

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0191.15
(398 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
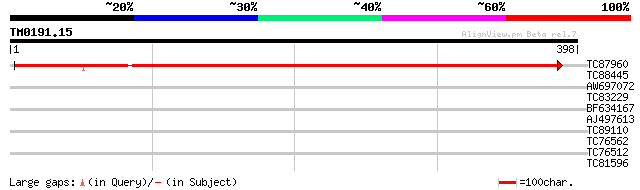
TC87960 homologue to PIR|S39523|S39523 coproporphyrinogen oxidas... 637 0.0
TC88445 similar to GP|16648903|gb|AAL24303.1 alpha glucosidase-l... 31 0.74
AW697072 weakly similar to GP|17979155|gb| unknown protein {Arab... 24 2.0
TC83229 30 2.1
BF634167 29 2.8
AJ497613 similar to GP|7295919|gb| Pgk gene product {Drosophila ... 29 3.6
TC89110 weakly similar to GP|17064780|gb|AAL32544.1 Unknown prot... 28 6.2
TC76562 similar to PIR|T11622|T11622 extensin class 1 precursor ... 28 8.1
TC76512 similar to GP|4959932|gb|AAD34548.1| cystathionine-gamma... 28 8.1
TC81596 similar to GP|21039012|dbj|BAB92985. protein kinase {Ara... 28 8.1
>TC87960 homologue to PIR|S39523|S39523 coproporphyrinogen oxidase (EC
1.3.3.3) precursor - soybean, partial (84%)
Length = 1581
Score = 637 bits (1643), Expect = 0.0
Identities = 313/389 (80%), Positives = 336/389 (85%), Gaps = 4/389 (1%)
Frame = +3
Query: 4 CATIVSPPSYAI-PFFTSTSSTSSPTAKLLSKRRWNPLIQTKTMTSLG--SKGAVRAGVS 60
C +S PSY+I PFFT TSS S+ + + + QT+ TS + +A VS
Sbjct: 132 CTISISAPSYSITPFFTFTSSKSTQISHISHTPPKSLTTQTRKTTSSHHFQRLITKATVS 311
Query: 61 IEKETPEAERPETFLRGGDTAEALSSSSSVRARFEKMIREAQDSVCDAIEAADGGAKFKE 120
IEKETPE+ERP+TFLRG D A ++SSVRARFEKMIREAQD VC+AIE ADGG KFKE
Sbjct: 312 IEKETPESERPDTFLRGVDNDGA--TASSVRARFEKMIREAQDKVCNAIEQADGGGKFKE 485
Query: 121 DVWSRPGGGGGISRVLQDGAVWEKAGVNVSVVYGVMPPDAYRAAKG-AAASPDEKPGPVP 179
DVWSRPGGGGGISRVLQDGAVWEKAGVNVSVVYG MPPDAYRAAKG AAAS D+KPGP+P
Sbjct: 486 DVWSRPGGGGGISRVLQDGAVWEKAGVNVSVVYGQMPPDAYRAAKGAAAASSDQKPGPIP 665
Query: 180 FFAAGISSVLHPKNPFAPTMHFNYRYFETDAPKDAPGAPKQWWFGGGTDLTPAYIFEEDV 239
FFAAGISSVLHPKNPFAPT+HFNYRYFETDAPKDAPGAP+QWWFGGGTDLTPAYIFEEDV
Sbjct: 666 FFAAGISSVLHPKNPFAPTLHFNYRYFETDAPKDAPGAPRQWWFGGGTDLTPAYIFEEDV 845
Query: 240 KHFHSIQKQACDKFDPTFYPRFKKWCDDYFYIKHRGERRGLGGIFFDDLNDYDQEMLLSF 299
KHFHS+QKQACDKFDPTFYPRFKKWCDDYFYIKHR ERRGLGGIFFDDLNDYDQEMLLSF
Sbjct: 846 KHFHSVQKQACDKFDPTFYPRFKKWCDDYFYIKHRDERRGLGGIFFDDLNDYDQEMLLSF 1025
Query: 300 ATECANSVVPSYIPIIEKRKDTPFTDHQKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRI 359
ATECANSV+P+YIPIIEKRKD PFTDHQKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRI
Sbjct: 1026ATECANSVIPAYIPIIEKRKDLPFTDHQKAWQQLRRGRYVEFNLVYDRGTTFGLKTGGRI 1205
Query: 360 ESILVSLPLTSRWEYDHKPEEGSEEWKLL 388
ESIL SLPLT+RWEYDHKP + L
Sbjct: 1206ESILGSLPLTARWEYDHKPRSRKRRMETL 1292
Score = 39.3 bits (90), Expect = 0.003
Identities = 16/17 (94%), Positives = 16/17 (94%)
Frame = +1
Query: 378 PEEGSEEWKLLDACINP 394
PE GSEEWKLLDACINP
Sbjct: 1261 PEVGSEEWKLLDACINP 1311
>TC88445 similar to GP|16648903|gb|AAL24303.1 alpha glucosidase-like protein
{Arabidopsis thaliana}, partial (31%)
Length = 1077
Score = 31.2 bits (69), Expect = 0.74
Identities = 12/26 (46%), Positives = 15/26 (57%)
Frame = -2
Query: 155 VMPPDAYRAAKGAAASPDEKPGPVPF 180
+ PP Y+A K ASP KP +PF
Sbjct: 572 IEPPFKYKAGKSGCASPKSKPNHIPF 495
>AW697072 weakly similar to GP|17979155|gb| unknown protein {Arabidopsis
thaliana}, partial (21%)
Length = 739
Score = 24.3 bits (51), Expect(2) = 2.0
Identities = 14/37 (37%), Positives = 19/37 (50%), Gaps = 1/37 (2%)
Frame = -1
Query: 162 RAAKGA-AASPDEKPGPVPFFAAGISSVLHPKNPFAP 197
RAA+ A P + P P+P A S PK+ F+P
Sbjct: 559 RAARVAHPLQPRQPPAPLPSQLASPPSSTRPKDTFSP 449
Score = 23.9 bits (50), Expect(2) = 2.0
Identities = 15/51 (29%), Positives = 20/51 (38%)
Frame = -2
Query: 191 PKNPFAPTMHFNYRYFETDAPKDAPGAPKQWWFGGGTDLTPAYIFEEDVKH 241
P+ PF PT HF + +P P GGG + I + KH
Sbjct: 468 PRTPFPPTQHFTHL-----SPPQGP-VILSHTLGGGIHVVHCQIHLQCAKH 334
>TC83229
Length = 692
Score = 29.6 bits (65), Expect = 2.1
Identities = 20/73 (27%), Positives = 26/73 (35%)
Frame = +1
Query: 160 AYRAAKGAAASPDEKPGPVPFFAAGISSVLHPKNPFAPTMHFNYRYFETDAPKDAPGAPK 219
+Y AA G + P P P P G H N + +FN AP+ P
Sbjct: 91 SYPAASGPSYQPPRTPPPPPTSNVGYPPNFHQSNNGLASSYFNMPSSSGTAPRQR--GPP 264
Query: 220 QWWFGGGTDLTPA 232
+ G G PA
Sbjct: 265 GFAMGAGAGALPA 303
>BF634167
Length = 221
Score = 29.3 bits (64), Expect = 2.8
Identities = 20/76 (26%), Positives = 28/76 (36%)
Frame = -2
Query: 157 PPDAYRAAKGAAASPDEKPGPVPFFAAGISSVLHPKNPFAPTMHFNYRYFETDAPKDAPG 216
PPD + G A P + P P FA + ++ P +F P+ G
Sbjct: 220 PPDVLATSPGQTAWPGCRACPSPAFACPPACLVAPAAS---------SFFLL*TPQGQAG 68
Query: 217 APKQWWFGGGTDLTPA 232
+ K WW G PA
Sbjct: 67 SGKPWWSGSQMVQPPA 20
>AJ497613 similar to GP|7295919|gb| Pgk gene product {Drosophila
melanogaster}, partial (41%)
Length = 607
Score = 28.9 bits (63), Expect = 3.6
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = -1
Query: 291 YDQEMLLSFATECANSVVPSYIPIIEKRKDTPF 323
+ L FA +N+ VP +IP I R+ TPF
Sbjct: 118 FSTNFLTIFAPSSSNNEVPIFIPFISFRRCTPF 20
>TC89110 weakly similar to GP|17064780|gb|AAL32544.1 Unknown protein
{Arabidopsis thaliana}, partial (21%)
Length = 706
Score = 28.1 bits (61), Expect = 6.2
Identities = 12/22 (54%), Positives = 15/22 (67%)
Frame = +2
Query: 16 PFFTSTSSTSSPTAKLLSKRRW 37
P F++TSS + P K LS RRW
Sbjct: 263 PRFSTTSSLNQPEFKSLSSRRW 328
>TC76562 similar to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (48%)
Length = 1391
Score = 27.7 bits (60), Expect = 8.1
Identities = 25/116 (21%), Positives = 37/116 (31%), Gaps = 3/116 (2%)
Frame = +2
Query: 158 PDAYRAAKGAAASPDEKPGPVPFFAAG---ISSVLHPKNPFAPTMHFNYRYFETDAPKDA 214
P AY + K KP P P++ S V +P P +++ Y P
Sbjct: 656 PFAYASKKHFKECEKPKPSPTPYYYKSPPPPSPVYKYNSPPPPVHYYSPPYTYKSPPPPV 835
Query: 215 PGAPKQWWFGGGTDLTPAYIFEEDVKHFHSIQKQACDKFDPTFYPRFKKWCDDYFY 270
AP +++ P Y + H K P P K Y+Y
Sbjct: 836 KAAPTPYYYKSPPPPAPVYKYNSPPPPVHYYSPPYTYKSPP---PPVKAAPTPYYY 994
>TC76512 similar to GP|4959932|gb|AAD34548.1| cystathionine-gamma-synthase
precursor {Glycine max}, partial (90%)
Length = 2103
Score = 27.7 bits (60), Expect = 8.1
Identities = 17/57 (29%), Positives = 24/57 (41%), Gaps = 3/57 (5%)
Frame = -2
Query: 101 AQDSVCDAIEAADG---GAKFKEDVWSRPGGGGGISRVLQDGAVWEKAGVNVSVVYG 154
+ D +C+ AA GA+ ++W + GGG G WE V VV G
Sbjct: 308 SSDDLCNTDVAAVTTRFGAELTLEIWRKTEDGGGDGSCRGKGGCWEMTTAVVVVVVG 138
>TC81596 similar to GP|21039012|dbj|BAB92985. protein kinase {Arabidopsis
thaliana}, partial (33%)
Length = 692
Score = 27.7 bits (60), Expect = 8.1
Identities = 11/22 (50%), Positives = 13/22 (59%)
Frame = -2
Query: 322 PFTDHQKAWQQLRRGRYVEFNL 343
P TDH + W Q RR RY + L
Sbjct: 211 PCTDHNERWDQFRRIRYRDHRL 146
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.135 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,703,334
Number of Sequences: 36976
Number of extensions: 204450
Number of successful extensions: 1076
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1063
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1073
length of query: 398
length of database: 9,014,727
effective HSP length: 98
effective length of query: 300
effective length of database: 5,391,079
effective search space: 1617323700
effective search space used: 1617323700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0191.15


