

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0201.10
(137 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
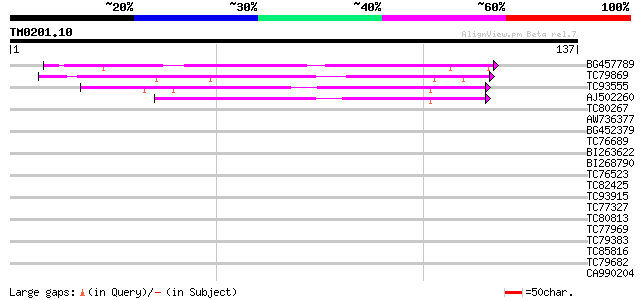
Score E
Sequences producing significant alignments: (bits) Value
BG457789 homologue to PIR|T06681|T066 hypothetical protein T17F1... 92 5e-20
TC79869 similar to GP|10177465|dbj|BAB10856. gene_id:MQB2.19~pir... 87 2e-18
TC93555 similar to PIR|T51434|T51434 hypothetical protein F2G14_... 66 4e-12
AJ502260 weakly similar to PIR|T51434|T514 hypothetical protein ... 64 1e-11
TC80267 similar to GP|6692128|gb|AAF24593.1| T19E23.14 {Arabidop... 29 0.009
AW736377 similar to PIR|E96794|E967 hypothetical protein F14G6.2... 33 0.046
BG452379 similar to SP|P51022|PNT1_ ETS-like protein pointed P1 ... 33 0.046
TC76689 MtN12 31 0.18
BI263622 homologue to GP|23496360|gb hypothetical protein {Plasm... 30 0.23
BI268790 30 0.30
TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 30 0.30
TC82425 similar to PIR|T06311|T06311 hypothetical protein F11C18... 30 0.30
TC93915 similar to GP|10178125|dbj|BAB11537. gene_id:MOP10.2~unk... 29 0.51
TC77327 homologue to GP|16224254|gb|AAL15651.1 dehydrin-like pro... 28 0.88
TC80813 similar to GP|20259227|gb|AAM14329.1 unknown protein {Ar... 28 0.88
TC77969 similar to GP|15450966|gb|AAK96754.1 Unknown protein {Ar... 28 0.88
TC79383 weakly similar to GP|12643043|gb|AAK00432.1 hypothetical... 28 0.88
TC85816 similar to GP|15021744|gb|AAK77899.1 root nodule extensi... 28 1.1
TC79682 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 28 1.5
CA990204 27 2.0
>BG457789 homologue to PIR|T06681|T066 hypothetical protein T17F15.110 -
Arabidopsis thaliana, partial (20%)
Length = 619
Score = 92.4 bits (228), Expect = 5e-20
Identities = 52/119 (43%), Positives = 70/119 (58%), Gaps = 9/119 (7%)
Frame = +1
Query: 9 QACICYLIPCSSS--SETSLSWWERVRSPENKEWWWAHGWNKVREWSEIIVGPKWKTFIR 66
+AC + PC S S T+ WWERVR+ + G+ K+REWSEI+ GP+WKTFIR
Sbjct: 67 RACT-FCFPCFGSRHSTTTEPWWERVRASSV-----SRGFMKIREWSEIVAGPRWKTFIR 228
Query: 67 RFNRNNNRAGAASYDKKGSFHYDSLSYALNFDDGEEDVY------SYGGFSTRF-ASVP 118
RFNRN + + G + YD LSYALNFD+G+ + + FS R+ A+VP
Sbjct: 229 RFNRNK----SGGFRHAGKYQYDPLSYALNFDEGQNGEFENDSPDGFRNFSARYVAAVP 393
>TC79869 similar to GP|10177465|dbj|BAB10856.
gene_id:MQB2.19~pir||T06681~similar to unknown protein
{Arabidopsis thaliana}, partial (13%)
Length = 754
Score = 87.0 bits (214), Expect = 2e-18
Identities = 49/124 (39%), Positives = 69/124 (55%), Gaps = 14/124 (11%)
Frame = +1
Query: 8 KQACICYLIPCSSSSETSLSWWERVRS--------PENKEWWWAHGWN---KVREWSEII 56
+ + +C+ P +S WWERVRS P + WW+ G K+R WSEI+
Sbjct: 139 RSSALCF--PTCFASRRRSVWWERVRSASFSQSHPPTTADRWWSRGLKALKKLRNWSEIV 312
Query: 57 VGPKWKTFIRRFNRNNNRAGAASYDKKGSFHYDSLSYALNFDDGE-EDVYSYG--GFSTR 113
GP+WK FIR+FN + ++ + YD LSYALNFD+G+ ED + G FSTR
Sbjct: 313 AGPRWKNFIRKFNNHRSK-------RMTKCQYDPLSYALNFDEGQNEDSHDDGFRNFSTR 471
Query: 114 FASV 117
+A+V
Sbjct: 472 YAAV 483
>TC93555 similar to PIR|T51434|T51434 hypothetical protein F2G14_10 -
Arabidopsis thaliana, partial (2%)
Length = 679
Score = 66.2 bits (160), Expect = 4e-12
Identities = 41/114 (35%), Positives = 58/114 (49%), Gaps = 15/114 (13%)
Frame = +2
Query: 18 CSSSSETSLSWWER----------VRSPENK--EWWWAHGWNKVREWSEIIVGPKWKTFI 65
C S + WW+R +S N+ E W K++E SE+I GPKWKTF+
Sbjct: 104 CGWSRIFTSKWWQRHDEEGKHLLNEKSEGNRGEETWMMDKLKKMKETSEVIAGPKWKTFL 283
Query: 66 RRFNRNNNRAGAASYDKKGSFHYDSLSYALNFDDG---EEDVYSYGGFSTRFAS 116
R+ +G +K F YD+ SYALNF+ G E++ Y FSTRF++
Sbjct: 284 RKI------SGYGKNQQKHRFQYDAHSYALNFNSGAQSEDEEYLPPSFSTRFSN 427
>AJ502260 weakly similar to PIR|T51434|T514 hypothetical protein F2G14_10 -
Arabidopsis thaliana, partial (3%)
Length = 667
Score = 64.3 bits (155), Expect = 1e-11
Identities = 34/83 (40%), Positives = 46/83 (54%), Gaps = 2/83 (2%)
Frame = +1
Query: 36 ENKEWWWAHGWNKVREWSEIIVGPKWKTFIRRFNRNNNRAGAASYDKKGSFHYDSLSYAL 95
+ ++ WW K++E SE+I GPKWKTFIR+ + + +KG F YD SYAL
Sbjct: 421 QREDSWWVCKLRKMKEVSEVIAGPKWKTFIRKISMYGKK------QQKGKFQYDEHSYAL 582
Query: 96 NFDDG--EEDVYSYGGFSTRFAS 116
NF G ED FS RF++
Sbjct: 583 NFSSGAQSEDDDLPHSFSARFSA 651
>TC80267 similar to GP|6692128|gb|AAF24593.1| T19E23.14 {Arabidopsis
thaliana}, partial (24%)
Length = 1625
Score = 28.9 bits (63), Expect(2) = 0.009
Identities = 12/21 (57%), Positives = 14/21 (66%)
Frame = -1
Query: 22 SETSLSWWERVRSPENKEWWW 42
SET + W ER+R E KE WW
Sbjct: 572 SETEIDWTERLR--ERKEGWW 516
Score = 25.0 bits (53), Expect(2) = 0.009
Identities = 6/17 (35%), Positives = 10/17 (58%)
Frame = -3
Query: 40 WWWAHGWNKVREWSEII 56
WWW + VR W +++
Sbjct: 495 WWWREDFPSVRYWIKVV 445
>AW736377 similar to PIR|E96794|E967 hypothetical protein F14G6.23 [imported]
- Arabidopsis thaliana, partial (7%)
Length = 588
Score = 32.7 bits (73), Expect = 0.046
Identities = 18/37 (48%), Positives = 21/37 (56%), Gaps = 2/37 (5%)
Frame = -2
Query: 82 KKGSFHYDSLSYALNFDDG--EEDVYSYGGFSTRFAS 116
+KG F YD SYALNF G ED FS RF++
Sbjct: 245 QKGKFQYDEHSYALNFSSGAQSEDDDLPHSFSARFSA 135
>BG452379 similar to SP|P51022|PNT1_ ETS-like protein pointed P1 (D-ETS-2).
[Fruit fly] {Drosophila melanogaster}, partial (3%)
Length = 625
Score = 32.7 bits (73), Expect = 0.046
Identities = 18/67 (26%), Positives = 33/67 (48%), Gaps = 2/67 (2%)
Frame = -1
Query: 15 LIPCSSSSETSLSWWERVRSPENKE--WWWAHGWNKVREWSEIIVGPKWKTFIRRFNRNN 72
L+ C ++S + +S R+ + NK W W++GW+ + KW + F NN
Sbjct: 526 LLRCMATSYSLISNISRISNRGNKGLMWVWSYGWDNTYVMES*VQPLKW*NHLCGFMSNN 347
Query: 73 NRAGAAS 79
+R+ A+
Sbjct: 346 SRSRLAT 326
>TC76689 MtN12
Length = 759
Score = 30.8 bits (68), Expect = 0.18
Identities = 14/43 (32%), Positives = 17/43 (38%), Gaps = 10/43 (23%)
Frame = -1
Query: 29 WERVRSPENKEWWWA----------HGWNKVREWSEIIVGPKW 61
W RVR WWW GW +V +W + VG W
Sbjct: 264 WLRVRDVGVNWWWWGVVDWFRVSVNWGWWRVVDWFRVRVGVNW 136
>BI263622 homologue to GP|23496360|gb hypothetical protein {Plasmodium
falciparum 3D7}, partial (0%)
Length = 655
Score = 30.4 bits (67), Expect = 0.23
Identities = 17/33 (51%), Positives = 18/33 (54%), Gaps = 4/33 (12%)
Frame = -2
Query: 13 CYLIPCSSSSETSLSWW----ERVRSPENKEWW 41
CYLI SSSSE S + W ER R E K W
Sbjct: 240 CYLIKNSSSSEQSATTWVVLSERCRECEKKSIW 142
>BI268790
Length = 416
Score = 30.0 bits (66), Expect = 0.30
Identities = 16/47 (34%), Positives = 19/47 (40%)
Frame = -2
Query: 6 FAKQACICYLIPCSSSSETSLSWWERVRSPENKEWWWAHGWNKVREW 52
FA+Q P S S WW R R P + W H W V +W
Sbjct: 331 FARQGLFPCWPPGR*RSPGSRRWWRR-RKPRRRAGWPWHSWCAVPDW 194
>TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (69%)
Length = 1615
Score = 30.0 bits (66), Expect = 0.30
Identities = 11/25 (44%), Positives = 13/25 (52%)
Frame = +2
Query: 28 WWERVRSPENKEWWWAHGWNKVREW 52
WW R R KEWWW+ W + W
Sbjct: 1112 WW*RWRW---KEWWWSIWWRWKKRW 1177
>TC82425 similar to PIR|T06311|T06311 hypothetical protein F11C18.90 -
Arabidopsis thaliana, partial (26%)
Length = 803
Score = 30.0 bits (66), Expect = 0.30
Identities = 11/23 (47%), Positives = 12/23 (51%)
Frame = -3
Query: 28 WWERVRSPENKEWWWAHGWNKVR 50
WWE V E EWWW W + R
Sbjct: 117 WWENVGGFE--EWWWRWRWPRTR 55
>TC93915 similar to GP|10178125|dbj|BAB11537. gene_id:MOP10.2~unknown
protein {Arabidopsis thaliana}, partial (29%)
Length = 1001
Score = 29.3 bits (64), Expect = 0.51
Identities = 14/32 (43%), Positives = 18/32 (55%)
Frame = -3
Query: 40 WWWAHGWNKVREWSEIIVGPKWKTFIRRFNRN 71
W W GW V + E +VGP W F+R F R+
Sbjct: 231 WGWLVGW-WVCYFYEFLVGPIWVRFLRVFLRD 139
>TC77327 homologue to GP|16224254|gb|AAL15651.1 dehydrin-like protein
{Medicago sativa}, partial (71%)
Length = 1330
Score = 28.5 bits (62), Expect = 0.88
Identities = 8/26 (30%), Positives = 14/26 (53%)
Frame = +1
Query: 41 WWAHGWNKVREWSEIIVGPKWKTFIR 66
WW WN+ R W+ + +W+ + R
Sbjct: 433 WWNRVWNRDRNWNNRLWSYRWRNWSR 510
>TC80813 similar to GP|20259227|gb|AAM14329.1 unknown protein {Arabidopsis
thaliana}, partial (8%)
Length = 498
Score = 28.5 bits (62), Expect = 0.88
Identities = 11/36 (30%), Positives = 16/36 (43%)
Frame = +3
Query: 27 SWWERVRSPENKEWWWAHGWNKVREWSEIIVGPKWK 62
SW E + WWW ++ +W + I P WK
Sbjct: 9 SW*EGLFKWSWSSWWWRRQRKRLPKWRQPIS*PAWK 116
>TC77969 similar to GP|15450966|gb|AAK96754.1 Unknown protein {Arabidopsis
thaliana}, partial (58%)
Length = 1033
Score = 28.5 bits (62), Expect = 0.88
Identities = 19/58 (32%), Positives = 25/58 (42%), Gaps = 11/58 (18%)
Frame = -3
Query: 9 QAC---ICYLIPCS---SSSETSL-----SWWERVRSPENKEWWWAHGWNKVREWSEI 55
QAC + Y I CS S ET + S WE + WWW W + R ++I
Sbjct: 281 QACGSLLHYKIACSLCHSRKETCILRRERSDWESEGKKRFEGWWWMQQWQRWRRKNKI 108
>TC79383 weakly similar to GP|12643043|gb|AAK00432.1 hypothetical protein
{Oryza sativa}, partial (8%)
Length = 1082
Score = 28.5 bits (62), Expect = 0.88
Identities = 23/74 (31%), Positives = 35/74 (47%), Gaps = 11/74 (14%)
Frame = -2
Query: 11 CICYLIPCSSSSETSLSWWERVRSPENKEW--WWAHGW-----NKVREWSEIIVGPKWKT 63
C+ L+ SSS + L W +R ++EW W A GW + V E +++ +
Sbjct: 1045 CLFKLVSRSSSIKQILQWKKRSIKGGSEEWARWMAVGWISVDCSLVEEEFRVLL*IF*EE 866
Query: 64 FIRRF----NRNNN 73
I RF NRNN+
Sbjct: 865 LITRFPLILNRNNS 824
>TC85816 similar to GP|15021744|gb|AAK77899.1 root nodule extensin {Pisum
sativum}, partial (32%)
Length = 603
Score = 28.1 bits (61), Expect = 1.1
Identities = 12/36 (33%), Positives = 18/36 (49%)
Frame = -2
Query: 27 SWWERVRSPENKEWWWAHGWNKVREWSEIIVGPKWK 62
SW + SP WWW W +V +W + V +W+
Sbjct: 230 SWCRSIESPTPVNWWW---W-RVVDWFRVSVN-RWR 138
>TC79682 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (17%)
Length = 1031
Score = 27.7 bits (60), Expect = 1.5
Identities = 14/44 (31%), Positives = 19/44 (42%)
Frame = -3
Query: 28 WWERVRSPENKEWWWAHGWNKVREWSEIIVGPKWKTFIRRFNRN 71
WW+R K WWW W W+ KW T+ ++ RN
Sbjct: 129 WWKR------K*WWWT*WWRLKWIWN------KWWTWWLKWIRN 34
>CA990204
Length = 479
Score = 27.3 bits (59), Expect = 2.0
Identities = 17/34 (50%), Positives = 20/34 (58%), Gaps = 3/34 (8%)
Frame = +3
Query: 86 FHYDSLSYALNFDDG---EEDVYSYGGFSTRFAS 116
F YDS SY NFDDG + D + FS RFA+
Sbjct: 162 FIYDSNSYLQNFDDGYSIDPDNF-LRSFSARFAA 260
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.133 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,551,220
Number of Sequences: 36976
Number of extensions: 95425
Number of successful extensions: 907
Number of sequences better than 10.0: 97
Number of HSP's better than 10.0 without gapping: 869
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 899
length of query: 137
length of database: 9,014,727
effective HSP length: 86
effective length of query: 51
effective length of database: 5,834,791
effective search space: 297574341
effective search space used: 297574341
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0201.10


