

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0185.17
(469 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
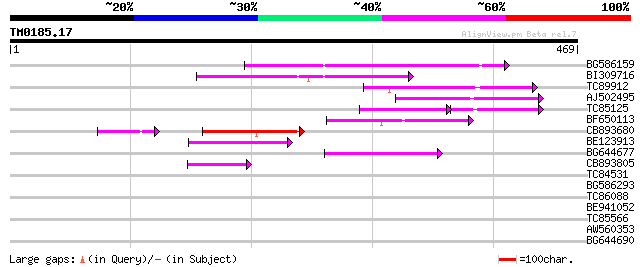
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 126 2e-29
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 100 1e-21
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 82 4e-16
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 78 6e-15
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 58 2e-14
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 74 1e-13
CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotia... 50 3e-09
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 52 6e-07
BG644677 weakly similar to GP|7682800|gb| Hypothetical protein T... 51 1e-06
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 45 5e-05
TC84531 37 0.021
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 35 0.047
TC86088 calmodulin-like protein 1 [Medicago truncatula] 29 3.4
BE941052 weakly similar to PIR|B85188|B85 retrotransposon like p... 29 3.4
TC85566 homologue to SP|P27880|HS12_MEDSA 18.2 kDa class I heat ... 29 4.4
AW560353 similar to PIR|T00660|T006 hypothetical protein F3I6.23... 28 9.9
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 28 9.9
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 126 bits (316), Expect = 2e-29
Identities = 77/221 (34%), Positives = 122/221 (54%), Gaps = 2/221 (0%)
Frame = +1
Query: 195 IKIIRDGEIIGLSQSHYIEKVLKKFDHFDCKSVSTPFDQNTKL-QPHKRCPVAQLEYSKA 253
+++I++ E I + Q Y+ +L++F P KL + V +Y +
Sbjct: 1 VEVIQNEEGIYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQI 180
Query: 254 IRCLMYAMTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV 313
+ CLMY + TRPD+ YV+ +SR+ P++ H H V RVL+YLNGTIN ++Y S
Sbjct: 181 VGCLMY-LAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMYKRNGSE 357
Query: 314 -LEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKE 372
LE YT + + + D STSG +F L GAVSW SKKQ + ST AEFIA A C+ +
Sbjct: 358 KLEAYTDSDYAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAAFCACQ 537
Query: 373 TV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYSQVYNG 413
+V +R +L ++ ++++CD+ T+ + + V +G
Sbjct: 538 SVWMRRVLEKLGYTQS--GSITMYCDNNSTIKLSKNPVLHG 654
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 100 bits (249), Expect = 1e-21
Identities = 59/183 (32%), Positives = 101/183 (54%), Gaps = 3/183 (1%)
Frame = +2
Query: 155 LYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQSHYIEK 214
+YVDD+++ G D++E++ K FL + F +KDLG LG+++ R + I L+Q Y +
Sbjct: 179 VYVDDIVLAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQGILLNQRKYTLE 358
Query: 215 VLKKFDHFDCKSVSTPFDQNTKLQPHKRCPV--AQLEYSKAIRCLMYAMTCTRPDIAYVV 272
+L+ + KS TP+D + KL + P+ + +Y + I L+Y +T TRPDI++ V
Sbjct: 359 LLEDSGNLAVKSTLTPYDISLKLH-NSDSPLYNDETQYRRLIGKLIY-LTTTRPDISFAV 532
Query: 273 GRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV-LEGYTYASWVTCVKDHAS 331
+LS++ P + H+ RVL+YL L Y+ ++ L + + W TC S
Sbjct: 533 QQLSQFVSKPQQVHYQAAIRVLQYLKTAPAKGLFYSATSNLKLSSFADSDWATCPTTRKS 712
Query: 332 TSG 334
+G
Sbjct: 713 VTG 721
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 82.0 bits (201), Expect = 4e-16
Identities = 51/147 (34%), Positives = 78/147 (52%), Gaps = 3/147 (2%)
Frame = +1
Query: 293 VLKYLNGTINYSLIYNGYPS---VLEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSK 349
VLKYLN ++ SL Y LEGY A + V S SG +F L G +SW +
Sbjct: 16 VLKYLNESLKSSLKYTKAAQEEDALEGYVDADYAGNVDTRKSLSGFVFTLYGTTISWKAN 195
Query: 350 KQTCIADSTMAAEFIALAACSKETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYSQ 409
+Q+ + ST AE+IA K+ + L+ ++ E+ I V IHCDSQ + A Q
Sbjct: 196 QQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELGI---TQEYVKIHCDSQSAIHLANHQ 366
Query: 410 VYNGKSRHIVLRHSHVKDLITNGVISI 436
VY+ +++HI +R ++D+I + I +
Sbjct: 367 VYHERTKHIDIRLHFIRDMIESKEIVV 447
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 78.2 bits (191), Expect = 6e-15
Identities = 41/122 (33%), Positives = 72/122 (58%)
Frame = +2
Query: 320 ASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNL 379
+ W + STSG F+LG GA+SW SKKQ +A ST AE+IA +C+ +TV LR +
Sbjct: 2 SDWAGDTETRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRI 181
Query: 380 LYEILIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFV 439
L ++ + I+CD++ ++ + + V++G+S+HI ++ +++LI + I +
Sbjct: 182 LE--VMHHEQNTPTKIYCDNKSAIALSKNPVFHGRSKHIDIQFHKIRELIAEKEVVIEYC 355
Query: 440 RT 441
T
Sbjct: 356 PT 361
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 58.2 bits (139), Expect(2) = 2e-14
Identities = 29/76 (38%), Positives = 44/76 (57%)
Frame = +1
Query: 290 VNRVLKYLNGTINYSLIYNGYPSVLEGYTYASWVTCVKDHASTSG*IFNLGGGAVSWGSK 349
V R+++Y+ GT ++ + G + GY + + ST+G +F L GGAVSW SK
Sbjct: 1 VKRIMRYIKGTSGVAVCFGGSELTVRGYVDSDFAGDHDKRKSTTGYVFTLAGGAVSWLSK 180
Query: 350 KQTCIADSTMAAEFIA 365
QT +A ST AE++A
Sbjct: 181 LQTVVALSTTEAEYMA 228
Score = 38.5 bits (88), Expect(2) = 2e-14
Identities = 17/77 (22%), Positives = 43/77 (55%)
Frame = +3
Query: 365 ALAACSKETV*LRNLLYEILIWPKLMRQVSIHCDSQCTLSKAYSQVYNGKSRHIVLRHSH 424
+L KE + ++ L+ E+ Q++++CDSQ L A + ++ +++HI +++
Sbjct: 228 SLPQACKEAIWMQRLMEEL---GHKQEQITVYCDSQSALHIARNPAFHSRTKHIGIQYHF 398
Query: 425 VKDLITNGVISIVFVRT 441
V++++ G + + + T
Sbjct: 399 VREVVEEGSVDMQKIHT 449
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 73.9 bits (180), Expect = 1e-13
Identities = 44/123 (35%), Positives = 66/123 (52%), Gaps = 2/123 (1%)
Frame = +1
Query: 263 CTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLI--YNGYPSVLEGYTYA 320
C RPDI Y V +S++ +P K H NR+L+Y+ GT+ Y L+ Y V E Y+
Sbjct: 115 C*RPDICYSVSVISKFMHDPRKPHLIAANRILRYVRGTMEYGLLFPYGAKSEVYELICYS 294
Query: 321 SWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADSTMAAEFIALAACSKETV*LRNLL 380
C D STSG +F A+SW +KKQ A S+ AE+IA + + + L +++
Sbjct: 295 DSDWC-GDRRSTSGYVFKFNDAAISWCTKKQPITALSSYEAEYIAGTFATFQALWLDSVI 471
Query: 381 YEI 383
E+
Sbjct: 472 KEL 480
>CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotiana tabacum},
partial (7%)
Length = 780
Score = 49.7 bits (117), Expect(2) = 3e-09
Identities = 30/87 (34%), Positives = 53/87 (60%), Gaps = 2/87 (2%)
Frame = -3
Query: 160 MLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGE--IIGLSQSHYIEKVLK 217
+L+ G++++E++ K+ S DMKDLG A I+G++I+ D + ++ LSQ YI +VL+
Sbjct: 271 LLVVGSNIDEIKNLKTRFSKEIDMKDLGPAKKIIGMQIMIDKQKGVL*LSQVEYITRVLQ 92
Query: 218 KFDHFDCKSVSTPFDQNTKLQPHKRCP 244
F+ + VST + L H++ P
Sbjct: 91 IFNMGNAILVSTTLASHFCLS-HEQSP 14
Score = 29.3 bits (64), Expect(2) = 3e-09
Identities = 16/52 (30%), Positives = 25/52 (47%)
Frame = -1
Query: 73 GC*DCFPKWRVG*GCVYETTWRVCYQRPRT*SV*ID*VVIWVKTSTQTMASK 124
GC DC WR+ *G ++ T R+ + + +W KT ++TM K
Sbjct: 474 GCEDCISSWRLS*GYIHAPT*RILIRSGENGGKTKE-EHVWTKTRSKTMYMK 322
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 51.6 bits (122), Expect = 6e-07
Identities = 25/86 (29%), Positives = 47/86 (54%)
Frame = +1
Query: 149 KAVMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRDGEIIGLSQ 208
K + +YVDD+ + G +++ K+ L+ F++KDLG LG+++ R + +SQ
Sbjct: 109 KKAILIVYVDDIFLTGDHGK*IKRLKNLLAEEFEIKDLGNLKYFLGMEVARWKKGSSISQ 288
Query: 209 SHYIEKVLKKFDHFDCKSVSTPFDQN 234
Y+ +LK+ CK++ P+ N
Sbjct: 289 RKYVLDLLKETRMIGCKTIRDPYGCN 366
>BG644677 weakly similar to GP|7682800|gb| Hypothetical protein T15F17.l
{Arabidopsis thaliana}, partial (3%)
Length = 539
Score = 50.8 bits (120), Expect = 1e-06
Identities = 33/99 (33%), Positives = 51/99 (51%), Gaps = 1/99 (1%)
Frame = -3
Query: 261 MTCTRPDIAYVVGRLSRYTCNPSKDHWHVVNRVLKYLNGTINYSLIYNGYPSV-LEGYTY 319
+T P+I + + LSRY+ P+ H + + + KYL G I+ L Y+ S L GY
Sbjct: 525 LTLQGPNITFSINLLSRYSSAPTMRH*NGIKHICKYLKGIIDMGLFYSKDCSPDLIGYVN 346
Query: 320 ASWVTCVKDHASTSG*IFNLGGGAVSWGSKKQTCIADST 358
A +++ S +G IF G +SW S K + IA S+
Sbjct: 345 A*YLSDPHKARS*TGYIFTCGNTVISWRSTK*STIATSS 229
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 45.4 bits (106), Expect = 5e-05
Identities = 19/53 (35%), Positives = 32/53 (59%)
Frame = +3
Query: 148 GKAVMTCLYVDDMLIFGTDLNEVEKTKSFLSNSFDMKDLGEADVILGIKIIRD 200
GK ++ LYVDD++ G D N E+ K + F+M DLG+ LG+++ ++
Sbjct: 597 GKILIISLYVDDLIFIGNDENMFEEFKKSMKKEFNMSDLGKMHYFLGVEVTQN 755
>TC84531
Length = 655
Score = 36.6 bits (83), Expect = 0.021
Identities = 14/34 (41%), Positives = 24/34 (70%)
Frame = -3
Query: 408 SQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRT 441
++ YNGK R I +HS +++ ++NG + + FVRT
Sbjct: 647 NRYYNGKRRQIRRKHSTIREYLSNGTVRVDFVRT 546
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 35.4 bits (80), Expect = 0.047
Identities = 23/80 (28%), Positives = 41/80 (50%)
Frame = +1
Query: 2 KIQTNRLQMDL*KENDSCWYH**I*S*TCC*KIYTKGRY*LF*YLCTCCKNCIN*SVVGT 61
+ +T+R ++DL* + W I S C ++ R+ L +CT C N + + +G
Sbjct: 82 RCETDRFEVDL*D*EE*RWNVDQIQSKASCKRLRETTRHRLRRSVCTSCSNRNHMTTLGV 261
Query: 62 SFSL*VCNP*NGC*DCFPKW 81
S + + +P + C +C PKW
Sbjct: 262 SSN*WMLDPSHRCKNCIPKW 321
>TC86088 calmodulin-like protein 1 [Medicago truncatula]
Length = 867
Score = 29.3 bits (64), Expect = 3.4
Identities = 20/63 (31%), Positives = 30/63 (46%), Gaps = 5/63 (7%)
Frame = -1
Query: 248 LEYSKAIRCLMYAMTCTRPDIAYVVGRLSRYTCNPS----KDHWHVVNRVLK-YLNGTIN 302
L Y I + +C R ++Y + + C PS H H +++LK YL+ TIN
Sbjct: 693 LSYLSHIIS*ILKFSCIRVPVSYRTRTRTMFICTPSDHNGSTHGHSFDKLLKVYLSITIN 514
Query: 303 YSL 305
SL
Sbjct: 513 ISL 505
>BE941052 weakly similar to PIR|B85188|B85 retrotransposon like protein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 480
Score = 29.3 bits (64), Expect = 3.4
Identities = 13/50 (26%), Positives = 25/50 (50%)
Frame = +2
Query: 395 IHCDSQCTLSKAYSQVYNGKSRHIVLRHSHVKDLITNGVISIVFVRTVKE 444
+ CD ++ VY+ + +HI + V+DL+ G + + V TV +
Sbjct: 23 LRCDYLSATYLTHNPVYHSRMKHISIDIHFVRDLVQQGKLKVQHVCTVDQ 172
>TC85566 homologue to SP|P27880|HS12_MEDSA 18.2 kDa class I heat shock
protein. [Alfalfa] {Medicago sativa}, complete
Length = 946
Score = 28.9 bits (63), Expect = 4.4
Identities = 13/46 (28%), Positives = 27/46 (58%)
Frame = -1
Query: 193 LGIKIIRDGEIIGLSQSHYIEKVLKKFDHFDCKSVSTPFDQNTKLQ 238
LG+++ +D + + + Q Y +++K DCK ++TP + KL+
Sbjct: 811 LGMEVRQDNK*VLICQMKYTREIMK-----DCKRINTPVNLKEKLE 689
>AW560353 similar to PIR|T00660|T006 hypothetical protein F3I6.23 -
Arabidopsis thaliana, partial (16%)
Length = 623
Score = 27.7 bits (60), Expect = 9.9
Identities = 17/39 (43%), Positives = 22/39 (55%), Gaps = 2/39 (5%)
Frame = +3
Query: 394 SIHCDSQCTLSKAYSQVYNGKSR-HIVLRHSH-VKDLIT 430
S HC C ++ SQV N + HI LRHS+ + LIT
Sbjct: 495 SSHCSINCYRFRSQSQVSNTLCKNHITLRHSYKMTSLIT 611
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 27.7 bits (60), Expect = 9.9
Identities = 29/92 (31%), Positives = 41/92 (44%)
Frame = -1
Query: 33 KIYTKGRY*LF*YLCTCCKNCIN*SVVGTSFSL*VCNP*NGC*DCFPKWRVG*GCVYETT 92
+I +K R L * TCC+N *+ V NGC +C WR G V + T
Sbjct: 392 RIQSKRRNRL**GFFTCCQNGSY*NFNSFCCIHGVQAVPNGCEECIY*WRSQRGGVCQAT 213
Query: 93 WRVCYQRPRT*SV*ID*VVIWVKTSTQTMASK 124
+ R V I+* IW + S+++M K
Sbjct: 212 SWI*RCRGTKSCVQIE*DTIWSEASSKSMV*K 117
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.351 0.154 0.546
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,875,052
Number of Sequences: 36976
Number of extensions: 246130
Number of successful extensions: 2034
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 1488
Number of HSP's successfully gapped in prelim test: 65
Number of HSP's that attempted gapping in prelim test: 523
Number of HSP's gapped (non-prelim): 1578
length of query: 469
length of database: 9,014,727
effective HSP length: 100
effective length of query: 369
effective length of database: 5,317,127
effective search space: 1962019863
effective search space used: 1962019863
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0185.17


