

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0197.5
(317 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
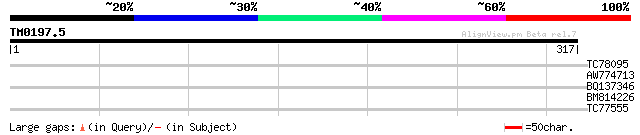
TC78095 weakly similar to PIR|G84776|G84776 hypothetical protein... 31 0.72
AW774713 weakly similar to PIR|A86340|A86 protein F2D10.24 [impo... 30 0.94
BQ137346 homologue to GP|15027095|em hypothetical protein LT.09 ... 30 1.6
BM814226 similar to GP|2094888|pdb|2 Cucumber Basic Protein A B... 29 2.1
TC77555 similar to GP|20260474|gb|AAM13135.1 26S proteasome subu... 28 3.6
>TC78095 weakly similar to PIR|G84776|G84776 hypothetical protein At2g36090
[imported] - Arabidopsis thaliana, partial (19%)
Length = 1467
Score = 30.8 bits (68), Expect = 0.72
Identities = 19/59 (32%), Positives = 28/59 (47%)
Frame = +3
Query: 192 GWVHNWLVGSLRMGTRWVVQINNVWLRSEQVCGDTQKAAKEDSWGASDASLQQVMVGEH 250
G V +LVG++ G RW V I+ R E+ CG ++ +E +MVG H
Sbjct: 996 GIVWGFLVGAMEKGERWKVDIDREKKRFEEFCG-RKRERREKKLRRERVMDMIIMVGCH 1169
>AW774713 weakly similar to PIR|A86340|A86 protein F2D10.24 [imported] -
Arabidopsis thaliana, partial (4%)
Length = 674
Score = 30.4 bits (67), Expect = 0.94
Identities = 19/51 (37%), Positives = 29/51 (56%), Gaps = 1/51 (1%)
Frame = +3
Query: 77 STARKPETVVGESSSERDQYRPESDWN-RSSVSRKANIAREVSSPVAVEQV 126
+T PE+V+GE S+ + P S W S S+K NI +SS +A E++
Sbjct: 231 ATPSMPESVIGEKSTSSEALFPASSWYFTSKESKKNNI---LSSNLAFEEL 374
>BQ137346 homologue to GP|15027095|em hypothetical protein LT.09 {Leishmania
major}, partial (0%)
Length = 1101
Score = 29.6 bits (65), Expect = 1.6
Identities = 29/120 (24%), Positives = 41/120 (34%), Gaps = 7/120 (5%)
Frame = +3
Query: 72 FSNGVSTARKPETVVGESSSERDQYRPESDWNRSSVSRKANIAREVSSPVAVEQVAGSKP 131
FS R+P G ++++R P + NR R + + P Q AG KP
Sbjct: 75 FSRQKEQKREPHRGGGRTTNQRS---PRAAGNRHEEGRPNTTKDKPAKP----QRAGPKP 233
Query: 132 QAENANHEEK-------PLDLGRMSHLVCGESKGATEEAMSVTAEARHVVKNADVGLMRK 184
+ ENA K G H +K T + T R +N G K
Sbjct: 234 KRENAKDAHKRQAPTTNKFSAGATQHPPARHTKNNTPKTQKQTTHDRQTTQNKSRGKRAK 413
>BM814226 similar to GP|2094888|pdb|2 Cucumber Basic Protein A Blue Copper
Protein, partial (46%)
Length = 591
Score = 29.3 bits (64), Expect = 2.1
Identities = 18/53 (33%), Positives = 26/53 (48%), Gaps = 1/53 (1%)
Frame = +3
Query: 265 VVDARNTNIQDGFRSGIAVDDDVSQVCVTPQEGSK-RRRGKPKGAAIRRPNMF 316
+VD N + G + +AV +V C TP+ GSK R GK + + N F
Sbjct: 15 LVDTLVFNYRQGTHNVVAVTKEVYDKCSTPRRGSKVYRSGKDRVRLAKGQNYF 173
>TC77555 similar to GP|20260474|gb|AAM13135.1 26S proteasome subunit-like
protein {Arabidopsis thaliana}, complete
Length = 1665
Score = 28.5 bits (62), Expect = 3.6
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = -3
Query: 269 RNTNIQDGFRSGIAVDDDVSQVCVTPQEGSKRRR 302
R I D +SG++ D + VC P + SKR R
Sbjct: 1252 REQKIHDPIKSGVSASMDNNAVCTFPVQPSKRSR 1151
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.131 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,930,417
Number of Sequences: 36976
Number of extensions: 89161
Number of successful extensions: 455
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 448
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 455
length of query: 317
length of database: 9,014,727
effective HSP length: 96
effective length of query: 221
effective length of database: 5,465,031
effective search space: 1207771851
effective search space used: 1207771851
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0197.5


