

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0192.11
(571 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
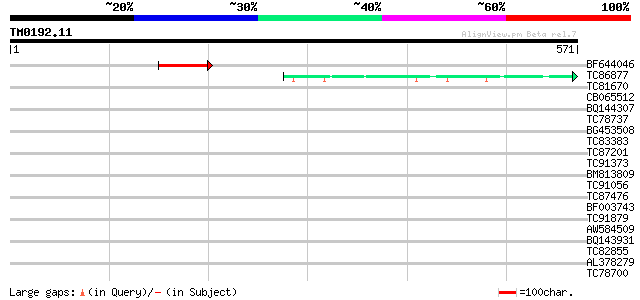
Score E
Sequences producing significant alignments: (bits) Value
BF644046 60 2e-09
TC86877 similar to GP|8953376|emb|CAB96649.1 putative protein {A... 50 3e-06
TC81670 weakly similar to PIR|E86293|E86293 hypothetical protein... 35 0.078
CB065512 similar to GP|3608154|gb| unknown protein {Arabidopsis ... 34 0.17
BQ144307 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 34 0.17
TC78737 similar to GP|1514443|emb|CAA67508.1 blue light receptor... 33 0.39
BG453508 weakly similar to GP|15625407|gb| like heterochromatin ... 32 0.66
TC83383 31 1.1
TC87201 similar to PIR|F86388|F86388 hypothetical protein AAG292... 30 1.9
TC91373 weakly similar to GP|12597773|gb|AAG60086.1 hypothetical... 30 1.9
BM813809 homologue to GP|15981743|emb hypothetical protein {Yers... 30 1.9
TC91056 similar to GP|16555790|emb|CAD10386. L-galactose dehydro... 30 2.5
TC87476 homologue to PIR|T11622|T11622 extensin class 1 precurso... 30 2.5
BF003743 similar to GP|17016406|gb| wsv008 {shrimp white spot sy... 30 2.5
TC91879 weakly similar to GP|5305335|gb|AAD41594.1| proline-rich... 30 2.5
AW584509 homologue to GP|13646986|db DNA-binding protein DF1 {Pi... 30 2.5
BQ143931 similar to PIR|T02229|T022 protein BYJ15 - common tobac... 30 2.5
TC82855 weakly similar to PIR|T45829|T45829 hypothetical protein... 30 3.3
AL378279 homologue to GP|17133068|dbj cobyrinic acid a c-diamide... 29 5.6
TC78700 similar to GP|21554835|gb|AAM63703.1 unknown {Arabidopsi... 29 5.6
>BF644046
Length = 597
Score = 60.5 bits (145), Expect = 2e-09
Identities = 26/54 (48%), Positives = 37/54 (68%)
Frame = +3
Query: 151 IFYVYEYQFTELGIKFPFSSFFCDVLADINVAPCQLHPNAWAFMRCFEILCVAI 204
+ ++Y + F ++G KFPF++F CD L +NVA QLHPN AFM FEI C ++
Sbjct: 3 MLHMYSFVFEDIGFKFPFTNFECDFLKALNVASSQLHPNCCAFMCGFEISCESL 164
>TC86877 similar to GP|8953376|emb|CAB96649.1 putative protein {Arabidopsis
thaliana}, partial (12%)
Length = 1558
Score = 49.7 bits (117), Expect = 3e-06
Identities = 75/324 (23%), Positives = 126/324 (38%), Gaps = 28/324 (8%)
Frame = +2
Query: 276 ASFPFYWTD----KPNGIVDPPVDSLTVEDKTIIGFFAQLPILD---CSKLAAARRLAKF 328
AS P++ D K D V LT + K +G +LP+LD SK+ A LA
Sbjct: 365 ASNPYHACDVNCQKRMSGADSGVIPLTFDRKKKLGSRPELPVLDSVPASKIGAIY-LADA 541
Query: 329 PPNFNTENPKKKRHQRDNEVCENSNPVRQGTGEKESRTALREKEGHFDLAEKEARPNLGE 388
+ KKK + NE+ S + K ++ K+G +LA + G+
Sbjct: 542 ASPISKYYDKKKLEPKSNEIVPASGELHT-LDVKPVNDKVQPKDGSENLAGQIQTNQAGD 718
Query: 389 KEKEVPSDPSTMLRDATDHV--------EFLLSRINHDYLEKEVFGKGIDDAGQECVAS- 439
K P T D + + F S + HD + + D+ G E V S
Sbjct: 719 KNSSNKVVPVTCFDDIGEGLTTSAGGSKHFCYSDVLHDNEDSD------DEEGSESVVSE 880
Query: 440 ------LFHTGCIFAHAFQKFGASTAE-GEKLQADYTALRTEHAKC-----DDRLDKVLT 487
+H FA Q + GE + +R+ + +C + +
Sbjct: 881 TRVPVGKYHVKESFAPILQSIFDKYGDIGESCHLESVVMRSYYIECVCFVIQELQSTSIM 1060
Query: 488 EAARDKITIDKKLVSIANELVEARFELVCRSERITDLEEEIKETKQEAQLALAAKDDALA 547
+ + K+ K+L++I N++ A+ + + ++ E IK + + + A A
Sbjct: 1061DLTKSKV---KELLAILNDVESAQLHVTWLRTVVNEIAEYIKLIDEHRSV-----EAAKA 1216
Query: 548 LKDQELTSLRAELEEKKRDLLKKK 571
D E+ SLR ELE K L +K+
Sbjct: 1217NSDSEMESLRKELESKVEILTQKE 1288
>TC81670 weakly similar to PIR|E86293|E86293 hypothetical protein AAF18488.1
[imported] - Arabidopsis thaliana, partial (41%)
Length = 1797
Score = 35.0 bits (79), Expect = 0.078
Identities = 30/85 (35%), Positives = 41/85 (47%), Gaps = 4/85 (4%)
Frame = +2
Query: 484 KVLTEAARDKITIDKKLVSIANE-LVEARFELVCRSERITDLEEEIKETKQEAQ---LAL 539
+VLT+ R K+ DKK NE L+ A E E + L EE K K+EA L L
Sbjct: 1088 EVLTDQERQKLEEDKKKKDSRNESLMLASKEQKIADENVFRLVEEQKREKEEALNKILQL 1267
Query: 540 AAKDDALALKDQELTSLRAELEEKK 564
+ DA + E+ LR +L+ K
Sbjct: 1268 EKQLDAKQKLEMEIEELRGKLQVMK 1342
>CB065512 similar to GP|3608154|gb| unknown protein {Arabidopsis thaliana},
partial (34%)
Length = 852
Score = 33.9 bits (76), Expect = 0.17
Identities = 30/107 (28%), Positives = 48/107 (44%), Gaps = 1/107 (0%)
Frame = +1
Query: 460 EGEKLQADYTALRT-EHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRS 518
E KL A + E AK + + K L + K + K++ + +++E R
Sbjct: 232 EASKLSASESVTACLEVAKKEKKYLKKLLAWEKQKAKLQKEISDLKEKILEDREVSAQNK 411
Query: 519 ERITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKR 565
+R + E + KE L A++DALAL D+E S A + KR
Sbjct: 412 QRQKEAEAKWKEE-------LKAQEDALALVDEERRSKEAAESDNKR 531
>BQ144307 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (24%)
Length = 1059
Score = 33.9 bits (76), Expect = 0.17
Identities = 40/141 (28%), Positives = 57/141 (40%), Gaps = 22/141 (15%)
Frame = -3
Query: 26 TSNGRSLFKSITRH--FRTEAEWSLHQSFAFIFINAPPALPNVIIRSCHHESRFPTNSTR 83
+S R+ ++ TRH +RT + S S + AP +LP++ I H T+S R
Sbjct: 835 SSRARASPRTTTRHSRYRTRSRISHIASHSLPRPRAPLSLPSLHISPVPHT----TSSNR 668
Query: 84 TPRAPCK---TSPYAYKNPAPSSLLTFQTPPYS-----------------PDFFEQTSGV 123
TP A K TSP + P P + PP S P F Q + +
Sbjct: 667 TPPARTKLMVTSPRPHTPPNP---IVTSLPPLSLIK*ADVHGPWSFCYHPPKSFAQPNSI 497
Query: 124 FLFRHRCQPATYPFFNPFLTP 144
L P + P +PFL P
Sbjct: 496 SL-----HPPSPPTLSPFLPP 449
>TC78737 similar to GP|1514443|emb|CAA67508.1 blue light receptor {Arabidopsis
thaliana}, partial (62%)
Length = 1721
Score = 32.7 bits (73), Expect = 0.39
Identities = 14/36 (38%), Positives = 20/36 (54%)
Frame = -3
Query: 44 AEWSLHQSFAFIFINAPPALPNVIIRSCHHESRFPT 79
A WSLHQ FA+ + + A P ++ + C H S T
Sbjct: 1173 ARWSLHQPFAYSWSSEVAACP*ILSKPCQHCSNLET 1066
>BG453508 weakly similar to GP|15625407|gb| like heterochromatin protein LHP1
{Arabidopsis thaliana}, partial (12%)
Length = 589
Score = 32.0 bits (71), Expect = 0.66
Identities = 23/93 (24%), Positives = 35/93 (36%), Gaps = 4/93 (4%)
Frame = -3
Query: 71 CHHESRFPTNSTRTPRAPCKTSP----YAYKNPAPSSLLTFQTPPYSPDFFEQTSGVFLF 126
C H PT S TP PC S +++N + Q + P F Q
Sbjct: 530 CLHAHPSPTISQSTPTYPCGISYDE*LQSHRNHHQEKEVFLQVQDHHPHFLHQNF----- 366
Query: 127 RHRCQPATYPFFNPFLTPSSARYHIFYVYEYQF 159
P+ YP F +P + H+F+ + + F
Sbjct: 365 -----PSGYPHFPTGYSPFQ*KNHLFHHHHHHF 282
>TC83383
Length = 534
Score = 31.2 bits (69), Expect = 1.1
Identities = 19/44 (43%), Positives = 24/44 (54%)
Frame = +2
Query: 11 SPSVHKSTSGEGRATTSNGRSLFKSITRHFRTEAEWSLHQSFAF 54
S SVH+S S A S RS KS T+H + ++HQSF F
Sbjct: 110 SSSVHRSRSSASLAQNSRQRS--KSQTKHH*LSEKGTVHQSFMF 235
>TC87201 similar to PIR|F86388|F86388 hypothetical protein AAG29221.1
[imported] - Arabidopsis thaliana, partial (41%)
Length = 991
Score = 30.4 bits (67), Expect = 1.9
Identities = 20/68 (29%), Positives = 31/68 (45%)
Frame = +1
Query: 47 SLHQSFAFIFINAPPALPNVIIRSCHHESRFPTNSTRTPRAPCKTSPYAYKNPAPSSLLT 106
S H +++ + PP P IR +R P +P P PY YK+P P +
Sbjct: 16 SHHLLRPYVYPSPPPPSPPPPIR-IQITTRAPPYVYPSPPPPSPPPPYIYKSPPPPPYV- 189
Query: 107 FQTPPYSP 114
+++PP P
Sbjct: 190 YKSPPPPP 213
Score = 28.1 bits (61), Expect = 9.5
Identities = 12/34 (35%), Positives = 19/34 (55%)
Frame = +1
Query: 85 PRAPCKTSPYAYKNPAPSSLLTFQTPPYSPDFFE 118
P +P PY YK+P P + +++PP P +E
Sbjct: 328 PPSPSPPPPYVYKSPPPPPYV-YKSPPPPPYVYE 426
>TC91373 weakly similar to GP|12597773|gb|AAG60086.1 hypothetical protein
{Arabidopsis thaliana}, partial (11%)
Length = 677
Score = 30.4 bits (67), Expect = 1.9
Identities = 16/60 (26%), Positives = 31/60 (51%)
Frame = +1
Query: 480 DRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSERITDLEEEIKETKQEAQLAL 539
+ L K EA +K I + +++ E+ + FE+ E++ + +E++E K LAL
Sbjct: 67 ETLKKETDEAKTEKELISQDIITTKLEIEKTEFEIDTSEEKLQGVMKELEEAKTTEALAL 246
>BM813809 homologue to GP|15981743|emb hypothetical protein {Yersinia
pestis}, partial (3%)
Length = 667
Score = 30.4 bits (67), Expect = 1.9
Identities = 17/48 (35%), Positives = 29/48 (60%), Gaps = 1/48 (2%)
Frame = +3
Query: 524 LEEEIKETKQEAQLALAAKDDA-LALKDQELTSLRAELEEKKRDLLKK 570
LE+EIK+ +AQL ++ L+ + E+ + EL +K+ DLLK+
Sbjct: 342 LEQEIKQEVMQAQLVEVMEEHRNLSARQAEMAKKQEELAKKQDDLLKQ 485
>TC91056 similar to GP|16555790|emb|CAD10386. L-galactose dehydrogenase
{Arabidopsis thaliana}, complete
Length = 1323
Score = 30.0 bits (66), Expect = 2.5
Identities = 17/60 (28%), Positives = 22/60 (36%)
Frame = +3
Query: 52 FAFIFINAPPALPNVIIRSCHHESRFPTNSTRTPRAPCKTSPYAYKNPAPSSLLTFQTPP 111
F ++ PP +VI+ CHH T P K +P LLT PP
Sbjct: 492 FTYVLDRVPPGTLDVILSYCHHSINDSTLEDIVPYLKSKGVGIISASPLAMGLLTEAGPP 671
>TC87476 homologue to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (48%)
Length = 939
Score = 30.0 bits (66), Expect = 2.5
Identities = 21/65 (32%), Positives = 26/65 (39%)
Frame = +3
Query: 48 LHQSFAFIFINAPPALPNVIIRSCHHESRFPTNSTRTPRAPCKTSPYAYKNPAPSSLLTF 107
LH FIN ++ HH T+ R P +P PY YK+P P S
Sbjct: 279 LHPHHLHTFINL------LLHHLLHHRLHISTSHHRPP-SPSPPPPYVYKSPPPPS--PS 431
Query: 108 QTPPY 112
PPY
Sbjct: 432 PPPPY 446
>BF003743 similar to GP|17016406|gb| wsv008 {shrimp white spot syndrome
virus}, partial (16%)
Length = 598
Score = 30.0 bits (66), Expect = 2.5
Identities = 24/132 (18%), Positives = 66/132 (49%), Gaps = 14/132 (10%)
Frame = +2
Query: 453 KFGASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAAR--DKIT-----IDKKLVSIAN 505
K T++ + L AD ++L + + ++++ TEA+ + IT + +++ S+ +
Sbjct: 140 KISDLTSQIDNLLADISSLHAQKNELEEQIIFKSTEASTRAESITNELNVLQQEVESLQH 319
Query: 506 ELVEARFELVCRSE-------RITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRA 558
+ + +LV +S+ +I L+EE+ E + + +++ +++EL+ +
Sbjct: 320 QKFDLEVQLVEKSQENSKCSIQIQSLKEEVDRKSLEQERLMEDRENFAKEREEELSDIMK 499
Query: 559 ELEEKKRDLLKK 570
+L++ + + K
Sbjct: 500 KLKDNENESSSK 535
>TC91879 weakly similar to GP|5305335|gb|AAD41594.1| proline-rich mucin
homolog {Mycobacterium tuberculosis}, partial (4%)
Length = 882
Score = 30.0 bits (66), Expect = 2.5
Identities = 15/40 (37%), Positives = 22/40 (54%)
Frame = -1
Query: 75 SRFPTNSTRTPRAPCKTSPYAYKNPAPSSLLTFQTPPYSP 114
SR+PT+S + +P SP +P +S + TPP SP
Sbjct: 591 SRYPTHSAPSCTSPHTASPRPPAHPPRTSRVPRHTPPQSP 472
>AW584509 homologue to GP|13646986|db DNA-binding protein DF1 {Pisum
sativum}, partial (22%)
Length = 694
Score = 30.0 bits (66), Expect = 2.5
Identities = 32/114 (28%), Positives = 48/114 (42%), Gaps = 17/114 (14%)
Frame = +2
Query: 15 HKSTSGEGRATTSNGRSLFKSITRHFRTEAEWSLHQSFAFI---------------FINA 59
HK T EGR S+G++ ++ + E S+HQS + F++A
Sbjct: 374 HKRTK-EGRGGKSDGKT-YRFFDQLQALENNPSMHQSPSITPPPISTIPTPTTTPSFLSA 547
Query: 60 PPA--LPNVIIRSCHHESRFPTNSTRTPRAPCKTSPYAYKNPAPSSLLTFQTPP 111
PP +P+ I + +S N T P P T+P NP P +T PP
Sbjct: 548 PPTTTVPSTTIPMSYTQS----NPTHFPPIPSSTNPINTINPIPQ--ITTTPPP 691
>BQ143931 similar to PIR|T02229|T022 protein BYJ15 - common tobacco (fragment),
partial (15%)
Length = 1239
Score = 30.0 bits (66), Expect = 2.5
Identities = 18/65 (27%), Positives = 30/65 (45%), Gaps = 1/65 (1%)
Frame = +1
Query: 126 FRHRCQPATYPFFNPFLTPSSAR-YHIFYVYEYQFTELGIKFPFSSFFCDVLADINVAPC 184
FR C P P + PS+ R YH ++ FT + FP + F ++A ++ +P
Sbjct: 988 FRAHCPPIRPPTY-----PSNPRLYHYPHLSRPNFTFPSLPFPIDAIFTPIVAALSTSPP 1152
Query: 185 QLHPN 189
P+
Sbjct: 1153 YKQPS 1167
>TC82855 weakly similar to PIR|T45829|T45829 hypothetical protein F2K15.100 -
Arabidopsis thaliana, partial (63%)
Length = 1447
Score = 29.6 bits (65), Expect = 3.3
Identities = 22/56 (39%), Positives = 25/56 (44%), Gaps = 8/56 (14%)
Frame = -1
Query: 99 PAPSSLLTFQTPPYSPDFFEQTSGV-FLFRHRCQPATY-------PFFNPFLTPSS 146
P+P L F TPP +T G+ RH C P TY P F P TPSS
Sbjct: 1195 PSPLLSLLFSTPP*------RTLGIPHQSRHHCHPTTYLQSEAHHPTFLPRSTPSS 1046
>AL378279 homologue to GP|17133068|dbj cobyrinic acid a c-diamide synthase
{Nostoc sp. PCC 7120}, partial (2%)
Length = 321
Score = 28.9 bits (63), Expect = 5.6
Identities = 17/44 (38%), Positives = 21/44 (47%), Gaps = 1/44 (2%)
Frame = +1
Query: 312 PILDCSKLAAARRLAKFP-PNFNTENPKKKRHQRDNEVCENSNP 354
PI + K R FP PNF +E+P R Q + ENS P
Sbjct: 103 PITEIPKTPPFRSHQNFPLPNFTSESPPSPRQQWR*HLSENSPP 234
>TC78700 similar to GP|21554835|gb|AAM63703.1 unknown {Arabidopsis
thaliana}, partial (5%)
Length = 687
Score = 28.9 bits (63), Expect = 5.6
Identities = 20/69 (28%), Positives = 38/69 (54%), Gaps = 5/69 (7%)
Frame = +2
Query: 329 PPNFNTENPKKKRHQRDNEVCENSNPVRQGTGEKESRTALREKEGHFDLAEKEARPNL-- 386
P N + K+K+++ + V + SNP+ + + +KE + ++K G + EKE NL
Sbjct: 311 PENGVVDEKKRKKNKEKSPVID-SNPIIEVSEKKEKKK--KKKSGSGEEEEKEKEVNLDG 481
Query: 387 ---GEKEKE 392
G+K+K+
Sbjct: 482 EGVGDKKKK 508
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.134 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,842,004
Number of Sequences: 36976
Number of extensions: 314750
Number of successful extensions: 1844
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 1740
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1833
length of query: 571
length of database: 9,014,727
effective HSP length: 101
effective length of query: 470
effective length of database: 5,280,151
effective search space: 2481670970
effective search space used: 2481670970
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0192.11


