

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0197.8
(242 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
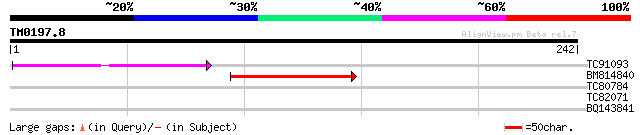
TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein... 50 6e-07
BM814840 weakly similar to GP|15128241|db helicase-like protein ... 45 2e-05
TC80784 similar to PIR|T50352|T50352 ornithine--oxo-acid transam... 29 1.9
TC82071 weakly similar to PIR|T00411|T01608 hypothetical protein... 27 5.5
BQ143841 27 9.3
>TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein T24M8.10 -
Arabidopsis thaliana, partial (11%)
Length = 701
Score = 50.4 bits (119), Expect = 6e-07
Identities = 34/85 (40%), Positives = 45/85 (52%)
Frame = +3
Query: 2 QFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLN 61
QF R ILT T + V+ IN Y+L +I E S+N RS + + FL
Sbjct: 453 QFLQSRAILTSTDEVVDQINDYVLKLIPGEERVIYSAN---RSEVNDVQAFDAIPPEFLQ 623
Query: 62 NTKS*GMSNHKLLLKVDIPIMLLRN 86
+ K+ + NHKL LKV PIMLLR+
Sbjct: 624 SLKTSDLPNHKLTLKVGTPIMLLRD 698
>BM814840 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (6%)
Length = 733
Score = 45.4 bits (106), Expect = 2e-05
Identities = 24/54 (44%), Positives = 33/54 (60%)
Frame = +3
Query: 95 NGTRLIVVELGERYIGATVITGTNANDKVHISIMDLVPSDPNFQLNSEEDSFQL 148
+GTRLI+V LG+ I A VI GT+A + +I M+L+PS N + E F L
Sbjct: 27 HGTRLIIVSLGKNVICARVIGGTHAGEVSYIPRMNLIPSGANVSITFERCQFPL 188
>TC80784 similar to PIR|T50352|T50352 ornithine--oxo-acid transaminase (EC
2.6.1.13) [imported] - fission yeast, partial (36%)
Length = 1206
Score = 28.9 bits (63), Expect = 1.9
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = -3
Query: 70 NHKLLLKVDIPIMLLRNIDQAAGLC 94
N++ LL +D+PI L I + +GLC
Sbjct: 946 NYRFLLLIDLPISLFSGIAKCSGLC 872
>TC82071 weakly similar to PIR|T00411|T01608 hypothetical protein At2g44820
[imported] - Arabidopsis thaliana, partial (36%)
Length = 764
Score = 27.3 bits (59), Expect = 5.5
Identities = 13/45 (28%), Positives = 26/45 (56%), Gaps = 4/45 (8%)
Frame = -3
Query: 201 SQANNRPVL*M*----CLRKFLEMFDVYNTISFIYFIFLLIYYYT 241
++ NN P+ + C + E+ DV++ + ++FIFLLI ++
Sbjct: 297 TKGNNLPIFNLFALFPCHMRVAEVLDVFHNLPDVHFIFLLIVLFS 163
>BQ143841
Length = 562
Score = 26.6 bits (57), Expect = 9.3
Identities = 11/31 (35%), Positives = 19/31 (60%), Gaps = 2/31 (6%)
Frame = +2
Query: 214 LRKFLEMFDVYNTISFIYFIFLLIY--YYTI 242
+ K++ +Y + FIY +F++IY YY I
Sbjct: 317 IAKYIYYISIYLYVLFIYILFIIIYIVYYYI 409
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.341 0.150 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,952,070
Number of Sequences: 36976
Number of extensions: 84366
Number of successful extensions: 801
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 792
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 798
length of query: 242
length of database: 9,014,727
effective HSP length: 93
effective length of query: 149
effective length of database: 5,575,959
effective search space: 830817891
effective search space used: 830817891
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0197.8


