

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0199.9
(56 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
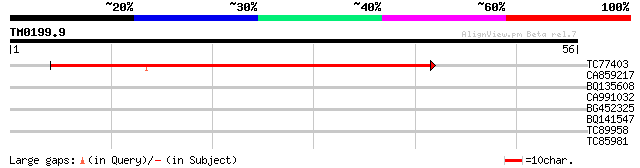
TC77403 similar to SP|Q9SDS7|VATC_ARATH Vacuolar ATP synthase su... 70 2e-13
CA859217 weakly similar to GP|23326466|gb| narrowly conserved hy... 27 2.1
BQ135608 similar to GP|4325337|gb|A F15P23.3 gene product {Arabi... 26 3.6
CA991032 weakly similar to GP|21592868|gb| unknown {Arabidopsis ... 26 3.6
BG452325 similar to GP|9294439|dbj gene_id:MMB12.21~unknown prot... 26 3.6
BQ141547 25 4.7
TC89958 similar to GP|12483623|dbj|BAB21445. NADH dehydrogenase ... 25 8.0
TC85981 similar to SP|Q00763|COMT_POPTM Caffeic acid 3-O-methylt... 25 8.0
>TC77403 similar to SP|Q9SDS7|VATC_ARATH Vacuolar ATP synthase subunit C (EC
3.6.3.14) (V-ATPase C subunit) (Vacuolar proton pump C
subunit)., partial (98%)
Length = 1783
Score = 70.1 bits (170), Expect = 2e-13
Identities = 31/40 (77%), Positives = 37/40 (92%), Gaps = 2/40 (5%)
Frame = +2
Query: 5 WWVVSLPV--HNSASELWNRLQQNVSKHSFDTPLYRFNIP 42
+WVVSLPV +NS+S +WN+LQQN+SKHSFDTPLYRFNIP
Sbjct: 62 YWVVSLPVQNNNSSSSIWNQLQQNISKHSFDTPLYRFNIP 181
>CA859217 weakly similar to GP|23326466|gb| narrowly conserved hypothetical
protein {Bifidobacterium longum NCC2705}, partial (5%)
Length = 766
Score = 26.6 bits (57), Expect = 2.1
Identities = 10/25 (40%), Positives = 18/25 (72%)
Frame = -3
Query: 3 VVWWVVSLPVHNSASELWNRLQQNV 27
V+ +V S+P+ +S+S +W+ L NV
Sbjct: 728 VLMYVYSIPLFSSSSRIWSLLSTNV 654
>BQ135608 similar to GP|4325337|gb|A F15P23.3 gene product {Arabidopsis
thaliana}, partial (4%)
Length = 759
Score = 25.8 bits (55), Expect = 3.6
Identities = 9/17 (52%), Positives = 13/17 (75%)
Frame = +3
Query: 26 NVSKHSFDTPLYRFNIP 42
+V K+ FDTP+YR +P
Sbjct: 330 DVPKYYFDTPVYRLGLP 380
>CA991032 weakly similar to GP|21592868|gb| unknown {Arabidopsis thaliana},
partial (19%)
Length = 403
Score = 25.8 bits (55), Expect = 3.6
Identities = 13/27 (48%), Positives = 17/27 (62%), Gaps = 3/27 (11%)
Frame = -1
Query: 13 HNSASELWNRLQQNVSKHS---FDTPL 36
HNS EL L QN+S+H +DTP+
Sbjct: 85 HNSHQELL*LLDQNMSQHEEHPYDTPI 5
>BG452325 similar to GP|9294439|dbj gene_id:MMB12.21~unknown protein
{Arabidopsis thaliana}, partial (25%)
Length = 666
Score = 25.8 bits (55), Expect = 3.6
Identities = 10/28 (35%), Positives = 15/28 (52%)
Frame = +2
Query: 5 WWVVSLPVHNSASELWNRLQQNVSKHSF 32
WW + P H S+ E+W L+ + SF
Sbjct: 239 WWHQNPPAHYSSHEIWPTLRVSFLLSSF 322
>BQ141547
Length = 849
Score = 25.4 bits (54), Expect = 4.7
Identities = 8/26 (30%), Positives = 16/26 (60%)
Frame = +1
Query: 19 LWNRLQQNVSKHSFDTPLYRFNIPIS 44
LW++ +N + HSF P++ F ++
Sbjct: 277 LWDKTTRNAATHSFPPPIHVFTFQMT 354
>TC89958 similar to GP|12483623|dbj|BAB21445. NADH dehydrogenase subunit 4L
{Gonostoma gracile}, partial (17%)
Length = 1040
Score = 24.6 bits (52), Expect = 8.0
Identities = 17/48 (35%), Positives = 25/48 (51%)
Frame = -1
Query: 2 AVVWWVVSLPVHNSASELWNRLQQNVSKHSFDTPLYRFNIPISGSEP* 49
A +WW++SL + SAS+ + + NV S L R N P+ P*
Sbjct: 737 AEMWWLISLDLTKSASQQSDCVTTNVPA*SH---LRRGNQPLHSYIP* 603
>TC85981 similar to SP|Q00763|COMT_POPTM Caffeic acid 3-O-methyltransferase
(EC 2.1.1.68), partial (93%)
Length = 1389
Score = 24.6 bits (52), Expect = 8.0
Identities = 8/26 (30%), Positives = 14/26 (53%)
Frame = +2
Query: 13 HNSASELWNRLQQNVSKHSFDTPLYR 38
H + S+ W++ + KH F P Y+
Sbjct: 170 H*NYSQSWSKCSSFIFKHCFSNPFYK 247
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.135 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,051,588
Number of Sequences: 36976
Number of extensions: 22229
Number of successful extensions: 186
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 184
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 185
length of query: 56
length of database: 9,014,727
effective HSP length: 32
effective length of query: 24
effective length of database: 7,831,495
effective search space: 187955880
effective search space used: 187955880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0199.9


