710 DGGDRAVRLGMNSQQFDRNLRVDRKSTRKG---------PSST----RRSSAVSSPKLSK 850
Query: 428 GKQSHLPPIRTKVSTQSRHYLDAQSKVLNLTSGHSMRKSRSEGT-LGAL 475
GKQ+ LPPIRTK STQSRH+ D+ SK LNL SGH MRKSRSE T LGAL
Sbjct: 851 GKQAQLPPIRTKTSTQSRHHFDSPSKPLNLNSGHFMRKSRSESTLLGAL 997
Score = 147 bits (371), Expect(3) = e-136
Identities = 74/89 (83%), Positives = 84/89 (94%)
Frame = +1
Query: 163 VTAAGVNHKITPGMEAAKRYISSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSI 222
V A+ V+HKITPGME AKRYISSLTANA+AAQLANHGL V+PFLSAFVSLKVLNL+GN+I
Sbjct: 82 VAASVVDHKITPGMETAKRYISSLTANATAAQLANHGLSVIPFLSAFVSLKVLNLSGNAI 261
Query: 223 VRITAGALPRGLHSLNLSRNNISSIEGLR 251
VRITAG LPRGLH+LNLS+NNIS+IEGL+
Sbjct: 262 VRITAGTLPRGLHTLNLSKNNISTIEGLQ 348
Score = 43.9 bits (102), Expect(3) = e-136
Identities = 19/24 (79%), Positives = 21/24 (87%)
Frame = +3
Query: 138 GPPVEEINELPESVDPVVDINTIN 161
GPP+E+I ELPESVDP VDINT N
Sbjct: 3 GPPLEDITELPESVDPKVDINTAN 74
>TC84264 similar to PIR|G96811|G96811 unknown protein T11I11.17 [imported] -
Arabidopsis thaliana, partial (31%)
Length = 752
Score = 223 bits (569), Expect = 1e-58
Identities = 121/243 (49%), Positives = 165/243 (67%), Gaps = 7/243 (2%)
Frame = +1
Query: 127 IEDWVVDLQHCGPPVEE--INELPESVD--PVVDINTINGVTAAGVNHK---ITPGMEAA 179
++ WV DL+ P E+ ++++ S+ P D ++ + + H + + A
Sbjct: 19 VDAWVKDLEIQEPVPEDDPLDDIAGSISFPPSPDAGRSKIISTSQLTHSNSNLPKDILLA 198
Query: 180 KRYISSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNL 239
+ SL +S A ++ G+ +P +S F +L+ +NL+ N IV I+ G LP+ + +LNL
Sbjct: 199 NSMVQSLNPASSVAHISGVGIKAIPVISHFSNLRSVNLSNNFIVTISPGCLPKSVQTLNL 378
Query: 240 SRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLL 299
SRN IS+IEGL+ELTRLRVLDLSYN I RIG GL+SC+ +KELYLA NKI +VEGLHRL
Sbjct: 379 SRNKISTIEGLKELTRLRVLDLSYNCISRIGQGLSSCTIVKELYLADNKISDVEGLHRLF 558
Query: 300 KLSILDLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQLKKYLQGLLPHLVY 359
KL++LDL FNKI+T K LGQL ANYNSLQA+NL GN Q+N+GDEQL K + GLLP LVY
Sbjct: 559 KLTVLDLSFNKITTTKALGQLVANYNSLQALNLLGNAIQRNIGDEQLNKAVSGLLPKLVY 738
Query: 360 YNR 362
N+
Sbjct: 739 LNK 747
>BG447915 similar to GP|7268809|emb leucine rich repeat-like protein
{Arabidopsis thaliana}, partial (4%)
Length = 669
Score = 60.8 bits (146), Expect = 1e-09
Identities = 63/200 (31%), Positives = 101/200 (50%), Gaps = 25/200 (12%)
Frame = +3
Query: 197 NHGLV-VVPFLSAFVSLKVLNLAGNSIVRITAGALPRGLHSL--NLSRNNISSIEGLREL 253
++G++ +P LSAF SLK L+L+ N + G +P GL ++ NLS + +
Sbjct: 3 SNGIIGTLPDLSAFTSLKTLDLSSNQLT----GEIP-GLQTILRNLSSGCVRN------- 146
Query: 254 TRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKI-GEVEGLHRLLK----------LS 302
L+VLDLS NRI L++ +SLK LYL+ N++ GE+ GL +L+ L
Sbjct: 147 -SLQVLDLSDNRITGTLPDLSAFTSLKTLYLSSNQLTGEIPGLQTILRNLSSGCVRNSLQ 323
Query: 303 ILDLRFNKISTAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQLKKY-----------LQ 351
+LDL N+I+ L L+A + SL+ ++L N + Y L+
Sbjct: 324 VLDLSDNRITGT--LPDLSA-FTSLKTLDLSSNQLSGEIPGGSSLPYQLEHLSITSNTLE 494
Query: 352 GLLPHLVYYNRQAMKVSTLK 371
G++P + N A K+ +LK
Sbjct: 495 GVIPKSFWMN--ACKLKSLK 548
>TC91071 similar to GP|21536755|gb|AAM61087.1 unknown {Arabidopsis
thaliana}, partial (58%)
Length = 684
Score = 58.9 bits (141), Expect = 4e-09
Identities = 45/158 (28%), Positives = 80/158 (50%), Gaps = 12/158 (7%)
Frame = +3
Query: 196 ANHGLVVVPFLSAFVSLKVLNLAGN-----SIVRITAGALPRGLHSLNLSRNNISSIEGL 250
AN + P ++ LK L+L N ++V +++ L L L N + +I +
Sbjct: 180 ANRLSTLDPRIAQLSDLKNLSLRQNLITDAAVVPLSSWNTLSSLEELVLRDNQLKNIPDV 359
Query: 251 RELTRLRVLDLSYNRILRIGHGLAS-CSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFN 309
+L V D+S+N I + HG++ C++LKELY++ N++ ++E + +L IL+L N
Sbjct: 360 SIFKKLLVFDVSFNEIASL-HGVSRVCNTLKELYVSKNEVTKIEEIEHFHQLQILELGSN 536
Query: 310 K------ISTAKCLGQLAANYNSLQAINLDGNPCQKNV 341
K + T L +L N ++ +NL G C K +
Sbjct: 537 KLRVMENLQTLVNLQELWLGRNRIKVVNLCGLKCIKKI 650
>CB893482 similar to PIR|T41744|T41 hypothetical protein F15J1.40 -
Arabidopsis thaliana, partial (39%)
Length = 658
Score = 57.4 bits (137), Expect = 1e-08
Identities = 58/210 (27%), Positives = 108/210 (50%), Gaps = 12/210 (5%)
Frame = +1
Query: 133 DLQHCGPPVEEINELPESVDPVVDINTINGVTAAGVNHKITPGMEAAKRYISSLTANASA 192
DL+ G +++++ LP+S+ + + T++ ++ + + +SSLT
Sbjct: 55 DLKLQGKLMDQVDWLPDSIGKLSSLVTLD------LSENRIVAIPSTIGGLSSLTK---- 204
Query: 193 AQLANHGLVVVP-FLSAFVSLKVLNLAGNSIVRITAGA--LPRGLHSLNLSRNNISSI-E 248
L ++ + +P + +SL L L GNS+ + A L R L L++S N I+ + +
Sbjct: 205 LDLHSNRITEIPDSVGNLLSLVHLYLRGNSLTTLPASVSRLIR-LEELDVSSNLITVLPD 381
Query: 249 GLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLSILDLR 307
+ L L+VL++ N I I + + +CSSL+EL+ NK+ + E L ++ L IL +R
Sbjct: 382 SIGSLVSLKVLNVETNDIEEIPYSIGNCSSLRELHADYNKLKALPEALGKIESLEILSVR 561
Query: 308 FNKI-------STAKCLGQLAANYNSLQAI 330
+N I ST L +L ++N L++I
Sbjct: 562 YNNIKQLPTTMSTLINLKELNVSFNELESI 651
Score = 40.4 bits (93), Expect = 0.001
Identities = 33/98 (33%), Positives = 51/98 (51%), Gaps = 5/98 (5%)
Frame = +1
Query: 221 SIVRITAGALPRGLHSLNLSRNNISSIEGLRE----LTRLRVLDLSYNRILRIGHGLASC 276
SI+ ++A +G L L + ++ L + L+ L LDLS NRI+ I +
Sbjct: 19 SIIEVSA---KKGTRDLKLQGKLMDQVDWLPDSIGKLSSLVTLDLSENRIVAIPSTIGGL 189
Query: 277 SSLKELYLAGNKIGEV-EGLHRLLKLSILDLRFNKIST 313
SSL +L L N+I E+ + + LL L L LR N ++T
Sbjct: 190 SSLTKLDLHSNRITEIPDSVGNLLSLVHLYLRGNSLTT 303
>TC93669 weakly similar to GP|20042892|gb|AAM08720.1 Putative Cf2/Cf5
disease resistance protein homolog {Oryza sativa},
partial (4%)
Length = 727
Score = 47.0 bits (110), Expect = 2e-05
Identities = 36/113 (31%), Positives = 65/113 (56%), Gaps = 12/113 (10%)
Frame = +3
Query: 206 LSAFVSLKVLNLAGNSIVRITAGALPRGLHSLN------LSRNNISSI--EGLRELTRLR 257
LS+F+ L L+L+GN++ ++ +P LH +N +S + +S I LR LT+L
Sbjct: 384 LSSFIYLSYLDLSGNNL---SSSPIPTFLHFMNQLEFLSISDSYLSGIIPNNLRNLTKLY 554
Query: 258 VLDLSYNRILRIG--HGLASCSSLKELYLAGNKIGEVEGLHRLLKL--SILDL 306
LDLS+N L + ++ S L+ LYL+ +G+ + L ++L + S+++L
Sbjct: 555 FLDLSFNSYLHSDDVNWVSKLSLLQNLYLSDVFLGKAQNLFKVLTMLPSLIEL 713
>AW695031 similar to GP|21536755|gb unknown {Arabidopsis thaliana}, partial
(35%)
Length = 656
Score = 44.3 bits (103), Expect = 1e-04
Identities = 24/61 (39%), Positives = 38/61 (61%)
Frame = -1
Query: 234 LHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVE 293
L L LS N I+ +EGL L LRVLD+S N++ + + + + L++L+L N+I +E
Sbjct: 587 LEELYLSHNGITKMEGLSSLANLRVLDVSSNKLTSV-DDIHNLTQLEDLWLNDNQIESLE 411
Query: 294 G 294
G
Sbjct: 410 G 408
Score = 39.3 bits (90), Expect = 0.003
Identities = 18/42 (42%), Positives = 28/42 (65%)
Frame = -1
Query: 272 GLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFNKIST 313
G C +L+ELYL+ N I ++EGL L L +LD+ NK+++
Sbjct: 608 GFEGCIALEELYLSHNGITKMEGLSSLANLRVLDVSSNKLTS 483
>TC81898 weakly similar to GP|20260576|gb|AAM13186.1 putative receptor-like
protein kinase {Arabidopsis thaliana}, partial (29%)
Length = 1103
Score = 42.4 bits (98), Expect = 4e-04
Identities = 45/136 (33%), Positives = 69/136 (50%), Gaps = 6/136 (4%)
Frame = +3
Query: 206 LSAFVSLKVLNLAGNSIVR-ITAGALP-RGLHSLNLSRNNISSI--EGLRE-LTRLRVLD 260
LS+ VSLKVL L N VR I +G L + L S++LS N +S G + +LR L+
Sbjct: 483 LSSLVSLKVLKLDHNMFVRSIPSGILKCQSLVSIDLSSNQLSGTLPHGFGDAFPKLRTLN 662
Query: 261 LSYNRILRIGHGLASCSSLKELYLAGNKI-GEVEGLHRLLKLSILDLRFNKISTAKCLGQ 319
L+ N I + S+ L ++GN G + + +LKL LDL N+ + Q
Sbjct: 663 LAENNIYGGVSNFSRLKSIVSLNISGNSFQGSIIEVF-VLKLEALDLSRNQFQGH--ISQ 833
Query: 320 LAANYNSLQAINLDGN 335
+ N++ L ++L N
Sbjct: 834 VKYNWSHLVYLDLSEN 881
Score = 35.8 bits (81), Expect = 0.037
Identities = 34/118 (28%), Positives = 48/118 (39%), Gaps = 13/118 (11%)
Frame = +3
Query: 252 ELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGE--VEGLHRLLKLSILDLRFN 309
+L +L LDLS N+I + S +SLK L L+ N I + L DL N
Sbjct: 276 KLNKLHSLDLSNNKITTLPSDFWSLTSLKSLNLSSNHISGSLTNNIGNFGLLENFDLSKN 455
Query: 310 KISTAKCLGQLAANYNSLQAINLDGN-----------PCQKNVGDEQLKKYLQGLLPH 356
S + + ++ SL+ + LD N CQ V + L G LPH
Sbjct: 456 SFSDE--IPEALSSLVSLKVLKLDHNMFVRSIPSGILKCQSLVSIDLSSNQLSGTLPH 623
>TC90965 similar to GP|3132477|gb|AAC16266.1| unknown protein {Arabidopsis
thaliana}, partial (5%)
Length = 652
Score = 39.7 bits (91), Expect = 0.003
Identities = 26/71 (36%), Positives = 42/71 (58%), Gaps = 3/71 (4%)
Frame = +2
Query: 197 NHGLVVVPFLSAFVSLKV-LNLAGNSIVRITAGAL--PRGLHSLNLSRNNISSIEGLREL 253
N L+V+P + S + L+L G+ + +TA L L + L N +S++EG+ L
Sbjct: 398 NSRLIVLPQIEVKASDDLRLDLRGHRVRSLTASGLNLSSNLEFVYLRDNLLSTLEGVEVL 577
Query: 254 TRLRVLDLSYN 264
TR++VLDLS+N
Sbjct: 578 TRVKVLDLSFN 610
>TC90678 weakly similar to GP|9759550|dbj|BAB11152.1 receptor protein
kinase-like protein {Arabidopsis thaliana}, partial
(48%)
Length = 1392
Score = 38.5 bits (88), Expect = 0.006
Identities = 31/80 (38%), Positives = 46/80 (56%), Gaps = 2/80 (2%)
Frame = +3
Query: 234 LHSLNLSRNNISSIEGLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKI-GEV 292
L SLNLS + SSI L L+ L + + +N+ + + L+ CSSLK L L+ N I G +
Sbjct: 249 LQSLNLSGDISSSICDLPSLSYLNLANNIFNQPIPLH--LSQCSSLKSLNLSNNLIWGTI 422
Query: 293 EG-LHRLLKLSILDLRFNKI 311
+ + + LS+LDL N I
Sbjct: 423 PSQISQFVSLSVLDLSRNHI 482
>TC90608 weakly similar to GP|10716599|gb|AAG21897.1 putative disease
resistance protein (3' partial) {Oryza sativa}, partial
(6%)
Length = 1494
Score = 37.4 bits (85), Expect = 0.013
Identities = 41/146 (28%), Positives = 69/146 (47%), Gaps = 17/146 (11%)
Frame = +1
Query: 183 ISSLTANASAAQLANHGLVVVPFLSAFVSLKVLNLAGNSIVRITAGALPR-----GLHSL 237
+S++++N L+ + L P + + + L G S R+ G L SL
Sbjct: 517 LSNISSNLVELDLSFNHLEAPPSIDYGIVMNSLERLGLSGNRLKGGVFKSFMNVCTLSSL 696
Query: 238 NLSRNNISSIEGLRELTR----------LRVLDLSYNRILRIGHGLASCSSLKELYLAGN 287
+LSR N + E L+ + + L+VLD+SYN I L+ +SLK L L+ N
Sbjct: 697 DLSRQN-NLTEDLQIILQNLSSGCVRNSLQVLDISYNEIAGTLPDLSIFTSLKTLDLSSN 873
Query: 288 KI-GEV-EGLHRLLKLSILDLRFNKI 311
++ G++ EG +L D+R N +
Sbjct: 874 QLSGKIPEGSSLPFQLEYFDIRSNSL 951
Score = 31.2 bits (69), Expect = 0.91
Identities = 34/104 (32%), Positives = 53/104 (50%), Gaps = 5/104 (4%)
Frame = +1
Query: 234 LHSLNLSRNNISSI--EGLRELTRLRVLDLSYNRIL-RIGHGLASCSSLKELYLAGNKIG 290
L LNLS N++ + L +L+ L+ LDLSYN + I L +L+ELYL G+
Sbjct: 19 LKYLNLSWNHLDGLIPHQLGDLSNLQFLDLSYNFLEGSIPSQLGKLVNLQELYL-GSAYY 195
Query: 291 EVEGL--HRLLKLSILDLRFNKISTAKCLGQLAANYNSLQAINL 332
++ L ++ L+I + ST GQ +N SL ++L
Sbjct: 196 DIANLTIDNIINLTIDN------STDHSGGQWLSNLTSLTHLHL 309
>AW693352 weakly similar to GP|20466532|g unknown protein {Arabidopsis
thaliana}, partial (10%)
Length = 499
Score = 37.4 bits (85), Expect = 0.013
Identities = 28/85 (32%), Positives = 49/85 (56%), Gaps = 2/85 (2%)
Frame = +1
Query: 253 LTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEV-EGLHRLLKLSILDLRFNKI 311
LT L LDLS+N++ + ++S +L + +A NK+ E+ L L +L LDL N++
Sbjct: 4 LTNLEYLDLSFNKMKTLPPEISSLKALISMKVANNKLVELPPALTSLSRLENLDLSNNRL 183
Query: 312 STAKCLGQL-AANYNSLQAINLDGN 335
++ LG L ++ + LQ ++L N
Sbjct: 184 TS---LGSLELSSMHRLQNLSLQHN 249
Score = 30.8 bits (68), Expect = 1.2
Identities = 21/63 (33%), Positives = 34/63 (53%), Gaps = 2/63 (3%)
Frame = +1
Query: 232 RGLHSLNLSRNNISSIE-GLRELTRLRVLDLSYNRILRIGH-GLASCSSLKELYLAGNKI 289
+ L S+ ++ N + + L L+RL LDLS NR+ +G L+S L+ L L NK+
Sbjct: 76 KALISMKVANNKLVELPPALTSLSRLENLDLSNNRLTSLGSLELSSMHRLQNLSLQHNKL 255
Query: 290 GEV 292
+
Sbjct: 256 PSI 264
>BQ139785
Length = 678
Score = 35.8 bits (81), Expect = 0.037
Identities = 21/54 (38%), Positives = 28/54 (50%)
Frame = +3
Query: 253 LTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDL 306
L L LDLS+ I ++ + SL+ L L GNK V +RL LS L+L
Sbjct: 12 LKSLIYLDLSFCNISKVPDAIGELKSLERLNLQGNKFTSVPSTYRLKNLSYLNL 173
>CA921917 similar to GP|9759078|dbj disease resistance protein-like
{Arabidopsis thaliana}, partial (19%)
Length = 569
Score = 35.0 bits (79), Expect = 0.063
Identities = 32/92 (34%), Positives = 49/92 (52%), Gaps = 10/92 (10%)
Frame = -2
Query: 211 SLKVLNLAGNSIVRITAGALPRG------LHSLNLSRNNISS--IEGLRELTRLRVLDLS 262
SLKVLNL N+I +G++P L L++SRN+I L +L +L+ LD+S
Sbjct: 562 SLKVLNLGSNNI----SGSIPDSISNLIELEMLDISRNHIMGKIPSSLGQLQKLQWLDVS 395
Query: 263 YNRIL-RIGHGLASCSSLKELYLAGNKI-GEV 292
N I +I L+ +LK N++ GE+
Sbjct: 394 INGITGQIPGSLSQIINLKHASFRANRLCGEI 299
>TC88229 similar to PIR|E84471|E84471 probable beta-1 3-glucanase [imported]
- Arabidopsis thaliana, partial (69%)
Length = 1368
Score = 35.0 bits (79), Expect = 0.063
Identities = 28/115 (24%), Positives = 47/115 (40%)
Frame = +3
Query: 355 PHLVYYNRQAMKVSTLKDGADRLVRLGTNDRSLRVDRKTTRKGSHGVAAARRPPSTSTHS 414
P L + QA K++ + G R G + +R+ ++ RK PP +T S
Sbjct: 138 PTLSIQSSQASKITRNRPGQTLRHRPGCSQILIRIRNQSNRK----------PPKRTTLS 287
Query: 415 HRSQTVESPKLSKGKQSHLPPIRTKVSTQSRHYLDAQSKVLNLTSGHSMRKSRSE 469
H +T+ P ++ K+ LPP +R + Q + T H + R E
Sbjct: 288 HSQKTLLRPHMAPKKRRRLPP-----QNPNRSHRRRQRSLRRHTQHHQIPHPRHE 437
>BF646806 weakly similar to PIR|T02565|T025 disease resistance protein
homolog At2g32660 [imported] - Arabidopsis thaliana,
partial (5%)
Length = 624
Score = 35.0 bits (79), Expect = 0.063
Identities = 33/100 (33%), Positives = 52/100 (52%), Gaps = 6/100 (6%)
Frame = +3
Query: 255 RLRVLDLSYNRI---LRIGHGLASCSSLKELYLAGNKIGEV---EGLHRLLKLSILDLRF 308
RL+ LDLSYN + L I GL++ +SLK L N++ ++ EG L +L IL+L
Sbjct: 105 RLKSLDLSYNNLNGSLDIS-GLSNLTSLKILDFTSNQLVDLIVREGSKNLSRLDILNLDS 281
Query: 309 NKISTAKCLGQLAANYNSLQAINLDGNPCQKNVGDEQLKK 348
N I+ + L Q + S++ + L N + + D K
Sbjct: 282 NMINGSN-LQQWLWAFPSIRNLTLRNNQFKGTILDGDWSK 398
>TC81060 similar to GP|22136192|gb|AAM91174.1 unknown protein {Arabidopsis
thaliana}, partial (95%)
Length = 1114
Score = 34.7 bits (78), Expect = 0.082
Identities = 27/82 (32%), Positives = 47/82 (56%), Gaps = 2/82 (2%)
Frame = +3
Query: 256 LRVLDLSYNRILRIGHGLASCSSLKELYLAGNKIGEVE-GLHRLLKLSILDLRFNKISTA 314
+R LDL++NRI+ I ++ +++ L LA N I + L +L L +++L N+IS+
Sbjct: 183 VRTLDLTHNRIVDIPVEISKLINVQRLILADNLIDRLPVNLGKLQSLKLVNLDGNRISSL 362
Query: 315 KC-LGQLAANYNSLQAINLDGN 335
LGQL L+ +++ GN
Sbjct: 363 PDELGQLV----RLERLSIAGN 416
Score = 32.3 bits (72), Expect = 0.41
Identities = 26/86 (30%), Positives = 47/86 (54%), Gaps = 2/86 (2%)
Frame = +3
Query: 230 LPRGLHSLNLSRNNISSIE-GLRELTRLRVLDLSYNRILRIGHGLASCSSLKELYLAGNK 288
L R + +L+L+ N I I + +L ++ L L+ N I R+ L SLK + L GN+
Sbjct: 171 LDRSVRTLDLTHNRIVDIPVEISKLINVQRLILADNLIDRLPVNLGKLQSLKLVNLDGNR 350
Query: 289 IGEV-EGLHRLLKLSILDLRFNKIST 313
I + + L +L++L L + N +++
Sbjct: 351 ISSLPDELGQLVRLERLSIAGNLLTS 428
>AW689938 weakly similar to GP|22296343|db similar to leucine-rich repeat
transmembrane protein kinase 1, partial (16%)
Length = 655
Score = 33.5 bits (75), Expect = 0.18
Identities = 30/82 (36%), Positives = 42/82 (50%), Gaps = 5/82 (6%)
Frame = +3
Query: 206 LSAFVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNIS-----SIEGLRELTRLRVLD 260
LS +SL+ L+L+ N I LP L SLNL+RNN++ S + LT L V +
Sbjct: 351 LSDLMSLRKLDLSDNKIHDQIPYQLPPNLTSLNLARNNLTGNLPYSFSAMVSLTYLNVSN 530
Query: 261 LSYNRILRIGHGLASCSSLKEL 282
+ + L IG A+ S L L
Sbjct: 531 NALS--LPIGXVFANHSHLDTL 590
>BQ148573 weakly similar to GP|8570048|dbj Similar to Lycopersicon esculentum
Hcr2-5D (AF053998) {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 689
Score = 33.1 bits (74), Expect = 0.24
Identities = 45/155 (29%), Positives = 65/155 (41%), Gaps = 28/155 (18%)
Frame = +1
Query: 209 FVSLKVLNLAGNSIVRITAGALPRGLHSLNLSRNNISSIEG-LRELTRLRVLDLSYNRI- 266
++ L +NL S+ + LP L L+LS + S L EL L LDLSYN+I
Sbjct: 136 YLYLSQINLIPFSLHNESDFTLPN-LLGLSLSSCKLKSFPSFLNELKTLENLDLSYNQIN 312
Query: 267 -------LRIGHGLASCSSLKELYLAGNKIGEVEGLHRLLKLSILDLRFN---------- 309
+G+G +L L L+ N + L + +S +DL FN
Sbjct: 313 GRVPSWFNNLGNG-----TLSSLDLSHNLLTSTGNLSH-MNISYIDLSFNMLEGEIPLPP 474
Query: 310 ------KISTAKCLGQLAA---NYNSLQAINLDGN 335
IS K G L++ N SL+ +NL N
Sbjct: 475 FGTSFFSISNNKLTGDLSSRICNARSLEILNLSHN 579
>AW690301 weakly similar to GP|18175662|gb putative receptor protein kinase
{Arabidopsis thaliana}, partial (18%)
Length = 673
Score = 33.1 bits (74), Expect = 0.24
Identities = 46/143 (32%), Positives = 74/143 (51%), Gaps = 13/143 (9%)
Frame = +1
Query: 206 LSAFVSLKVLNLAGNSIVRITAGALPRGLHSL-NLSRNNISS--IEGL-----RELTRLR 257
LS +L+ L+L+ N++ G++P+ L L NL++ + S +EGL T L
Sbjct: 49 LSECQNLQALDLSYNNLT----GSIPKQLFVLRNLTQLMLISNDLEGLIPPDIGNCTSLY 216
Query: 258 VLDLSYNRIL-RIGHGLASCSSLKELYLAGNK-IGEVEG-LHRLLKLSILDLRFNKISTA 314
L L+ NR++ I +A+ +L L L N +GE+ L KL +LDL NK+S
Sbjct: 217 RLRLNQNRLVGTIPSEIANLKNLNFLDLHYNHLVGEIPSQFSGLSKLGVLDLSHNKLS-- 390
Query: 315 KCLGQLAA--NYNSLQAINLDGN 335
G L A N ++L ++N+ N
Sbjct: 391 ---GNLDAISNLHNLVSLNVSFN 450
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.132 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,096,781
Number of Sequences: 36976
Number of extensions: 169688
Number of successful extensions: 858
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 826
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 842
length of query: 475
length of database: 9,014,727
effective HSP length: 100
effective length of query: 375
effective length of database: 5,317,127
effective search space: 1993922625
effective search space used: 1993922625
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0203.3


