

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0190.3
(329 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
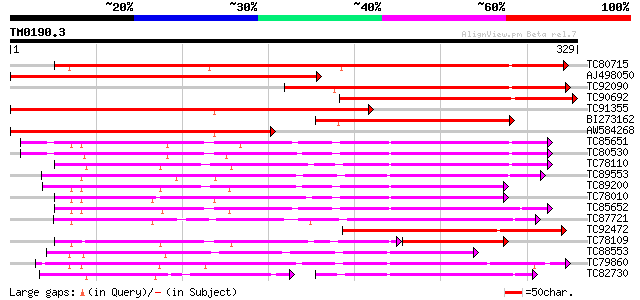
Score E
Sequences producing significant alignments: (bits) Value
TC80715 similar to GP|5579092|gb|AAD45424.1| gibberellin 2-oxida... 308 2e-84
AJ498050 similar to GP|6478200|gb| gibberellin 2 beta-hydroxylas... 281 2e-76
TC92090 similar to GP|4678586|emb|CAB41036.1 GA 2-oxidase {Phase... 252 1e-67
TC90692 homologue to GP|5579094|gb|AAD45425.1| gibberellin 2-oxi... 248 2e-66
TC91355 similar to GP|6446413|gb|AAF08609.1| gibberellin 2-beta-... 244 4e-65
BI273162 similar to GP|21553670|gb gibberellin 2- oxidase {Arabi... 168 2e-42
AW584268 similar to GP|4678586|emb GA 2-oxidase {Phaseolus cocci... 168 2e-42
TC85651 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis ... 147 6e-36
TC80530 weakly similar to PIR|S44261|S44261 SRG1 protein - Arabi... 141 3e-34
TC78110 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis ... 141 3e-34
TC89553 weakly similar to SP|P24397|HY6H_HYONI Hyoscyamine 6-dio... 140 5e-34
TC89200 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis ... 140 9e-34
TC78010 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis ... 140 9e-34
TC85652 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis ... 139 2e-33
TC87721 weakly similar to PIR|H96594|H96594 hypothetical protein... 138 3e-33
TC92472 similar to GP|5579092|gb|AAD45424.1| gibberellin 2-oxida... 137 7e-33
TC78109 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis ... 102 2e-32
TC88553 weakly similar to GP|21327037|gb|AAM48133.1 putative fla... 135 2e-32
TC79860 homologue to PIR|T06533|T06533 probable gibberellin 20-o... 134 5e-32
TC82730 similar to GP|10177853|dbj|BAB11205. flavanone 3-hydroxy... 88 1e-31
>TC80715 similar to GP|5579092|gb|AAD45424.1| gibberellin 2-oxidase-like
protein {Pisum sativum}, partial (97%)
Length = 1272
Score = 308 bits (788), Expect = 2e-84
Identities = 158/313 (50%), Positives = 209/313 (66%), Gaps = 15/313 (4%)
Frame = +2
Query: 27 IPVVDLS--KPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKT 84
IP +DLS + + ++VKACEE+GFFKV+NHG+ E IS+LE+E FFS TEK +
Sbjct: 101 IPTIDLSMERSELSELVVKACEEYGFFKVVNHGIPKEVISRLENEGTEFFSKNSTEKLQA 280
Query: 85 GLANPFGYGNKRIGHNGDVGWVEYLLLKTN----QEVNVPVHSQNPEKFRCVLDDYLCAV 140
G + PFGYG K IG NGD G +EYLLL TN E + + +P KF C++ DY+ AV
Sbjct: 281 GTSTPFGYGCKNIGPNGDKGDLEYLLLHTNPNSISERSKTIAKDHPIKFSCIVTDYIEAV 460
Query: 141 RNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDG--------- 191
+ + CEIL+L AEGL + K+ LSK++ D SDSV R+NHYP ++ D
Sbjct: 461 KELACEILELAAEGLWVPDKSSLSKVIKDVHSDSVLRINHYPPVKKLSKDNLDPSKFQNN 640
Query: 192 ENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGR 251
N IGFGEH+DPQI+++LRSNN GLQIS + G WI V PD + F++ VGDSLQV+TNGR
Sbjct: 641 NNTIGFGEHSDPQILTILRSNNVGGLQISTQHGLWIPVHPDPNEFYVMVGDSLQVLTNGR 820
Query: 252 FRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSY 311
F SV+HRVL N + R+SM+YF PPL+ I+PL ++ SLYK FTW +YK ++Y
Sbjct: 821 FVSVRHRVLTNTTKPRMSMMYFAAPPLNWWISPLSKMVT-AHNPSLYKPFTWAQYKQAAY 997
Query: 312 GSRLADNRLGHFE 324
RL D+RL F+
Sbjct: 998 ALRLGDSRLDQFK 1036
>AJ498050 similar to GP|6478200|gb| gibberellin 2 beta-hydroxylase {Pisum
sativum}, partial (71%)
Length = 667
Score = 281 bits (720), Expect = 2e-76
Identities = 139/182 (76%), Positives = 158/182 (86%), Gaps = 1/182 (0%)
Frame = +1
Query: 1 MVLLSKQATEQYSYVKNCM-ATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVS 59
MVLLSK ++EQY+YV+N M AT S +IP+VDLSKPDAKS+IVKACE+FGFFKVINHG+
Sbjct: 121 MVLLSKPSSEQYTYVRNNMQATTFSSSIPLVDLSKPDAKSLIVKACEDFGFFKVINHGIP 300
Query: 60 MEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNV 119
MEAISQLESEA FFS+P+TEKEK G ANPFGYGNKRIG NGDVGWVEYLLL TNQE N
Sbjct: 301 MEAISQLESEAFKFFSLPLTEKEKAGPANPFGYGNKRIGPNGDVGWVEYLLLNTNQEHNF 480
Query: 120 PVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVN 179
+H ++ +KFRC+L+DY CA+RNM CEILDLMAEGLKIQPKNV SK +MDKQSDS FRVN
Sbjct: 481 SLHGKDIDKFRCLLNDYKCAMRNMACEILDLMAEGLKIQPKNVFSKPVMDKQSDSAFRVN 660
Query: 180 HY 181
Y
Sbjct: 661 RY 666
>TC92090 similar to GP|4678586|emb|CAB41036.1 GA 2-oxidase {Phaseolus
coccineus}, partial (48%)
Length = 776
Score = 252 bits (644), Expect = 1e-67
Identities = 123/169 (72%), Positives = 146/169 (85%), Gaps = 3/169 (1%)
Frame = +3
Query: 160 KNVLSKLLMDKQSDSVFRVNHYPACPEM---GLDGENLIGFGEHTDPQIISLLRSNNTSG 216
KNVLS+LL D++SDS F++NHYP CPE+ L+G NL+GFGEHTDPQ+IS+LRSN+TS
Sbjct: 3 KNVLSRLLKDEKSDSCFKINHYPPCPEVQQAALNGRNLLGFGEHTDPQVISVLRSNSTSR 182
Query: 217 LQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGP 276
LQI L DG+W+SVPPD++SFFINVGD+LQV+TNGRF+SVKHRVLA+ +SRLSMIYFGGP
Sbjct: 183 LQICLTDGTWVSVPPDHTSFFINVGDTLQVLTNGRFKSVKHRVLADTTKSRLSMIYFGGP 362
Query: 277 PLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGSRLADNRLGHFER 325
PLSEKI PLPSL+ KEESLYKEFTW EYK + Y SRLAD RLG FE+
Sbjct: 363 PLSEKIVPLPSLML-KKEESLYKEFTWLEYKKAMYNSRLADYRLGPFEK 506
>TC90692 homologue to GP|5579094|gb|AAD45425.1| gibberellin 2-oxidase {Pisum
sativum}, partial (41%)
Length = 652
Score = 248 bits (634), Expect = 2e-66
Identities = 122/138 (88%), Positives = 130/138 (93%)
Frame = +1
Query: 192 ENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGR 251
ENLIGFGEHTDPQIISLLRSNNTSG QISLRDGSWISVPPD+ SFFINVGDSLQVMTNGR
Sbjct: 1 ENLIGFGEHTDPQIISLLRSNNTSGFQISLRDGSWISVPPDHRSFFINVGDSLQVMTNGR 180
Query: 252 FRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSY 311
F+SV+HRVLANG RLSMIYFGGPPLSEKIAPLPSL+KG ESLYKEFTWFEYKNS+Y
Sbjct: 181 FKSVRHRVLANGINPRLSMIYFGGPPLSEKIAPLPSLMKG--NESLYKEFTWFEYKNSTY 354
Query: 312 GSRLADNRLGHFERIAAS 329
G+RLADNRLG++ERIAAS
Sbjct: 355 GTRLADNRLGNYERIAAS 408
>TC91355 similar to GP|6446413|gb|AAF08609.1| gibberellin 2-beta-hydroxylase
{Pisum sativum}, partial (46%)
Length = 753
Score = 244 bits (622), Expect = 4e-65
Identities = 125/218 (57%), Positives = 160/218 (73%), Gaps = 7/218 (3%)
Frame = +2
Query: 1 MVLLSKQAT-EQYSYVKNCMATPCSFT-IPVVDLSKPDAKSIIVKACEEFGFFKVINHGV 58
MV+LS+QAT + ++ C T F +P VDLS P+AK++IV AC EFGFFKV+NH V
Sbjct: 74 MVVLSQQATLNELFHINTCKPTSYVFKGVPEVDLSDPEAKTLIVNACTEFGFFKVVNHQV 253
Query: 59 SMEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEV- 117
+E I+ LE+E L FF P EKEK G +PFGYG+K IG NGDVGWVEYLLL TN +V
Sbjct: 254 PLELITNLENETLKFFGQPQLEKEKAGPPDPFGYGSKIIGTNGDVGWVEYLLLNTNPDVI 433
Query: 118 ---NVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDS 174
++ + QN + FRC ++Y+ AV+ + CE+L+LMA+GL+I+PKNV SKL+ D++SDS
Sbjct: 434 SLKSLFLLQQNTKNFRCAAEEYIVAVKEVCCEVLELMADGLRIEPKNVFSKLVRDERSDS 613
Query: 175 VFRVNHYPACPEM-GLDGENLIGFGEHTDPQIISLLRS 211
RVNHY AC E+ L G NLIGFGEHTDPQIIS+LRS
Sbjct: 614 CLRVNHYAACGELQALSGGNLIGFGEHTDPQIISVLRS 727
>BI273162 similar to GP|21553670|gb gibberellin 2- oxidase {Arabidopsis
thaliana}, partial (32%)
Length = 372
Score = 168 bits (426), Expect = 2e-42
Identities = 83/120 (69%), Positives = 96/120 (79%), Gaps = 4/120 (3%)
Frame = +2
Query: 178 VNHYPACPEMGL----DGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDY 233
+N YP P + EN+IGFGEHTDPQI SLLRSNNT GLQI L+D SWISVP D+
Sbjct: 2 LNXYPXXPPKSNXNNNENENVIGFGEHTDPQIXSLLRSNNTXGLQIRLKDKSWISVPSDH 181
Query: 234 SSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGK 293
+SFF+NVGDSL VMTNGRF+SV+H VLANGF+S LSMIYFGGP L+ KIAPLP L+KG +
Sbjct: 182 NSFFVNVGDSLXVMTNGRFKSVRHXVLANGFKSXLSMIYFGGPSLNXKIAPLPCLIKGNE 361
>AW584268 similar to GP|4678586|emb GA 2-oxidase {Phaseolus coccineus},
partial (41%)
Length = 619
Score = 168 bits (426), Expect = 2e-42
Identities = 82/159 (51%), Positives = 115/159 (71%), Gaps = 5/159 (3%)
Frame = +3
Query: 1 MVLLSKQATEQYSYVKNCM-ATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVS 59
MV+LSK + VK+C +T IPVVDL+ P+AK++IVKAC+EFGFFKV+NHGV
Sbjct: 141 MVVLSKPTLNNFLLVKSCKPSTTLLNGIPVVDLADPEAKTLIVKACKEFGFFKVVNHGVP 320
Query: 60 MEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEV-- 117
+E +S LE+EAL FF P +EK++ G +PFGYG+KRIG NGDVGWVEY+LL TN +V
Sbjct: 321 LEFMSNLENEALRFFKKPQSEKDRAGPPDPFGYGSKRIGSNGDVGWVEYILLNTNPDVIS 500
Query: 118 --NVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEG 154
++ + +N + R ++DY+ A++ M C +L+LMA+G
Sbjct: 501 NKSLSFYRENRQNLRSAVEDYIAAMKKMCCLVLELMADG 617
>TC85651 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis thaliana,
partial (39%)
Length = 1269
Score = 147 bits (371), Expect = 6e-36
Identities = 97/325 (29%), Positives = 167/325 (50%), Gaps = 16/325 (4%)
Frame = +3
Query: 7 QATEQYSYVKNCMATPCSFTIPVVDLSK---PDAKSI--IVKACEEFGFFKVINHGVSME 61
Q + V N + P +PV+DLSK DA + + AC+++GFF++INHGV+
Sbjct: 108 QPNQDSILVSNTTSLP---QLPVIDLSKLLCEDAIELENLDHACKDWGFFQLINHGVNPL 278
Query: 62 AISQLESEALNFFSMPVTEKEKTGLANPF--GYGNKRIGHNGD-VGWVEYLLLKTNQEVN 118
+ ++ FF++P+ EK+K G+G + + + W + +KT
Sbjct: 279 LVENIKIGVQKFFNLPIEEKKKFWQTTEEMQGFGQVYVALEEEKLRWGDMFFIKT----- 443
Query: 119 VPVHSQNPEKFRCV-------LDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQ 171
P+H+++P C+ L+ Y V + +++ M++ LKI+P +L + ++
Sbjct: 444 FPLHTRHPHLIPCIPQPFRDNLESYSLEVNKLCVTLIEFMSKALKIKPNELLD---IFEE 614
Query: 172 SDSVFRVNHYPACPEMGLDGENLIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVP 230
R+N+YP CP+ + +IG H+D ++ LL+ N GLQI +DG WI +
Sbjct: 615 GSQAMRMNYYPPCPQP----DQVIGLNPHSDAGTLTILLQVNEMEGLQIK-KDGIWIPIR 779
Query: 231 PDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLK 290
P +F +NVGD L+V TNG +RS++HR N + R+S+ F P + I P PSL+
Sbjct: 780 PLSDAFVVNVGDILEVQTNGIYRSIEHRATVNSEKERISVAAFHAPHMGGYIGPTPSLVT 959
Query: 291 GGKEESLYKEFTWFEYKNSSYGSRL 315
+ +L+K ++ N SR+
Sbjct: 960 -PQSPALFKTMPTADFLNGYIASRI 1031
>TC80530 weakly similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis
thaliana, partial (52%)
Length = 1219
Score = 141 bits (356), Expect = 3e-34
Identities = 94/325 (28%), Positives = 164/325 (49%), Gaps = 16/325 (4%)
Frame = +2
Query: 7 QATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIV-----KACEEFGFFKVINHGVSME 61
Q + V N ++P +P++D K + I AC+E+GFF++INHGV+
Sbjct: 83 QPNQDSILVSNTTSSP---QLPIIDFDKLLCEDGIELEKLDNACKEWGFFQLINHGVNPS 253
Query: 62 AISQLESEALNFFSMPVTEKEKTGLANPF--GYGNKRIG-HNGDVGWVEYLLLKTNQEVN 118
+ ++ FF +P+ EK+K G+G + + W + +KT
Sbjct: 254 LVESVKIGVQQFFHLPMEEKKKFWQTEEELQGFGQVYVALEEQKLRWGDMFYVKT----- 418
Query: 119 VPVHSQ-------NPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQ 171
P+H + P+ FR ++Y ++ + +I++ M + LKIQ N L ++
Sbjct: 419 FPLHIRLPHLIPCMPQPFRDDFENYSLELKKLCFKIIERMTKALKIQQPNELLDFF--EE 592
Query: 172 SDSVFRVNHYPACPEMGLDGENLIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVP 230
D R+N+YP CP+ + +IG H+D ++ LL+ N GLQI +DG W+ +
Sbjct: 593 GDQSIRMNYYPPCPQP----DQVIGLNPHSDASALTILLQVNEMQGLQIK-KDGMWVPIN 757
Query: 231 PDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLK 290
P ++F +N+GD L++MTNG +RS++HR AN + R+S+ F + + P PSL+
Sbjct: 758 PLPNAFVVNIGDLLEIMTNGIYRSIEHRATANSEKERISVAGFHNIQMGRDLGPAPSLVT 937
Query: 291 GGKEESLYKEFTWFEYKNSSYGSRL 315
+ +++K T EY N S++
Sbjct: 938 -PETPAMFKTITLEEYVNGYLASKI 1009
>TC78110 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis thaliana,
partial (53%)
Length = 1303
Score = 141 bits (356), Expect = 3e-34
Identities = 94/309 (30%), Positives = 156/309 (50%), Gaps = 20/309 (6%)
Frame = +2
Query: 27 IPVVDLSKPDAKSIIVK---------ACEEFGFFKVINHGVSMEAISQLESEALNFFSMP 77
+PV+D SK ++ + +K AC+E+GFF++INHGVS + ++ A F+++P
Sbjct: 209 LPVIDFSKLFSQDLTIKGLELDKLHSACKEWGFFQLINHGVSTSLVENVKMGAKEFYNLP 388
Query: 78 VTEKEKTGL--ANPFGYGNKRI-GHNGDVGWVEYLLLKTNQEVNVPVHSQNPE------- 127
+ EK+K + GYG + + W + + + +P H + P
Sbjct: 389 IEEKKKFSQKEGDVEGYGQAFVMSEEQKLDWADMFFM-----ITLPSHMRKPHLFPKLPL 553
Query: 128 KFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEM 187
FR L+ Y ++ + +I+D MA LK+ K + QS R+N+YP CP+
Sbjct: 554 PFRDDLETYSAELKKLAIQIIDFMANALKVDAKEIRELFGEGTQST---RINYYPPCPQP 724
Query: 188 GLDGENLIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQV 246
L +IG H+D ++ LL+ N GLQI +DG WI V P ++F IN+GD L++
Sbjct: 725 EL----VIGLNSHSDGGGLTILLQGNEMDGLQIK-KDGFWIPVKPLPNAFIINLGDMLEI 889
Query: 247 MTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEY 306
+TNG + S++HR N + RLS+ F P + + P P+L+ K +L+K ++
Sbjct: 890 ITNGIYPSIEHRATVNLKKERLSIATFYSPSSAVILRPSPTLVT-PKTPALFKPIGVTDF 1066
Query: 307 KNSSYGSRL 315
G L
Sbjct: 1067YKGYLGKEL 1093
>TC89553 weakly similar to SP|P24397|HY6H_HYONI Hyoscyamine 6-dioxygenase
(EC 1.14.11.11) (Hyoscyamine 6-beta- hydroxylase).
[Henbane], partial (49%)
Length = 1119
Score = 140 bits (354), Expect = 5e-34
Identities = 101/304 (33%), Positives = 152/304 (49%), Gaps = 11/304 (3%)
Frame = +1
Query: 19 MATPCSFTIPVVDLSKPD-AKSI--IVKACEEFGFFKVINHGVSMEAISQLESEALNFFS 75
+ P S TIP++DL D A +I I+KA EE+GFF+VINHGVS + + + + F +
Sbjct: 106 VTNPSSKTIPLIDLGGHDHAHTILQILKASEEYGFFQVINHGVSKDLMDEAFNIFQEFHA 285
Query: 76 MPVTEKEKTGLANPFGYGNK--RIGHNGDVGWVEYLLLKTNQEVN-----VPVHSQNPEK 128
MP EK +P G K N + V+Y + Q P K
Sbjct: 286 MPPKEKISECSRDPNGINCKIYASSENYKIDAVQYWKDTLTHPCPPSGEFMEFWPQKPPK 465
Query: 129 FRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMG 188
+R V+ Y + +G EIL+++ EGL ++ + + +L + V +H+P CPE
Sbjct: 466 YREVVGKYTQELNKLGHEILEMLCEGLGLKLEYFIGEL----SENPVILGHHFPPCPEPS 633
Query: 189 LDGENLIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVM 247
L +G +H DP II+ LL+ GLQI L+D WI V P ++ +N+G LQ++
Sbjct: 634 LT----LGLAKHRDPTIITILLQDQEVHGLQI-LKDDEWIPVEPIPNALVVNIGLILQII 798
Query: 248 TNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYK 307
TNGR +HRV+ N R S+ YF P S I P L+ G +YK ++ E++
Sbjct: 799 TNGRLVGAEHRVVTNSKSVRTSVAYFIYPSFSRMIEPAKELV-DGNNPPIYKSMSFGEFR 975
Query: 308 NSSY 311
Y
Sbjct: 976 KKFY 987
>TC89200 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis thaliana,
partial (47%)
Length = 1323
Score = 140 bits (352), Expect = 9e-34
Identities = 88/287 (30%), Positives = 149/287 (51%), Gaps = 17/287 (5%)
Frame = +3
Query: 20 ATPCSFTIPVVDLSK----PDAKSI--IVKACEEFGFFKVINHGVSMEAISQLESEALNF 73
+T S +P++DL+K D + AC+E+GFF++INHGV + + + F
Sbjct: 261 STISSQQVPIIDLNKLLSEEDGTELQKFDLACKEWGFFQLINHGVDLSLVENFKKHVQEF 440
Query: 74 FSMPVTEKEKTGL--ANPFGYGNKRI-GHNGDVGWVEYLLLKTNQEVNVPVHSQNP---- 126
F++ EK+ G + G+G + + W + + V +P + +NP
Sbjct: 441 FNLSAEEKKIFGQKPGDMEGFGQMFVVSEEHKLEWADLFYI-----VTLPTYIRNPNLFP 605
Query: 127 ---EKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPA 183
+ FR L+ Y ++ + I++ MA+ LKIQP +L ++ R+N+YP
Sbjct: 606 SIPQPFRDNLEMYAIEIKKLCVTIVEFMAKALKIQPNELLDFF---EEGSQAMRMNYYPP 776
Query: 184 CPEMGLDGENLIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGD 242
CP+ E +IG H+D ++ LL+ N GLQ+ +DG WI + P ++F IN+GD
Sbjct: 777 CPQP----EQVIGLNPHSDVGALTILLQLNEIEGLQVR-KDGMWIPIKPLSNAFVINIGD 941
Query: 243 SLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLL 289
L++MTNG +RS++HR N + R+S+ F L+ + P PSL+
Sbjct: 942 MLEIMTNGIYRSIEHRATINAEKERISIATFNSGRLNAILCPAPSLV 1082
>TC78010 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis thaliana,
partial (47%)
Length = 1321
Score = 140 bits (352), Expect = 9e-34
Identities = 92/277 (33%), Positives = 151/277 (54%), Gaps = 14/277 (5%)
Frame = +3
Query: 27 IPVVDLSK----PDAKSI--IVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTE 80
+PV+DLSK D + + AC+E+GFF++INHGV+ + + +FF++PV E
Sbjct: 219 VPVIDLSKLLSEEDETELQKLDHACKEWGFFQLINHGVNPLLVENFKKLVQDFFNLPVEE 398
Query: 81 K----EKTGLANPFGYGNKRI-GHNGDVGWVEYLLLKTNQEV--NVPVHSQNPEKFRCVL 133
K +K G N G+G + + + W + + T+ N + P+ FR L
Sbjct: 399 KKILSQKPG--NIEGFGQLFVVSEDHKLEWADLFHIITHPSYMRNPQLFPSIPQPFRESL 572
Query: 134 DDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGEN 193
+ Y ++ + I++ M++ LKIQ +L QS R+N+YP CP+ +
Sbjct: 573 EMYSLVLKKLCVMIIEFMSKALKIQKNELLEFFEEGGQS---MRMNYYPPCPQP----DK 731
Query: 194 LIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRF 252
+IG H+D ++ LL+ N GLQI +DG WI + P ++F IN+GD L++MTNG +
Sbjct: 732 VIGLNPHSDGTALTILLQLNEIEGLQIK-KDGMWIPIKPLTNAFVINIGDMLEIMTNGIY 908
Query: 253 RSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLL 289
RS++HR N + R+S+ F L+ +AP+PSL+
Sbjct: 909 RSIEHRATINSEKERISIATFHSARLNAILAPVPSLI 1019
>TC85652 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis thaliana,
partial (50%)
Length = 1489
Score = 139 bits (350), Expect = 2e-33
Identities = 97/305 (31%), Positives = 159/305 (51%), Gaps = 16/305 (5%)
Frame = +1
Query: 27 IPVVDLSK---PDAKSI--IVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK 81
+PV+DL K DA + + AC+E+GFF++INHGV+ I +++ F S+PV EK
Sbjct: 400 VPVIDLGKLLGEDATELEKLDLACKEWGFFQIINHGVNTSLIEKVKIGIKEFLSLPVEEK 579
Query: 82 EKTGLA--NPFGYGNKRI-GHNGDVGWVEYLLLKTNQEVNVPVHSQNP-------EKFRC 131
+K + G+G + + + W + L+ T +P+ +NP + FR
Sbjct: 580 KKFWQTPNDMEGFGQMFVVSDDQKLEWADLFLITT-----LPLDERNPRLFPSIFQPFRD 744
Query: 132 VLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDG 191
L+ Y V+ + I+ M + LKI+ V + R N YP CP+
Sbjct: 745 NLEIYCSEVQKLCFTIISQMEKALKIESNEVTE---LFNHITQAMRWNLYPPCPQP---- 903
Query: 192 ENLIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNG 250
EN+IG H+D ++ LL++N GLQI +DG WI V P ++F IN+GD L+++TNG
Sbjct: 904 ENVIGLNPHSDVGALTILLQANEIEGLQIR-KDGQWIPVQPLPNAFVINIGDMLEIVTNG 1080
Query: 251 RFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSS 310
+RS++HR N + R+S+ F P ++ ++P PSL+ + S +K EY +
Sbjct: 1081IYRSIEHRATVNSEKERISIAAFHRPHVNTILSPRPSLVTPERPAS-FKSIAVGEYLKAY 1257
Query: 311 YGSRL 315
+ +L
Sbjct: 1258FSRKL 1272
>TC87721 weakly similar to PIR|H96594|H96594 hypothetical protein F7A10.24
[imported] - Arabidopsis thaliana, partial (50%)
Length = 1365
Score = 138 bits (348), Expect = 3e-33
Identities = 93/303 (30%), Positives = 157/303 (51%), Gaps = 20/303 (6%)
Frame = +1
Query: 26 TIPVVDLSK---PDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK- 81
+IP++D + PD + I A + GFF+++NHG+ + + L++ FF +PV EK
Sbjct: 361 SIPIIDFTNWDDPDVQDSIFSAATKLGFFQIVNHGIPINVLDDLKASVHKFFELPVEEKK 540
Query: 82 --------EKTGLANPFGYGNKRIGHNGDVGWVEYL-LLKTNQEVNVPVHSQNPEKFRCV 132
E LA F + + + W +YL L+ T++E +H+ P +
Sbjct: 541 SVKENSPPEVVRLATSFSPHAESV-----LEWKDYLQLVYTSEE---KIHAYWPVVCKNQ 696
Query: 133 LDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSD------SVFRVNHYPACPE 186
+Y+ ++L ++ + L ++ +DK+ + + N+YPACPE
Sbjct: 697 ALEYMKYADAFIRKLLKVLLKKLNVKE--------LDKEREYALMGAMILGFNYYPACPE 852
Query: 187 MGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDG-SWISVPPDYSSFFINVGDSLQ 245
L G G H+D I++L ++ GL + +DG SWI+VPP + IN+GD LQ
Sbjct: 853 PELGS----GVGPHSDISSITVLLQDDIGGLYVRGKDGDSWINVPPVNGALVINIGDVLQ 1020
Query: 246 VMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFE 305
+M+NGR++S++HRV+ +G ++R+SM F P I LP +L+ G EE YK+ + E
Sbjct: 1021IMSNGRYKSIEHRVVVDGNKTRISMPIFVNPAPDAVIGTLPEVLENG-EEPHYKQVVFSE 1197
Query: 306 YKN 308
Y N
Sbjct: 1198YFN 1206
>TC92472 similar to GP|5579092|gb|AAD45424.1| gibberellin 2-oxidase-like
protein {Pisum sativum}, partial (33%)
Length = 671
Score = 137 bits (344), Expect = 7e-33
Identities = 65/130 (50%), Positives = 90/130 (69%)
Frame = +3
Query: 194 LIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFR 253
++ +GEH+DP I+++LRSN+ S LQISL+ G WI V PD + IN+GD L+VMTNGRF
Sbjct: 63 MLCYGEHSDPHILTILRSNDVSALQISLQHGLWIPVNPDPEALCINIGDVLEVMTNGRFV 242
Query: 254 SVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGS 313
SV+HR + N ++SR+SM YFG PPL+ I P +L SL++ FTW +YK ++Y
Sbjct: 243 SVRHRAMTNSYKSRMSMAYFGAPPLNASIV-APPVLVTPTRPSLFRPFTWADYKKATYSL 419
Query: 314 RLADNRLGHF 323
RL D R+ F
Sbjct: 420 RLGDTRIQLF 449
>TC78109 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis thaliana,
partial (48%)
Length = 1314
Score = 102 bits (254), Expect(2) = 2e-32
Identities = 74/221 (33%), Positives = 112/221 (50%), Gaps = 20/221 (9%)
Frame = +3
Query: 27 IPVVDLSK---------PDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMP 77
+PV+DLSK P+ K + AC+E+GFF++INHGVS + ++ A FF++P
Sbjct: 192 LPVIDLSKLLSSHDLNEPELKKLHY-ACKEWGFFQLINHGVSESLMENVKKGAEEFFNLP 368
Query: 78 VTEKEKTGL--ANPFGYGNK-RIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPE------- 127
+ EK+K G + GYG + + W + L+L T +P H + P
Sbjct: 369 MEEKKKFGQTEGDVEGYGQAFVVSEEQKLDWADMLVLFT-----LPPHKRKPHLFSNIPL 533
Query: 128 KFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEM 187
FR L++Y +R + +I+DLMA L + P + QS R+N+YP CP+
Sbjct: 534 PFRVDLENYCEKMRTLAIQIMDLMAHSLAVDPMEIREFFGQATQST---RMNYYPPCPQ- 701
Query: 188 GLDGENLIGFGEHTD-PQIISLLRSNNTSGLQISLRDGSWI 227
E +IG H+D + LL+ N GLQI +DG WI
Sbjct: 702 ---PEFVIGLNSHSDGGGLTILLQGNEMDGLQIK-KDGLWI 812
Score = 54.7 bits (130), Expect(2) = 2e-32
Identities = 25/61 (40%), Positives = 39/61 (62%)
Frame = +1
Query: 229 VPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSL 288
V P ++F IN+GD L++MTNG FRS++HR N + R S+ F P + ++P PS+
Sbjct: 817 VKPLPNAFIINLGDMLEMMTNGIFRSIEHRATVNSEKERFSIASFYSPAFNTILSPAPSI 996
Query: 289 L 289
+
Sbjct: 997 V 999
>TC88553 weakly similar to GP|21327037|gb|AAM48133.1 putative flavanone
3-hydroxylase {Saussurea medusa}, partial (26%)
Length = 1017
Score = 135 bits (341), Expect = 2e-32
Identities = 89/267 (33%), Positives = 141/267 (52%), Gaps = 16/267 (5%)
Frame = +1
Query: 22 PCSFTIPVVDLS---KPDAKSII---VKACEEFGFFKVINHGVSMEAISQLESEALNFFS 75
P S +IPV+DLS K D +II +KA EEFGFF+VINHG+SM+ + + S F
Sbjct: 250 PFSNSIPVIDLSEACKGDRSNIIKKIIKAAEEFGFFQVINHGISMDQMKETMSIFKEIFH 429
Query: 76 MPVTEKEKTGLANP----------FGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQN 125
MP K+ +P F Y +++ D L+ Q + +N
Sbjct: 430 MPDEYKKNLCTNDPSKPCKMFTSSFNYATEKVHLWRDSLRHPCYPLEQWQHL----WPEN 597
Query: 126 PEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACP 185
P +R + D+ V+ +G +++L++EGL ++ D + VNHYP CP
Sbjct: 598 PTTYRECVGDFSNEVKELGSRLMNLISEGLGLK----CGYFDNDLTGSMILSVNHYPRCP 765
Query: 186 EMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQ 245
E L +G +H+DP +I++L ++ SGLQ+ L+DG WI++ +F INVG LQ
Sbjct: 766 EPNL----ALGMSKHSDPNLITILMQDDVSGLQV-LKDGKWIALEARPHAFIINVGYQLQ 930
Query: 246 VMTNGRFRSVKHRVLANGFQSRLSMIY 272
+++N + +SV+HRV+ N Q R + +
Sbjct: 931 IISNDKLKSVEHRVVTNASQGRTTAAF 1011
>TC79860 homologue to PIR|T06533|T06533 probable gibberellin 20-oxidase -
garden pea, partial (93%)
Length = 1454
Score = 134 bits (337), Expect = 5e-32
Identities = 101/333 (30%), Positives = 156/333 (46%), Gaps = 23/333 (6%)
Frame = +3
Query: 16 KNCMATPCSFTIPVVDLS---KPDAKSI------IVKACEEFGFFKVINHGVSMEAISQL 66
K C+ TP +P +DL D K+I + AC++ GFF V+NHGV + ++Q
Sbjct: 153 KPCL-TPPKLQVPPIDLKAFLSGDPKAISNACSQVNDACKKHGFFLVVNHGVDKKLLAQA 329
Query: 67 ESEALNFFSMPVTEKEKTG--LANPFGYGNKRIGH-NGDVGWVEYLLLK--------TNQ 115
+FF M + EK+K + GY N IG + + W E L + T +
Sbjct: 330 HKLVDDFFCMQLCEKQKAQRKVGEHCGYANSFIGRFSSKLPWKETLSFRYSDDKSCRTVE 509
Query: 116 EVNVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSV 175
+ V V ++ +F V DY A+ N+ I++L+ L + + + +DSV
Sbjct: 510 DYFVNVMGEDFRQFGSVYQDYCEAMSNLSLGIMELLGMSLGVDKEYFRHFF---EANDSV 680
Query: 176 FRVNHYPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSS 235
R+N+YP C L +G G H DP +++L + GLQ+ L DG W S+ P +
Sbjct: 681 MRLNYYPPCKNPDL----ALGTGPHCDPTSLTILHQDQVEGLQV-LVDGIWHSIVPKEDA 845
Query: 236 FFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEE 295
F +N+GD+ ++NG F+S HR + N R S+ +F P EKI P L +
Sbjct: 846 FVVNIGDTFMALSNGIFKSCLHRAVVNDTIVRKSLAFFLCPN-EEKIVTPPKELINKENP 1022
Query: 296 SLYKEFTW---FEYKNSSYGSRLADNRLGHFER 325
+Y FTW E+ Y R + L F R
Sbjct: 1023MIYPNFTWPSLLEFTQKHY--RADERTLDAFSR 1115
>TC82730 similar to GP|10177853|dbj|BAB11205. flavanone 3-hydroxylase-like
protein {Arabidopsis thaliana}, partial (81%)
Length = 1406
Score = 87.8 bits (216), Expect(2) = 1e-31
Identities = 50/130 (38%), Positives = 76/130 (58%), Gaps = 1/130 (0%)
Frame = +1
Query: 178 VNHYPACPEMGLDGENLIGFGEHTDPQIIS-LLRSNNTSGLQISLRDGSWISVPPDYSSF 236
VN+YP CP+ L G HTDP ++ LL+ + +GLQ+ L+DG W+++ P +F
Sbjct: 682 VNYYPPCPQPELT----YGLPGHTDPNALTILLQDLHVAGLQV-LKDGKWLAINPIPDAF 846
Query: 237 FINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEES 296
IN+GD LQ ++NG ++SV HR + N + RLS+ F P I P L + G +
Sbjct: 847 VINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDG-SGA 1023
Query: 297 LYKEFTWFEY 306
+Y+ FT+ EY
Sbjct: 1024VYRGFTYPEY 1053
Score = 66.6 bits (161), Expect(2) = 1e-31
Identities = 50/162 (30%), Positives = 78/162 (47%), Gaps = 14/162 (8%)
Frame = +3
Query: 18 CMATPCSF-TIPVVDLSKPDAKSIIVK---ACEEFGFFKVINHGVSMEAISQLESEALNF 73
C++ F +P++DL + I+ + AC +GFF+V+NHGV +E + + A +F
Sbjct: 183 CLSQVSEFENVPIIDLGSHNRTQIVQQIGEACSSYGFFQVVNHGVPLEELKKTAEVAYDF 362
Query: 74 FSMPVTEKEK---------TGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVN-VPVHS 123
F +PV EK K L+ F NK HN W +YL L N VP
Sbjct: 363 FKLPVEEKMKLYSDDPTKTMRLSTSFNV-NKEEVHN----WRDYLRLHCYPLDNYVPEWP 527
Query: 124 QNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSK 165
NP F+ + +Y VR +G I + ++ +++ PK L K
Sbjct: 528 SNPTSFKETVANYCKEVRELGLRIEEYIS--VELGPKKRLLK 647
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,185,774
Number of Sequences: 36976
Number of extensions: 138479
Number of successful extensions: 956
Number of sequences better than 10.0: 168
Number of HSP's better than 10.0 without gapping: 800
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 807
length of query: 329
length of database: 9,014,727
effective HSP length: 96
effective length of query: 233
effective length of database: 5,465,031
effective search space: 1273352223
effective search space used: 1273352223
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0190.3


