

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
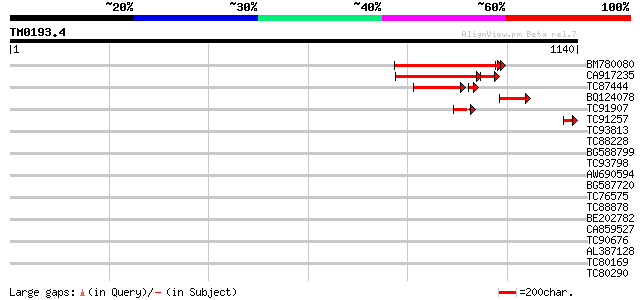
Query= TM0193.4
(1140 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BM780080 similar to GP|12322632|gb unknown protein 5' partial {... 395 e-110
CA917235 similar to GP|12322632|gb unknown protein 5' partial {... 301 4e-96
TC87444 similar to GP|21555695|gb|AAM63916.1 unknown {Arabidopsi... 202 6e-52
BQ124078 similar to GP|12322632|gb unknown protein 5' partial {... 99 1e-20
TC91907 49 1e-05
TC91257 similar to GP|21434|emb|CAA36616.1|| ORF4 {Solanum tuber... 44 4e-04
TC93813 similar to GP|11935197|gb|AAG42014.1 unknown protein {Ar... 34 0.37
TC88228 similar to GP|13605916|gb|AAK32943.1 AT4g00830/A_TM018A1... 33 0.82
BG588799 GP|945050|gb| transcription factor {Drosophila melanoga... 32 1.1
TC93798 similar to GP|11994748|dbj|BAB03077. gene_id:T5M7.2~unkn... 32 1.1
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 32 1.4
BG587720 homologue to GP|18075960|em putative metallophosphatase... 30 4.1
TC76575 homologue to SP|Q40465|IF4V_TOBAC Eukaryotic initiation ... 30 5.3
TC88878 similar to PIR|C96777|C96777 F25A4.24 [imported] - Arabi... 30 5.3
BE202782 weakly similar to GP|22093801|dbj hypothetical protein~... 30 5.3
CA859527 30 5.3
TC90676 similar to GP|6503308|gb|AAF14684.1| Contains PF|00646 F... 30 5.3
AL387128 similar to PIR|A36621|A366 NADH dehydrogenase (ubiquino... 29 9.0
TC80169 29 9.0
TC80290 weakly similar to GP|13937149|gb|AAK50068.1 At1g79380/T8... 29 9.0
>BM780080 similar to GP|12322632|gb unknown protein 5' partial {Arabidopsis
thaliana}, partial (42%)
Length = 672
Score = 395 bits (1016), Expect = e-110
Identities = 190/217 (87%), Positives = 197/217 (90%), Gaps = 1/217 (0%)
Frame = +1
Query: 775 PGLLARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQ 834
PGLLARYMKEVHAPILSIWGVKI VIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQ
Sbjct: 1 PGLLARYMKEVHAPILSIWGVKIAVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQ 180
Query: 835 GYFNNVSEYLRIGPPLYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETS 894
GYFNNV+EYLRIGPPLYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEI+KA+LVP+TS
Sbjct: 181 GYFNNVTEYLRIGPPLYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEIAKAALVPDTS 360
Query: 895 YIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPCCAPEDDSCVS-GACKDCT 953
YIAKPAASWLDDFLVW+SPEAFGCCRKFTNGSYCPPDDQPPCCAPEDDSCVS G C DCT
Sbjct: 361 YIAKPAASWLDDFLVWVSPEAFGCCRKFTNGSYCPPDDQPPCCAPEDDSCVSVGVCNDCT 540
Query: 954 TCFRHSDLRNDRTSTMQFRDKLPWFLSALPSADCAKG 990
TCFRHSDL +DR FR+KLPW P KG
Sbjct: 541 TCFRHSDLSHDRPFHNTFREKLPWVS*CSPIC*LCKG 651
Score = 44.3 bits (103), Expect = 3e-04
Identities = 19/19 (100%), Positives = 19/19 (100%)
Frame = +2
Query: 978 FLSALPSADCAKGGHGAYT 996
FLSALPSADCAKGGHGAYT
Sbjct: 614 FLSALPSADCAKGGHGAYT 670
>CA917235 similar to GP|12322632|gb unknown protein 5' partial {Arabidopsis
thaliana}, partial (37%)
Length = 637
Score = 301 bits (770), Expect(2) = 4e-96
Identities = 141/172 (81%), Positives = 154/172 (88%)
Frame = +1
Query: 776 GLLARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQG 835
GLL RYMKEVHAP L +WGVKI+VIAIF AF LASIAL TRIEPGLEQ+I LPRDSYLQG
Sbjct: 7 GLLTRYMKEVHAPFLGLWGVKILVIAIFGAFTLASIALCTRIEPGLEQQIALPRDSYLQG 186
Query: 836 YFNNVSEYLRIGPPLYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSY 895
YF+N+SEYLR+GPPLYFVVK+YNYS ES HTNQLCSIS C+S+SLLNEIS+ASLVPE+SY
Sbjct: 187 YFSNISEYLRVGPPLYFVVKDYNYSLESKHTNQLCSISHCDSNSLLNEISRASLVPESSY 366
Query: 896 IAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPCCAPEDDSCVSG 947
IAKPAASWLDDFLVWISPEAF CCRKFTN SYCPPDDQPPCC ++ C G
Sbjct: 367 IAKPAASWLDDFLVWISPEAFSCCRKFTNDSYCPPDDQPPCCFLDEGPCGLG 522
Score = 70.5 bits (171), Expect(2) = 4e-96
Identities = 30/38 (78%), Positives = 32/38 (83%)
Frame = +2
Query: 947 GACKDCTTCFRHSDLRNDRTSTMQFRDKLPWFLSALPS 984
G CKDCTTCFRHSDL NDR ST QF++KLPWFL AL S
Sbjct: 524 GVCKDCTTCFRHSDLVNDRPSTAQFKEKLPWFLDALSS 637
>TC87444 similar to GP|21555695|gb|AAM63916.1 unknown {Arabidopsis
thaliana}, partial (68%)
Length = 1607
Score = 202 bits (514), Expect = 6e-52
Identities = 97/104 (93%), Positives = 102/104 (97%)
Frame = +3
Query: 812 ALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYFVVKNYNYSSESTHTNQLCS 871
ALSTRIEPGLEQEIVLPRDSYLQGYFNNV+EYLRIGPPLYFVVKNYNYSSESTHTNQLCS
Sbjct: 54 ALSTRIEPGLEQEIVLPRDSYLQGYFNNVTEYLRIGPPLYFVVKNYNYSSESTHTNQLCS 233
Query: 872 ISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWISPEA 915
ISQCNSDSLLNEI+KA+LVP+TSYIAKPAASWLDDFLVW+S A
Sbjct: 234 ISQCNSDSLLNEIAKAALVPDTSYIAKPAASWLDDFLVWVSSGA 365
Score = 46.2 bits (108), Expect = 7e-05
Identities = 17/19 (89%), Positives = 17/19 (89%)
Frame = -2
Query: 923 TNGSYCPPDDQPPCCAPED 941
T SYCPPDDQPPCCAPED
Sbjct: 409 TERSYCPPDDQPPCCAPED 353
>BQ124078 similar to GP|12322632|gb unknown protein 5' partial {Arabidopsis
thaliana}, partial (31%)
Length = 529
Score = 98.6 bits (244), Expect = 1e-20
Identities = 45/62 (72%), Positives = 54/62 (86%)
Frame = +2
Query: 986 DCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDYVNSMRAAREFSSKVSDSL 1045
DCAKG GAYT+S+DL GY+ G+IQAS FRTYHTPLN+Q DYVN++RAAREF SK+S SL
Sbjct: 2 DCAKGVPGAYTNSIDLSGYEGGVIQASEFRTYHTPLNRQGDYVNAIRAAREFCSKISASL 181
Query: 1046 KV 1047
K+
Sbjct: 182 KM 187
>TC91907
Length = 166
Score = 48.5 bits (114), Expect = 1e-05
Identities = 22/45 (48%), Positives = 28/45 (61%), Gaps = 1/45 (2%)
Frame = -3
Query: 892 ETSYIAKPAASWLDDFLVWISPE-AFGCCRKFTNGSYCPPDDQPP 935
E SYI+ PAASW+DD+ +WI+P+ CC NG C D PP
Sbjct: 164 EVSYISSPAASWIDDYFLWINPDLGDSCC--VDNGKACFADRNPP 36
>TC91257 similar to GP|21434|emb|CAA36616.1|| ORF4 {Solanum tuberosum},
partial (8%)
Length = 854
Score = 43.9 bits (102), Expect = 4e-04
Identities = 21/28 (75%), Positives = 24/28 (85%)
Frame = -1
Query: 1113 LPVVLSIFGPPSRCTIIEQEENRSSTSS 1140
L VVLS+FGPPSRCT +Q E+RSSTSS
Sbjct: 410 LQVVLSMFGPPSRCTNTDQGEDRSSTSS 327
>TC93813 similar to GP|11935197|gb|AAG42014.1 unknown protein {Arabidopsis
thaliana}, partial (70%)
Length = 664
Score = 33.9 bits (76), Expect = 0.37
Identities = 24/95 (25%), Positives = 38/95 (39%), Gaps = 5/95 (5%)
Frame = +1
Query: 919 CRKFTNGSYCPPDDQP-----PCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQFRD 973
CR+ SY PP D+P C + +SC++G C R+ + R+
Sbjct: 208 CRQPLPESYAPPADEPWMTGIFACVEDRESCLTGLFCPCVLFGRNVE---------SLRE 360
Query: 974 KLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGI 1008
PW + A +GG ++V + SGI
Sbjct: 361 NTPWTTPCICHAIFVEGGISVAIATVIATSFISGI 465
>TC88228 similar to GP|13605916|gb|AAK32943.1 AT4g00830/A_TM018A10_14
{Arabidopsis thaliana}, partial (67%)
Length = 1712
Score = 32.7 bits (73), Expect = 0.82
Identities = 21/63 (33%), Positives = 34/63 (53%), Gaps = 3/63 (4%)
Frame = -1
Query: 157 RKAAPNSPGSPYAIMFRPNATKSSGMKPMNVSAYSCSDT---SLGCSCGDCPSSSVCSNS 213
R +S +P+++ F P+ S VS+YSCS T + S PSSS+CS++
Sbjct: 404 RAGLSSSSSAPWSL-FSPSGNWSF*PASAAVSSYSCSSTLPSAFSSSPPSTPSSSICSST 228
Query: 214 AST 216
+S+
Sbjct: 227 SSS 219
>BG588799 GP|945050|gb| transcription factor {Drosophila melanogaster},
partial (0%)
Length = 781
Score = 32.3 bits (72), Expect = 1.1
Identities = 19/70 (27%), Positives = 32/70 (45%)
Frame = +3
Query: 182 MKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVDFILA 241
+ P+ + Y+ S T S C SS+ +S+++ CS+ SL +DF+L
Sbjct: 195 INPLYIYLYNSSITLTKMSIRSCAMSSLILHSSNSISLIGGICSVLFCSLCYPYLDFVLR 374
Query: 242 VLYIILICVF 251
+ IC F
Sbjct: 375 HSSLFFICTF 404
>TC93798 similar to GP|11994748|dbj|BAB03077. gene_id:T5M7.2~unknown protein
{Arabidopsis thaliana}, partial (15%)
Length = 991
Score = 32.3 bits (72), Expect = 1.1
Identities = 20/55 (36%), Positives = 27/55 (48%), Gaps = 2/55 (3%)
Frame = +3
Query: 179 SSGMKPMNVSAYSCSDTSLGCSCGDCPSS--SVCSNSASTTINKANSCSIKVGSL 231
SS + AY + + C DC S CS S T N+ NSCS++VGS+
Sbjct: 414 SSSSSSGGICAYCLKERLVKLVCSDCGEQRLSSCSCSDEITSNR-NSCSVEVGSV 575
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 32.0 bits (71), Expect = 1.4
Identities = 22/73 (30%), Positives = 40/73 (54%), Gaps = 2/73 (2%)
Frame = -3
Query: 155 IGRKAAP-NSPGSPYAIMFRPNATKSSGMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNS 213
+GRK + N+ GS ++ +++ S + S+ S S +S SC SSS S+S
Sbjct: 313 VGRKESICNNSGSGRVLILSISSSSPSSSSSSSSSSSSSSSSSSSSSCSSSSSSSSSSSS 134
Query: 214 AST-TINKANSCS 225
+S+ + + ++SCS
Sbjct: 133 SSSCSSSSSSSCS 95
Score = 29.6 bits (65), Expect = 6.9
Identities = 18/52 (34%), Positives = 30/52 (57%), Gaps = 1/52 (1%)
Frame = -3
Query: 175 NATKSSGMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNSASTTINKANS-CS 225
+++ SS + S+ S S +S SC SSS S+S+ST+ + ++S CS
Sbjct: 202 SSSSSSSSSSCSSSSSSSSSSSSSSSCSSSSSSSCSSSSSSTSPSSSSSLCS 47
Score = 29.3 bits (64), Expect = 9.0
Identities = 20/65 (30%), Positives = 35/65 (53%)
Frame = -3
Query: 159 AAPNSPGSPYAIMFRPNATKSSGMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNSASTTI 218
++P+S S + +++ SS + S+ S S +S CS SSS CS+S+S+T
Sbjct: 244 SSPSSSSSSSSSSSSSSSSSSSSSCSSSSSSSSSSSSSSSCSSS---SSSSCSSSSSSTS 74
Query: 219 NKANS 223
++S
Sbjct: 73 PSSSS 59
>BG587720 homologue to GP|18075960|em putative metallophosphatase {Lupinus
luteus}, partial (36%)
Length = 768
Score = 30.4 bits (67), Expect = 4.1
Identities = 15/43 (34%), Positives = 20/43 (45%)
Frame = -1
Query: 746 AEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAP 788
A D+R C PCI +F ++Q P L RY + AP
Sbjct: 351 AHDRRWQCPPCISATAFEGSLSPLVQQMTPLLHNRYKRSSQAP 223
>TC76575 homologue to SP|Q40465|IF4V_TOBAC Eukaryotic initiation factor
4A-11 (eIF-4A-11) (eIF4A-11). [Common tobacco]
{Nicotiana tabacum}, complete
Length = 1627
Score = 30.0 bits (66), Expect = 5.3
Identities = 14/28 (50%), Positives = 17/28 (60%), Gaps = 1/28 (3%)
Frame = -2
Query: 71 CLRNFLNLFCELTC-SPNQSLFINVTSV 97
C N LNL C+L C NQSL +N T +
Sbjct: 525 CSHNLLNLLCQLPCWGQNQSLALNQTII 442
>TC88878 similar to PIR|C96777|C96777 F25A4.24 [imported] - Arabidopsis
thaliana, partial (20%)
Length = 839
Score = 30.0 bits (66), Expect = 5.3
Identities = 14/45 (31%), Positives = 20/45 (44%)
Frame = +2
Query: 37 CPTITGNVCCTKAQFDTLQTQVQQAIPFLVGCPACLRNFLNLFCE 81
CP G+ CC Q +Q Q QQ C + L++ L C+
Sbjct: 248 CPYNNGSTCCNSIQDGQIQKQFQQMNVSDTACASLLKSILCARCD 382
>BE202782 weakly similar to GP|22093801|dbj hypothetical protein~predicted by
GlimmerM~similar to Arabidopsis thaliana chromosome3
At3g23330, partial (13%)
Length = 548
Score = 30.0 bits (66), Expect = 5.3
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = -3
Query: 165 GSPYAIMFRPNATKSSGMKPMNVSAYSCSDTS 196
G P ++ PN T + + P N+ + SCS TS
Sbjct: 129 GCPGLVLISPNTTANFSLCPHNICSLSCSSTS 34
>CA859527
Length = 501
Score = 30.0 bits (66), Expect = 5.3
Identities = 16/34 (47%), Positives = 20/34 (58%), Gaps = 3/34 (8%)
Frame = +3
Query: 197 LGCSCG---DCPSSSVCSNSASTTINKANSCSIK 227
+GC CG DC SS CSNS +NKA+ + K
Sbjct: 108 MGCPCGLQCDCSSSCNCSNS-KVLVNKASETNPK 206
>TC90676 similar to GP|6503308|gb|AAF14684.1| Contains PF|00646 F-box
domain. ESTs gb|AA586135 gb|Z26675 gb|AI993239 and
gb|AA585907 come from, partial (5%)
Length = 1119
Score = 30.0 bits (66), Expect = 5.3
Identities = 16/53 (30%), Positives = 23/53 (43%)
Frame = +1
Query: 917 GCCRKFTNGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTM 969
GCCR C D + +DD + + CT R+ +LR D ST+
Sbjct: 901 GCCRSAHATMRCCVDRRRAVATADDDHTRTRLRRHCTGVRRNGELRRDARSTV 1059
>AL387128 similar to PIR|A36621|A366 NADH dehydrogenase (ubiquinone) (EC
1.6.5.3) 22K chain precursor - Neurospora crassa,
partial (24%)
Length = 476
Score = 29.3 bits (64), Expect = 9.0
Identities = 13/40 (32%), Positives = 19/40 (47%)
Frame = +3
Query: 199 CSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVDF 238
CSC DCP+ ++ TT S + +G+ C DF
Sbjct: 69 CSCRDCPNENLLGREVGTTSAPLKSAAFFIGAY---CKDF 179
>TC80169
Length = 811
Score = 29.3 bits (64), Expect = 9.0
Identities = 13/25 (52%), Positives = 20/25 (80%)
Frame = +2
Query: 676 LEGRISNALVEVGPSITLASLSEVL 700
LEG+++ A +EVG ++LA LSE+L
Sbjct: 458 LEGKMNKAYLEVGLGVSLAHLSELL 532
>TC80290 weakly similar to GP|13937149|gb|AAK50068.1 At1g79380/T8K14_20
{Arabidopsis thaliana}, partial (10%)
Length = 1735
Score = 29.3 bits (64), Expect = 9.0
Identities = 36/150 (24%), Positives = 62/150 (41%), Gaps = 11/150 (7%)
Frame = +1
Query: 421 VDAIRANYSGLMVSLQDICMKPLDKDCATQSVLQYFKMDPRNFDDSGAVEHLNYCFQQYS 480
+++I+ GL +C P + C+T ++Y ++FD L C ++ S
Sbjct: 463 IESIKETAIGLQHGGCAVCGNPCSQKCSTCKAIKYCSQACQHFDWK-LGHKLQCCVKKTS 639
Query: 481 SADQCMSAFKAPLDPSTVL----GGFSGKDYSGASAFIVTYPVNNA-------IDEEGNE 529
S S + P D + ++ D+S + Y N I +E N
Sbjct: 640 STQVTTSNQEWPTDENVIMPPNPDEVVDNDHSYDPLCLEFYSEENTNNKALILISQEANN 819
Query: 530 TAKAVAWEKAFIQLVKDELLPMAQSRNLTL 559
+ V EK ++ +KDE L M +S N+TL
Sbjct: 820 KVQEVE-EK--MRNLKDE-LEMIRSENMTL 897
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 36,901,881
Number of Sequences: 36976
Number of extensions: 575570
Number of successful extensions: 3124
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 3062
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3117
length of query: 1140
length of database: 9,014,727
effective HSP length: 107
effective length of query: 1033
effective length of database: 5,058,295
effective search space: 5225218735
effective search space used: 5225218735
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0193.4


