

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0195a.11
(377 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
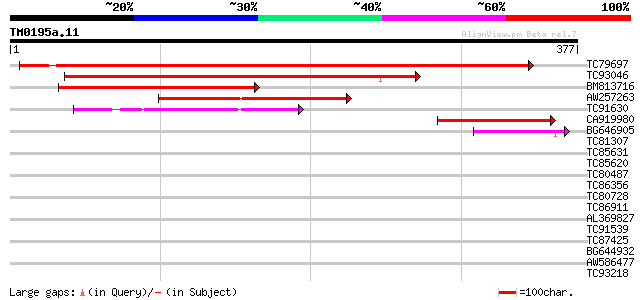
Score E
Sequences producing significant alignments: (bits) Value
TC79697 weakly similar to GP|2160150|gb|AAB60772.1| EST gb|T4382... 355 2e-98
TC93046 similar to GP|11320840|dbj|BAB18323. hypothetical protei... 308 3e-84
BM813716 similar to GP|11320840|dbj hypothetical protein {Oryza ... 224 6e-59
AW257263 similar to GP|9758227|dbj contains similarity to unknow... 110 9e-25
TC91630 similar to GP|9758227|dbj|BAB08726.1 contains similarity... 102 3e-22
CA919980 weakly similar to GP|11320840|dbj hypothetical protein ... 92 2e-19
BG646905 weakly similar to GP|20804419|dbj contains EST C73272(E... 45 6e-05
TC81307 weakly similar to GP|4335772|gb|AAD17449.1| unknown prot... 35 0.062
TC85631 homologue to SP|Q9BBN9|NU1C_LOTJA NADH-plastoquinone oxi... 32 0.52
TC85620 similar to SP|O22591|PSBQ_ONOVI Oxygen-evolving enhancer... 30 1.5
TC80487 30 2.0
TC86356 similar to GP|18654278|gb|AAL77575.1 dehydroquinate synt... 30 2.0
TC80728 similar to GP|15810469|gb|AAL07122.1 unknown protein {Ar... 28 4.4
TC86911 homologue to GP|20339362|gb|AAM19354.1 ribulose-5-phosph... 28 4.4
AL369827 weakly similar to PIR|T48594|T485 hypothetical protein ... 28 5.8
TC91539 28 5.8
TC87425 weakly similar to GP|20161478|dbj|BAB90402. contains EST... 28 5.8
BG644932 28 5.8
AW586477 28 7.6
TC93218 similar to PIR|G86185|G86185 hypothetical protein [impor... 28 7.6
>TC79697 weakly similar to GP|2160150|gb|AAB60772.1| EST gb|T43829 comes
from this gene. {Arabidopsis thaliana}, partial (55%)
Length = 1086
Score = 355 bits (911), Expect = 2e-98
Identities = 176/343 (51%), Positives = 234/343 (67%), Gaps = 1/343 (0%)
Frame = +3
Query: 7 KPEIALVIDNTVSTIISREPPSFIVARRLLASLNSVLTKLHAGYFRISLSLGGQALLWKT 66
KP I +++ T + + +P + + + LTK+HAGYF I +S G QALLWK+
Sbjct: 69 KPTIDIIVCATTNNTTNAKP----ITKPSHSQSQPFLTKIHAGYFFICVSFGAQALLWKS 236
Query: 67 LIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHV 126
L ++ TL H + +PS A+L+LW LA+ A LS LY+L+ + HFN V EF HH+
Sbjct: 237 LSEHNNESQTLWHGFNFMPSVAYLLLWCLAVLIAATLSFLYMLKSILHFNAVNDEFAHHI 416
Query: 127 GVNYLFAPWISWFLLLQ-SAPFVAPKTATYLVLWWVFAVPVVVLDVKIYGQWFTKGKRFL 185
GVNY++ PWIS+ L+LQ S P++ +T Y L F+ + +LDVK++GQWFT KRFL
Sbjct: 417 GVNYMYTPWISYLLMLQASPPWIVSRTCYYEFLCLAFSFVIFLLDVKLFGQWFTTEKRFL 596
Query: 186 STAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYLVLFVTLYQRLSGGNRLP 245
S ANP + +SVIGNLV AQ +GW E A+ ++SLGMVHYL+LFVTLYQRL+ N+ P
Sbjct: 597 SVVANPVNLVSVIGNLVAAQVMTEIGWNEIAISMYSLGMVHYLILFVTLYQRLTSSNQFP 776
Query: 246 VLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSMKRF 305
V+LRP FL FAAP +ASLAW SI G F KMLFFLSLFLF+S CRP LF+++MKR
Sbjct: 777 VVLRPANFLVFAAPSMASLAWKSISGAFLIS*KMLFFLSLFLFISQACRPALFKKTMKRL 956
Query: 306 NVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVL 348
NV WW YSFP+T L +A +YA EVK +++ LMLV+ +SVL
Sbjct: 957 NVTWWIYSFPLTFLGLACAEYAHEVKSSMASALMLVICIVSVL 1085
>TC93046 similar to GP|11320840|dbj|BAB18323. hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (37%)
Length = 829
Score = 308 bits (788), Expect = 3e-84
Identities = 153/239 (64%), Positives = 179/239 (74%), Gaps = 2/239 (0%)
Frame = +1
Query: 37 ASLNSVLTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLA 96
+SL+ +LTK+HAGYFRISLSL QALLWK LI P D LRH+ M+PS+AF +LWSLA
Sbjct: 97 SSLSIILTKIHAGYFRISLSLSVQALLWKILIEPIKDAHILRHIFTMIPSTAFTLLWSLA 276
Query: 97 LFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYL 156
LFTL LS LYLL+CL HF+ VK EF + +GVNY+FAPWISW LLLQS+P V P Y
Sbjct: 277 LFTLLTLSFLYLLKCLLHFDKVKEEFFNQIGVNYMFAPWISWLLLLQSSPIVPPAALHYK 456
Query: 157 VLWWVFAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESA 216
+LW +F VPVV+LDVKIYGQWFTKGK FLS ANPTSQ+SVIGNLV A AAA MGWKESA
Sbjct: 457 ILWLLFVVPVVILDVKIYGQWFTKGKMFLSMVANPTSQMSVIGNLVAALAAAQMGWKESA 636
Query: 217 VCLFSLGMVHYLVLFVTLYQRLSGGNRLP--VLLRPVFFLFFAAPGVASLAWGSIVGGF 273
+C FSLG+ HYLVLFVTLYQRL G N++P V R V AS +G++ G+
Sbjct: 637 ICFFSLGIAHYLVLFVTLYQRLPGNNKIPGDVATRGVLSCSLLLLVWASFGFGNLFCGY 813
>BM813716 similar to GP|11320840|dbj hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (17%)
Length = 469
Score = 224 bits (570), Expect = 6e-59
Identities = 106/134 (79%), Positives = 116/134 (86%)
Frame = +1
Query: 33 RRLLASLNSVLTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVL 92
R L+ S+ VL KLHAGYFRISLS GGQALLWKTLI PT+D S RHV+ M+PSS F+VL
Sbjct: 67 RSLIRSITCVLKKLHAGYFRISLSFGGQALLWKTLIDPTNDTSKSRHVLSMLPSSVFIVL 246
Query: 93 WSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKT 152
WS++LF LALLSLLYLLRCLF F MVK EFLHHVGVNYLFAPWISWFLLLQSAPF+APKT
Sbjct: 247 WSMSLFILALLSLLYLLRCLFFFKMVKEEFLHHVGVNYLFAPWISWFLLLQSAPFIAPKT 426
Query: 153 ATYLVLWWVFAVPV 166
TYL+LWWVF VPV
Sbjct: 427 ITYLILWWVFTVPV 468
>AW257263 similar to GP|9758227|dbj contains similarity to unknown
protein~gene_id:MZF18.9~pir||T02898 {Arabidopsis
thaliana}, partial (20%)
Length = 405
Score = 110 bits (275), Expect = 9e-25
Identities = 51/128 (39%), Positives = 78/128 (60%)
Frame = +1
Query: 100 LALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLW 159
+A + +Y+L+ L +F V+ E+ H + VN+ FAPWI+ L P K + LW
Sbjct: 16 IATIGAVYILKLLSYFEAVRREYYHPIRVNFFFAPWIALLFLALGVPPSVTKNL-HQSLW 192
Query: 160 WVFAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCL 219
++ P++ L++KIYGQW + G+R LS ANP++ LSV+GN VGA A MG E +
Sbjct: 193 YILMAPILFLELKIYGQWMSGGQRRLSKVANPSNHLSVVGNFVGALLGASMGLIEGPIFF 372
Query: 220 FSLGMVHY 227
F++G+ HY
Sbjct: 373 FAVGLAHY 396
>TC91630 similar to GP|9758227|dbj|BAB08726.1 contains similarity to unknown
protein~gene_id:MZF18.9~pir||T02898 {Arabidopsis
thaliana}, partial (31%)
Length = 1346
Score = 102 bits (253), Expect = 3e-22
Identities = 54/153 (35%), Positives = 83/153 (53%)
Frame = +3
Query: 43 LTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLAL 102
L + F I L + QA+LWKTL S +H+ P L+LW ++ +A
Sbjct: 909 LLRFPVSSFGICLGVSSQAILWKTLA-----TSPSTEFLHISPKIN-LILWYISTILIAT 1070
Query: 103 LSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWWVF 162
+ +Y+L+ L +F V+ E+ H + VN+ FAPWI+ L P K + LW++
Sbjct: 1071 VFAVYILKLLLYFEAVRREYYHPIRVNFFFAPWIALLFLALGVPPSVTKN-LHQSLWYIL 1247
Query: 163 AVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQL 195
VP+ L++KIYGQW + G+R LS ANP++ L
Sbjct: 1248 MVPIFFLELKIYGQWMSGGQRXLSKVANPSNHL 1346
>CA919980 weakly similar to GP|11320840|dbj hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (18%)
Length = 669
Score = 92.4 bits (228), Expect = 2e-19
Identities = 43/79 (54%), Positives = 60/79 (75%)
Frame = -2
Query: 285 LFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLA 344
+FL + V RP LF++SM++F+VAWWAYSFP+T LA+AS YA EVKG ++HV+ML L
Sbjct: 329 VFLNNNQVSRPLLFKKSMRKFSVAWWAYSFPLTALAIASAQYAHEVKGIMAHVIMLFLSL 150
Query: 345 LSVLVSLALTVFTFINSKM 363
+SVLV + L + T +N +M
Sbjct: 149 ISVLVCIMLMIVTALNIRM 93
>BG646905 weakly similar to GP|20804419|dbj contains EST
C73272(E3606)~similar to Arabidopsis thaliana chromosome
5 MZF18.9~unknown protein, partial (10%)
Length = 735
Score = 44.7 bits (104), Expect = 6e-05
Identities = 22/67 (32%), Positives = 39/67 (57%), Gaps = 3/67 (4%)
Frame = +2
Query: 309 WWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVLVSLALTVFTFINS---KMLL 365
WWAY+FP+T A+A+ Y+ +V ++ L + L +S +AL + T +++ + L
Sbjct: 14 WWAYTFPMTGAAIATIRYSNQVPNIVTKSLCIALALISTFTVIALLLSTILHAFVFRDLF 193
Query: 366 PDDDPIA 372
P+D IA
Sbjct: 194 PNDIAIA 214
>TC81307 weakly similar to GP|4335772|gb|AAD17449.1| unknown protein
{Arabidopsis thaliana}, partial (31%)
Length = 1171
Score = 34.7 bits (78), Expect = 0.062
Identities = 32/117 (27%), Positives = 58/117 (49%), Gaps = 1/117 (0%)
Frame = +1
Query: 244 LPVLLRPVFFLFF-AAPGVASLAWGSIVGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSM 302
LP+LL P+FFLFF + +A++ G F K+L ++ L L T +
Sbjct: 361 LPLLLLPLFFLFFLSLCSLATITHSVFHGFFGRPVKLLSTVTSLLSSFLPLLTTTILSHL 540
Query: 303 KRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVLVSLALTVFTFI 359
F + S P+ +L + S ++E ++ T+S +L+L L L + ++ T+ + I
Sbjct: 541 ILFTI-----SLPIPLLRL-SFSFSEIIR-TVSAILVLALSLLIFYIRISWTLSSVI 690
>TC85631 homologue to SP|Q9BBN9|NU1C_LOTJA NADH-plastoquinone oxidoreductase
chain 1 chloroplast (EC 1.6.5.3). {Lotus japonicus},
complete
Length = 3706
Score = 31.6 bits (70), Expect = 0.52
Identities = 28/90 (31%), Positives = 44/90 (48%), Gaps = 1/90 (1%)
Frame = +3
Query: 230 LFVTLYQRLSGGN-RLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLF 288
LFVT+ L G N +P + FF GV +G+ + F TL+K FFL F
Sbjct: 2787 LFVTVLY-LGGSNISIPYIFVSEFFEINKTYGV----FGTTIDLFITLAKTYFFL----F 2939
Query: 289 MSLVCRPTLFRRSMKRFNVAWWAYSFPVTV 318
+S++ R +L R M + W + P+++
Sbjct: 2940 VSIITRWSLPRLRMDQLLNLGWKFLLPISL 3029
>TC85620 similar to SP|O22591|PSBQ_ONOVI Oxygen-evolving enhancer protein 3
chloroplast precursor (OEE3), partial (84%)
Length = 1005
Score = 30.0 bits (66), Expect = 1.5
Identities = 30/112 (26%), Positives = 46/112 (40%), Gaps = 20/112 (17%)
Frame = -2
Query: 233 TLYQRLSGGNRLPVLLRPVF---------------FLFFAAPGVASLAWGSIVGGFDTLS 277
+L+++ + P+LLR F L FAA S++W S F S
Sbjct: 815 SLHKKARNHSY*PILLRTSFRVDTAIA*YFSASEGLLTFAA*SRLSMSWKS----FPVSS 648
Query: 278 KMLFFLSLFLFMSLVCR-----PTLFRRSMKRFNVAWWAYSFPVTVLAMAST 324
FF S +V R P L RRS R+ A+W+ +F ++ A +
Sbjct: 647 LRDFFWSFGFAEMMVLRSYRR*PALRRRSF*RYGQAFWSMNFLAATISFADS 492
>TC80487
Length = 817
Score = 29.6 bits (65), Expect = 2.0
Identities = 14/61 (22%), Positives = 27/61 (43%)
Frame = -2
Query: 95 LALFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTAT 154
L + +++L+ C F + + +H+ V + SW+ +Q PF AP T
Sbjct: 693 LETLNIRTMNILFRSPCFFTLHSIILNPFNHISVGFCLPTRCSWWR*IQR*PF*APLVTT 514
Query: 155 Y 155
+
Sbjct: 513 H 511
>TC86356 similar to GP|18654278|gb|AAL77575.1 dehydroquinate synthase
{Lycopersicon esculentum}, partial (43%)
Length = 881
Score = 29.6 bits (65), Expect = 2.0
Identities = 17/51 (33%), Positives = 24/51 (46%)
Frame = +3
Query: 115 FNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWWVFAVP 165
F + K+ FLH +G Y P SW Q + V P T ++W + A P
Sbjct: 204 FFLKKSLFLHLLGHQYALVPLNSWTPFQQKSNLVFPPLLT--LIWVIGATP 350
>TC80728 similar to GP|15810469|gb|AAL07122.1 unknown protein {Arabidopsis
thaliana}, partial (87%)
Length = 1523
Score = 28.5 bits (62), Expect = 4.4
Identities = 26/104 (25%), Positives = 45/104 (43%), Gaps = 13/104 (12%)
Frame = +2
Query: 267 GSIVGGFDTLSKMLFFLSLFLFMSLVCRP--------TLFRRSMKR-----FNVAWWAYS 313
G +GG+ + + LS F+ MSL+ P +L ++ +K+ FN +W +
Sbjct: 461 GLTIGGYVAVVLAIVSLSPFVLMSLIAIPKINPHKWLSLGQKGVKKDWNLFFNTLFWNLN 640
Query: 314 FPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVLVSLALTVFT 357
F V +A EE K T L++ ++ V + L T
Sbjct: 641 FWDNVSTLAGE--VEEPKKTFPLALLIAVIFTCVSYLIPLFAVT 766
>TC86911 homologue to GP|20339362|gb|AAM19354.1
ribulose-5-phosphate-3-epimerase {Pisum sativum},
complete
Length = 1233
Score = 28.5 bits (62), Expect = 4.4
Identities = 25/96 (26%), Positives = 42/96 (43%), Gaps = 20/96 (20%)
Frame = +2
Query: 36 LASLNSVLTKLHAGYFRISLSLGGQALLWKTL-IGPTHDKSTLRHVVH------------ 82
L S++ T++ + +SLS GG+ LW L + T + + +H
Sbjct: 83 LRSMDFPFTRILSSIPVLSLSPGGKFQLWLRLHLVLTSFQKAISLFLHLFFLPTLQNWEN 262
Query: 83 -------MVPSSAFLVLWSLALFTLALLSLLYLLRC 111
+V L+LW +AL + LL LL+L+ C
Sbjct: 263 R*KQ*SWLVVIGFMLMLWMVALSQILLLDLLWLMHC 370
>AL369827 weakly similar to PIR|T48594|T485 hypothetical protein T22N19.120 -
Arabidopsis thaliana, partial (19%)
Length = 527
Score = 28.1 bits (61), Expect = 5.8
Identities = 13/21 (61%), Positives = 17/21 (80%)
Frame = -2
Query: 273 FDTLSKMLFFLSLFLFMSLVC 293
FD ++KMLFFL LFL S++C
Sbjct: 79 FDFINKMLFFL-LFLLFSILC 20
>TC91539
Length = 1095
Score = 28.1 bits (61), Expect = 5.8
Identities = 17/46 (36%), Positives = 27/46 (57%)
Frame = -3
Query: 331 KGTISHVLMLVLLALSVLVSLALTVFTFINSKMLLPDDDPIATFLI 376
K ++H+L LV L +LVSL+ + I SK L+ + I++F I
Sbjct: 163 KHQMNHLLHLVAPILGILVSLSCHWSSCIQSKFLVDEPVAISSFSI 26
>TC87425 weakly similar to GP|20161478|dbj|BAB90402. contains ESTs
C74501(E31745) AU094804(E31745)~unknown protein, partial
(24%)
Length = 929
Score = 28.1 bits (61), Expect = 5.8
Identities = 15/47 (31%), Positives = 26/47 (54%), Gaps = 5/47 (10%)
Frame = +2
Query: 77 LRHVVHMVPSSAFLVLWSLALF-----TLALLSLLYLLRCLFHFNMV 118
+R + +PSS+FLV+ L LF LA+ +++ FH +M+
Sbjct: 788 IRTYITTLPSSSFLVMCYLFLFFLPNYILAISVIIFYFNMTFHLSMI 928
>BG644932
Length = 641
Score = 28.1 bits (61), Expect = 5.8
Identities = 20/60 (33%), Positives = 29/60 (48%), Gaps = 3/60 (5%)
Frame = +3
Query: 59 GQALLWKTL---IGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLRCLFHF 115
G L +KTL + H+ TL H + + F + W L+ + L SLLY +CL F
Sbjct: 120 GVLLGFKTLN*AVRNFHEIKTLNHTILL-----F*LFWILSAISPLLFSLLYETQCLLTF 284
>AW586477
Length = 522
Score = 27.7 bits (60), Expect = 7.6
Identities = 18/59 (30%), Positives = 29/59 (48%), Gaps = 9/59 (15%)
Frame = -2
Query: 78 RHVVHMV-PSSAFL--------VLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHVG 127
+H +H+V P + FL VLW++ + T L+ L + L HF + +FL G
Sbjct: 506 KHGIHLVEPLNNFLSHFVA*NIVLWAITIQTSVTLNFLKIWNILHHFLLKFLKFLKPTG 330
>TC93218 similar to PIR|G86185|G86185 hypothetical protein [imported] -
Arabidopsis thaliana, partial (16%)
Length = 732
Score = 27.7 bits (60), Expect = 7.6
Identities = 12/31 (38%), Positives = 19/31 (60%)
Frame = -2
Query: 90 LVLWSLALFTLALLSLLYLLRCLFHFNMVKA 120
L+ S L ++ L S+L+LL+C HF+ A
Sbjct: 524 LIFHSFTLISILLKSVLFLLKCRRHFSFTVA 432
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.330 0.141 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,610,394
Number of Sequences: 36976
Number of extensions: 232473
Number of successful extensions: 1853
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 1819
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1845
length of query: 377
length of database: 9,014,727
effective HSP length: 98
effective length of query: 279
effective length of database: 5,391,079
effective search space: 1504111041
effective search space used: 1504111041
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0195a.11


