

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0193.7
(848 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
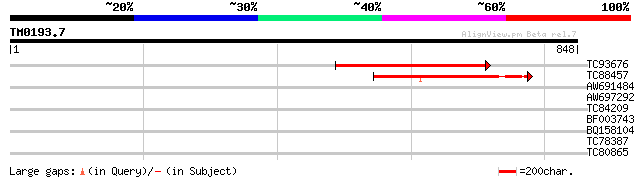
Score E
Sequences producing significant alignments: (bits) Value
TC93676 similar to GP|20977172|gb|AAM33305.1 glycogen branching ... 295 4e-80
TC88457 homologue to PIR|T06493|T06493 1 4-alpha-glucan branchin... 246 3e-65
AW691484 weakly similar to PIR|T48283|T48 ankyrin-like protein -... 33 0.46
AW697292 homologue to GP|22136708|gb putative isoamylase {Arabid... 32 1.3
TC84209 similar to GP|4204311|gb|AAD10692.1| lcl|prt_seq No defi... 31 2.3
BF003743 similar to GP|17016406|gb| wsv008 {shrimp white spot sy... 30 5.1
BQ158104 similar to GP|22758351|gb| hypothetical protein {Oryza ... 30 5.1
TC78387 similar to GP|22136492|gb|AAM91324.1 unknown protein {Ar... 30 5.1
TC80865 weakly similar to GP|13940211|emb|CAC37923. fructan 1-ex... 29 6.6
>TC93676 similar to GP|20977172|gb|AAM33305.1 glycogen branching enzyme
{Glomus intraradices}, partial (37%)
Length = 911
Score = 295 bits (756), Expect = 4e-80
Identities = 135/232 (58%), Positives = 176/232 (75%)
Frame = +2
Query: 488 GIGFDYRLAMAIPDKWIDYLKNKKDHEWSMKEISLSLTNRRYSEKCVSYAESHDQSIVGD 547
GIGFDYRL MAIPD WI LK K+D +W+M I +LTNRR+ EK ++YAESHDQ++VGD
Sbjct: 2 GIGFDYRLGMAIPDMWIKLLKEKRDDDWNMGNICWTLTNRRHKEKTIAYAESHDQALVGD 181
Query: 548 KTFSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMSLGGEGYLNFMGNEFGHPE 607
KT +F LMD+E+Y+ MS + +P I+RG+ALHKMI IT LGGEGYLNF GNEFGHPE
Sbjct: 182 KTIAFWLMDKEMYTNMSDITPLTPIIDRGLALHKMIRLITHGLGGEGYLNFEGNEFGHPE 361
Query: 608 WIDFPREGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSFLASTKQIVSST 667
W+DFPREGN SY RRQW++VD LRYK++N FDKAM ++K+ +L+S + VS
Sbjct: 362 WLDFPREGNNNSYHYARRQWNVVDDPLLRYKYLNEFDKAMQHSEEKYKWLSSPQAYVSLK 541
Query: 668 NEEDKVIVFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSDAREFGG 719
NE+ K+IVFER +L+++FNFHP +Y YK+G + GKY + L++D +FGG
Sbjct: 542 NEDHKLIVFERANLLWIFNFHPTNSYPDYKIGTEWAGKYSIVLNTDNPKFGG 697
>TC88457 homologue to PIR|T06493|T06493 1 4-alpha-glucan branching enzyme
(EC 2.4.1.18) I - garden pea, partial (27%)
Length = 1201
Score = 246 bits (627), Expect = 3e-65
Identities = 126/249 (50%), Positives = 163/249 (64%), Gaps = 11/249 (4%)
Frame = +3
Query: 545 VGDKTFSFLLMDEEIYSGMSCLADASPTIERGIALHKMIHFITMSLGGEGYLNFMGNEFG 604
VGDKT +F LMD+++Y M+ ++P I+RGIALHKMI ITM LGGEGYLNFMGNEFG
Sbjct: 3 VGDKTLAFWLMDKDMYDFMALDRPSTPLIDRGIALHKMIRLITMGLGGEGYLNFMGNEFG 182
Query: 605 HPEWIDFPR-----------EGNGWSYEKCRRQWSLVDTDHLRYKFMNAFDKAMNLLDDK 653
HPEWIDFPR GN SY+KCRR++ L D ++LRY M FD+AM L+++
Sbjct: 183 HPEWIDFPRGDQHLPNGTVVPGNNNSYDKCRRRFDLGDAEYLRYHGMQEFDRAMQHLEER 362
Query: 654 FSFLASTKQIVSSTNEEDKVIVFERGDLVFVFNFHPETTYEGYKVGCDLPGKYRVALDSD 713
+ F+ S Q +S NE D+VI+FER +LVFVFNFH +Y YKVGC PGKY++ LDSD
Sbjct: 363 YGFMISEHQYISRKNEGDRVIIFERDNLVFVFNFHWTNSYSDYKVGCLKPGKYKIVLDSD 542
Query: 714 AREFGGHGRVGHNVDHFTAPEGIPGVPESNFNNRPNSFKILSPPRTCVVYYRVDESQEEN 773
FGG R+ H ++FT+ E +++RP SF + +P RT VVY VD E
Sbjct: 543 ESLFGGFNRLNHTAEYFTS--------EGWYDDRPRSFLVYAPCRTAVVYALVD-GVESE 695
Query: 774 SISNLVGVQ 782
+ VGV+
Sbjct: 696 PVELSVGVE 722
>AW691484 weakly similar to PIR|T48283|T48 ankyrin-like protein - Arabidopsis
thaliana, partial (27%)
Length = 646
Score = 33.1 bits (74), Expect = 0.46
Identities = 16/43 (37%), Positives = 27/43 (62%)
Frame = -1
Query: 1 MITSFSLQSFNIASTAHNSRNKQDLAKQNSVELVLGYRNPKGC 43
+++SFSL+ S++H S+NK+ + + E+VLG N GC
Sbjct: 217 IVSSFSLRKSTKLSSSHFSQNKRSIFMKKI*EIVLGN*NRSGC 89
>AW697292 homologue to GP|22136708|gb putative isoamylase {Arabidopsis
thaliana}, partial (13%)
Length = 343
Score = 31.6 bits (70), Expect = 1.3
Identities = 12/24 (50%), Positives = 17/24 (70%)
Frame = +2
Query: 394 VLRFLLSNLRWWLEEFKFDGFRFD 417
V+ +L +LR W+ E+ DGFRFD
Sbjct: 65 VMDLILDSLRHWVTEYHVDGFRFD 136
>TC84209 similar to GP|4204311|gb|AAD10692.1| lcl|prt_seq No definition line
found {Arabidopsis thaliana}, partial (36%)
Length = 701
Score = 30.8 bits (68), Expect = 2.3
Identities = 29/109 (26%), Positives = 46/109 (41%), Gaps = 2/109 (1%)
Frame = +1
Query: 692 TYEGYKVGCDLPGKYRVALDSDAREFGGHGRVG--HNVDHFTAPEGIPGVPESNFNNRPN 749
T E + + P R L++ + GHG G H+ +H + P+ F P+
Sbjct: 37 TVESHHAEAEKPHSNRRRLEA----WLGHGGKGQ*HHSNHIS--------PQKKFKKPPS 180
Query: 750 SFKILSPPRTCVVYYRVDESQEENSISNLVGVQETSTAADIVANIPDGS 798
+FKILSP RT NSI+ + T+T + + N + S
Sbjct: 181 TFKILSPLRT------------TNSINYELSSMTTATLSALFRNFLNNS 291
>BF003743 similar to GP|17016406|gb| wsv008 {shrimp white spot syndrome
virus}, partial (16%)
Length = 598
Score = 29.6 bits (65), Expect = 5.1
Identities = 14/34 (41%), Positives = 22/34 (64%)
Frame = +3
Query: 623 CRRQWSLVDTDHLRYKFMNAFDKAMNLLDDKFSF 656
C R+WSL +T +L +KF + + KA +L +F F
Sbjct: 291 CSRKWSLYNTRNLTWKF-SLWKKAKKILSVQFKF 389
>BQ158104 similar to GP|22758351|gb| hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (9%)
Length = 993
Score = 29.6 bits (65), Expect = 5.1
Identities = 16/52 (30%), Positives = 28/52 (53%), Gaps = 1/52 (1%)
Frame = -1
Query: 235 PPLSERYQFKYPRPPKPKAPRIYEAH-VGMSSSEPRINSYKEFADDILPRIR 285
PP+ R+ + PP +Y AH +G +S+ PR S+ + ++ PR+R
Sbjct: 330 PPMLARFLLPHAVPPP-----LYRAHHLGGTSASPRPTSFLPWGINLYPRVR 190
>TC78387 similar to GP|22136492|gb|AAM91324.1 unknown protein {Arabidopsis
thaliana}, partial (58%)
Length = 2019
Score = 29.6 bits (65), Expect = 5.1
Identities = 20/57 (35%), Positives = 28/57 (49%), Gaps = 5/57 (8%)
Frame = +1
Query: 219 VDPTKFAAPYDGVY--WDPPLSERYQFKYPRPPKPKA---PRIYEAHVGMSSSEPRI 270
V+P K A GV+ W+ P + + PR K + P E H+G S+SEP I
Sbjct: 1813 VNPRKLAGGGTGVFLPWNVPSRKPTKHLPPRAQKGRLLALPSPVEPHIGESTSEPSI 1983
>TC80865 weakly similar to GP|13940211|emb|CAC37923. fructan 1-exohydrolase
IIb {Cichorium intybus}, partial (26%)
Length = 809
Score = 29.3 bits (64), Expect = 6.6
Identities = 29/125 (23%), Positives = 52/125 (41%), Gaps = 2/125 (1%)
Frame = +3
Query: 57 RVSTGFKGVAVITDNKSAMSATEEDLENIGILHIDPAIKPFKDHFKCRLKRYIDQKKLIE 116
R GFK + + +S++ E I IDP +K L+ ID + +IE
Sbjct: 285 RAGDGFKCLMISDQTRSSLREDVEKTSYATIFDIDPNLKTIS------LRSLID-RSIIE 443
Query: 117 EYEGGLEE--FAQGYLKFGFNREEGGIVYREWAPAAQEAQIIGDFNEWNGSNHPMEKNQF 174
+ G ++ Y F +++ V+ + ++ +I N W+ M++ QF
Sbjct: 444 SFGDGGRACITSRAYPLFATDKDAHLFVFND----GSQSVVISQLNAWS-----MKQAQF 596
Query: 175 GVWSI 179
G SI
Sbjct: 597 GTESI 611
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.136 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,006,568
Number of Sequences: 36976
Number of extensions: 410977
Number of successful extensions: 1935
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1861
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1923
length of query: 848
length of database: 9,014,727
effective HSP length: 104
effective length of query: 744
effective length of database: 5,169,223
effective search space: 3845901912
effective search space used: 3845901912
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0193.7


