

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.7
(1813 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
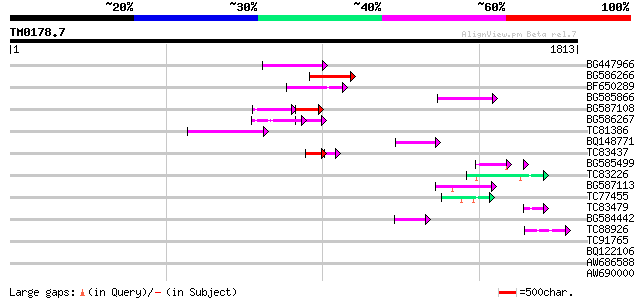
Score E
Sequences producing significant alignments: (bits) Value
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 152 1e-36
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 139 1e-32
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 132 2e-30
BG585866 122 2e-27
BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - ... 74 1e-22
BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At... 96 1e-19
TC81386 similar to GP|10140689|gb|AAG13524.1 putative non-LTR re... 91 5e-18
BQ148771 90 7e-18
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 62 8e-16
BG585499 59 6e-09
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 60 8e-09
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 57 5e-08
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 52 3e-06
TC83479 similar to GP|23476992|emb|CAD48949. hypothetical protei... 49 2e-05
BG584442 49 2e-05
TC88926 similar to GP|7290507|gb|AAF45960.1| peb gene product {D... 46 1e-04
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 40 0.006
BQ122106 similar to GP|9366656|emb|C probable similar to ring-h2... 40 0.006
AW686588 40 0.008
AW690000 40 0.008
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 152 bits (383), Expect = 1e-36
Identities = 79/208 (37%), Positives = 119/208 (56%)
Frame = +2
Query: 808 DRNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDRQLLAEEIQNYFQSLFAADPDNTDVNM 867
D+NT +FH+ RRK N+I +LK G+WC + + + YF +LF +
Sbjct: 20 DKNTKFFHSKASQRRKVNEIKKLKDETGNWCKGEENVERLLITYFNNLFTSSNPTAIEET 199
Query: 868 QHVMMPRLTEAERHKAEAEVIKEEVFQALLRMRSYSALGPDGFHPFFFKKYWELVGDDLW 927
V+ +L+ E E +EEV +A+ +M A GPDG FF+KYW +VG ++
Sbjct: 200 CEVVKGKLSHEHIVWCEKEFTEEEVLEAINQMHPVKAPGPDGLPALFFQKYWHIVGKEVQ 379
Query: 928 KLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVVVNRIRPL 987
++V M + L +T IVLIPK +P+T K++RPISLCNVV K+ITKV+ NR++
Sbjct: 380 QMVLQVLNNSMETEELNKTFIVLIPKGKNPNTPKDYRPISLCNVVMKIITKVIANRVKQT 559
Query: 988 LNRIIGPMQNSFLPGMGIMDNAFLAQEI 1015
L +I Q++F+ G I DNA +A +
Sbjct: 560 LPDVIDVEQSAFVQGRLITDNALIAWSV 643
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 139 bits (350), Expect = 1e-32
Identities = 68/148 (45%), Positives = 95/148 (63%), Gaps = 1/148 (0%)
Frame = -3
Query: 960 VKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHM 1019
V E+R I+ CN YK+I K++ R++PLL II P Q++F+PG I DN + +I+H++
Sbjct: 778 VSEYRTIAPCNTQYKIIAKILSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKILHYL 599
Query: 1020 GASMAKKG-TLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWN 1078
S AKK ++A K D+ KAYD ++W FL + L + GF + IM CVS + S L N
Sbjct: 598 RQSGAKKHVSMAVKTDMTKAYDRIAWNFLREVLTRLGFHGIWISWIMECVSTVSYSFLIN 419
Query: 1079 GSRLPSFKPGRGLRKGDPLSPYLFVLCS 1106
G P RGLR+GDPLSPYLF+LC+
Sbjct: 418 GGPQGRVLPSRGLRQGDPLSPYLFILCT 335
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 132 bits (331), Expect = 2e-30
Identities = 69/198 (34%), Positives = 117/198 (58%), Gaps = 3/198 (1%)
Frame = +3
Query: 885 AEVIKEEVFQALLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLL 944
+E EV AL M S A G DG++ FFK W ++GD + + D F+TG + +
Sbjct: 18 SEFTAVEVKNALFSMDSSKAPGIDGYNVHFFKCSWNIIGDSVIDAILDFFKTGFMPKIIN 197
Query: 945 ETLIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMG 1004
T + L+PK + ++VK FRPI+ C+V+YK+I+K++ +R++ +LN ++ Q++F+ G
Sbjct: 198 CTYVTLLPKEVNVTSVKNFRPIACCSVIYKIISKILTSRMQGVLNSVVSENQSAFVKGRV 377
Query: 1005 IMDNAFLAQEIIHHMGASMAKKG---TLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLV 1061
I DN L+ E++ S ++KG KIDL KAYDS W F++ +++ GFP + V
Sbjct: 378 IFDNIILSHELV----KSYSRKGISPRCMVKIDLXKAYDSXEWPFIKHLMLELGFPYKFV 545
Query: 1062 QLIMRCVSGSNLSMLWNG 1079
+M ++ ++ + NG
Sbjct: 546 NWVMAXLTTASYTFNXNG 599
>BG585866
Length = 828
Score = 122 bits (305), Expect = 2e-27
Identities = 60/193 (31%), Positives = 95/193 (49%)
Frame = +3
Query: 1368 ILKARDQLQEGFQFRMGDGSSSVWYHDWTGKGKLAESLDFVHISDTKLCLHELVDNNQWN 1427
I++ ++ L+ G+ +R G G+SS WY +W+ G L FV I D L + ++ +
Sbjct: 84 IIRDKNVLKSGYTWRAGSGNSSFWYTNWSSLGLLGTQAPFVDIHDLHLTVKDVFTTGGQH 263
Query: 1428 LNRCMTVFPLHIRDWFARVEPRVVANGIDRWSWKDNSDGISSVHDGYQWLREKKMAGVVH 1487
T+ P I + A+ D + W NS+G+ + GY W+ + +
Sbjct: 264 TQSLYTILPTDIAEVINNTHLNFNASIGDAYIWPHNSNGVYTAKSGYSWILSQTETVNYN 443
Query: 1488 GDSWLWI*KLHAPEKL*VFIWLVLHDALPTNANRYRCKLAVSPGCTRCSSPEEDTIHCLR 1547
SW WI +L PEK F+WL H+A+PT + + S C+RC EE HC+R
Sbjct: 444 NSSWSWIWRLKIPEKYKFFLWLACHNAVPTLSLLNHRNMVNSAICSRCGEHEESFFHCVR 623
Query: 1548 DCPHSQELWGKLG 1560
DC S+ +W K+G
Sbjct: 624 DCRFSKIIWHKIG 662
>BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - Arabidopsis
thaliana, partial (17%)
Length = 677
Score = 73.6 bits (179), Expect(2) = 1e-22
Identities = 37/89 (41%), Positives = 54/89 (60%)
Frame = +3
Query: 914 FFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVY 973
FF+ W ++ DL K+V +G +D L T I LIPK P+ + E RPISLCNV Y
Sbjct: 411 FFQHSWHIIKMDLLKMVNSFLASGKLDTRLNTTNICLIPKKKRPTRMTELRPISLCNVGY 590
Query: 974 KLITKVVVNRIRPLLNRIIGPMQNSFLPG 1002
K+I+KV+ R++ L +I Q++F+ G
Sbjct: 591 KIISKVLCQRLKVCLPSLISETQSAFVHG 677
Score = 53.1 bits (126), Expect(2) = 1e-22
Identities = 41/140 (29%), Positives = 58/140 (41%)
Frame = +2
Query: 776 LETELQAEYRFLLRQEELLWFQKARENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDG 835
L+ +LQ Y+ EE W QK+R D N ++HA T R RN+I L DG
Sbjct: 8 LKEKLQEAYK----DEEDYWQQKSRNMWHISGDLNKKFYHALTKQRHARNRIVGLYDYDG 175
Query: 836 SWCDDRQLLAEEIQNYFQSLFAADPDNTDVNMQHVMMPRLTEAERHKAEAEVIKEEVFQA 895
+W + Q + + +YF+ LF + +T + +EEV A
Sbjct: 176 NWITEEQGVEKVAVDYFEDLFQRTTPTGFDGFLDEITSSITPQMNQRLLRLATEEEVRLA 355
Query: 896 LLRMRSYSALGPDGFHPFFF 915
L M A GPDG F
Sbjct: 356 LFIMHPEKAPGPDGMTTLLF 415
>BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (16%)
Length = 794
Score = 95.9 bits (237), Expect = 1e-19
Identities = 56/182 (30%), Positives = 102/182 (55%), Gaps = 5/182 (2%)
Frame = +3
Query: 772 DMTRLETELQAEYRFLLRQEELLWFQKARENRVCFDDRNTAYFHAHTIIRRKRNKIHRLK 831
+++ L +EL EY EE+ W QK+R N + DRNT +FHA T RR +N+I L
Sbjct: 78 ELSLLRSELNEEYH----NEEIFWMQKSRLNWLRSGDRNTKFFHAVTKNRRAQNRI--LS 239
Query: 832 LNDGSWCDDRQLLAEE-----IQNYFQSLFAADPDNTDVNMQHVMMPRLTEAERHKAEAE 886
L D DD++ EE ++F+ L++++ + + + +TE + + A+
Sbjct: 240 LIDD---DDKEWFVEEDLGRLADSHFKLLYSSEDVGITLEDWNSIPAIVTEEQNAQLMAQ 410
Query: 887 VIKEEVFQALLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLET 946
+ +EEV +A+ + + GPDG + FFF+++W+ +GDDL + ++ TG +++ + +T
Sbjct: 411 ISREEVREAVFDINPHKCPGPDGMNVFFFQQFWDTMGDDLTSMAQEFLRTGKLEEGINKT 590
Query: 947 LI 948
I
Sbjct: 591 NI 596
Score = 56.2 bits (134), Expect = 1e-07
Identities = 34/98 (34%), Positives = 53/98 (53%)
Frame = +1
Query: 915 FKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVYK 974
F WE++ LW+ +++ E G + + L+PK + EFRPISLCNV YK
Sbjct: 505 FGTQWEMISH-LWR--KNSSEQGNLKRESTKQTSGLVPKKLEAKRLVEFRPISLCNVAYK 675
Query: 975 LITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLA 1012
+++KV+ R++ +L II Q +F I DN +A
Sbjct: 676 IVSKVLSKRLKSVLPWIITETQAAFGRRQLISDNILIA 789
>TC81386 similar to GP|10140689|gb|AAG13524.1 putative non-LTR retroelement
reverse transcriptase {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 798
Score = 90.5 bits (223), Expect = 5e-18
Identities = 68/268 (25%), Positives = 116/268 (42%), Gaps = 9/268 (3%)
Frame = -1
Query: 569 KREELWSHLLHIRSNLVLPWMIVGDMNEILHPNEVRGG*FLPTRASR--FAAILEGCGMV 626
KR +LWS + +I++ L W +GD N IL +E +G P R F + +
Sbjct: 798 KRRQLWSAISNIQTQHKLSWCCIGDFNTILGSHEHQGS-HTPARLPMLDFQQWSDVNNLF 622
Query: 627 DLGMSGGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHRIHSDHCPLLINC 686
L G +TW + R KRLDR++ + S L ++ SDH P+L
Sbjct: 621 HLPTRGSAFTWTNGRRGRNNTRKRLDRSIVNQLMIDNCESLSACTLTKLRSDHFPILFEL 442
Query: 687 SGPEIVHHDRPFRFLASWVEHPQYKEVVQSAWTGETE-----TVAGKLNKVRTKSCKFNK 741
I F+F+ W HP +++ W + + KL ++ +NK
Sbjct: 441 QTQNI-QFSSSFKFMKMWSAHPDCINIIKQCWANQVVGCPMFVLNQKLKNLKEVLKVWNK 265
Query: 742 EVYGSIFQRKRWVEGRLRGVQRELNYRVTFDMTRLETELQAEYRF--LLRQEELLWFQKA 799
+G++ + L +Q +++ + + ++ E A+ L EE+ W +K+
Sbjct: 264 NTFGNVHSQVDNAYKELDDIQVKID-SIGYSDVLMDQEKAAQLNLESALNIEEVFWHEKS 88
Query: 800 RENRVCFDDRNTAYFHAHTIIRRKRNKI 827
+ N C DRNTA+FH I+R + I
Sbjct: 87 KVNWHCEGDRNTAFFHRVAKIKRTSSLI 4
>BQ148771
Length = 680
Score = 90.1 bits (222), Expect = 7e-18
Identities = 43/145 (29%), Positives = 82/145 (55%), Gaps = 1/145 (0%)
Frame = -3
Query: 1233 LSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQRG 1292
L++WK N L++ +V LAKSVI A+P Y M +P++ +I + R+F+W R
Sbjct: 573 LANWKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQKLQRKFVWGDTEVSRR 394
Query: 1293 WHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHGESI 1352
+H V W+T+++PK GL ++ + N + + K+ W + + S+ L +V+ KY ES+
Sbjct: 393 YHAVGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSNSLCTEVMRGKYQRSESL 214
Query: 1353 FSVH-RRANASPIWQGILKARDQLQ 1376
+ + S +W+ ++K +++
Sbjct: 213 EEIFLEKPTDSSLWKALVKLWPEIE 139
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 61.6 bits (148), Expect(2) = 8e-16
Identities = 28/68 (41%), Positives = 45/68 (66%)
Frame = +2
Query: 946 TLIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGI 1005
T I LIPKVD+P + +FRPISL +YK++ K++ NR+R ++ +I Q++F+ I
Sbjct: 59 TFIALIPKVDNPQRLNDFRPISLVGSLYKILGKLLANRLRVVIGSVISDAQSAFVKNRQI 238
Query: 1006 MDNAFLAQ 1013
++ FL Q
Sbjct: 239 LEMVFL*Q 262
Score = 42.0 bits (97), Expect(2) = 8e-16
Identities = 21/50 (42%), Positives = 28/50 (56%)
Frame = +1
Query: 1007 DNAFLAQEIIHHMGASMAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGF 1056
D +A E + A KK L FK+D EKAY SV WA+L+ L ++ F
Sbjct: 244 DGILIANEAVDE--AKKLKKDLLLFKVDFEKAYHSVDWAYLDSVLGRYVF 387
>BG585499
Length = 792
Score = 59.3 bits (142), Expect(2) = 6e-09
Identities = 36/125 (28%), Positives = 53/125 (41%), Gaps = 10/125 (8%)
Frame = +3
Query: 1491 WLWI*KLHAPEKL*VFIWLVLHDALPTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCP 1550
W W P + F+WLV H + TN R R V C C + +E +H L DC
Sbjct: 237 WGW----RGPHRTQTFMWLVAHGCILTNYRRSRWGTRVLATCPCCGNADETVLHVLCDCR 404
Query: 1551 HSQELWGKLGAWGWST--FRLPDLKA*V-KTQLQGSNNI-------RFLAGLWGVWKWRN 1600
+ ++W +L W T F D + V K + SN + F+ W +W WRN
Sbjct: 405 PASQVWIRLVPSDWITNFFSFDDCRDWVFKNLSKRSNGVSKFKWQPTFMTTCWHMWTWRN 584
Query: 1601 NVVLD 1605
+ +
Sbjct: 585 KAIFE 599
Score = 20.8 bits (42), Expect(2) = 6e-09
Identities = 8/15 (53%), Positives = 9/15 (59%)
Frame = +2
Query: 1644 WKPPPPEFVKLNTDG 1658
WK P + KLN DG
Sbjct: 725 WKRPLDGWAKLNCDG 769
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 60.1 bits (144), Expect = 8e-09
Identities = 72/285 (25%), Positives = 111/285 (38%), Gaps = 23/285 (8%)
Frame = +3
Query: 1460 WKDNSDGISSVHDGYQWLREKKMAGVVHGDS-------WLWI*KLHAPEKL*VFIWLVLH 1512
W N GI SV GY LR + + + + W I LH + V +W +L+
Sbjct: 6 WMHNPTGIYSVKSGYNTLRTWQTQQINNTSTSSDETLIWKKIWSLHTIPRHKVLLWRILN 185
Query: 1513 DALPTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELW-GKLGAWGWSTFRLPD 1571
D+LP ++ + + P C RC S E H CP S+ +W G + P+
Sbjct: 186 DSLPVRSSLRKRGIQCYPLCPRCHSKTETITHLFMSCPLSKRVWFGSNLCINFDNLPNPN 365
Query: 1572 LKA*VKTQLQGSN---NIRFLAGLWGVWKWRNNVVLDPLPWTLDGAWQKLRHDHDELLQF 1628
*+ + + I A ++ +W RN VL+ Q+ + + Q
Sbjct: 366 FIN*LYEAIL*KDECITI*IAAIIYNLWHARNLSVLEDQTILEMDIIQRASNCISDYKQA 545
Query: 1629 SSG----------DLLDEHHFLINL-WKPPPPEFVKLNTDGSFQDTASSMGGGGLIRDAK 1677
++ D +H N WK P VK+NTD + Q+ G G +IRD
Sbjct: 546 NTQAPPSMARTGYDPRSQHRPAKNTKWKRPNLGLVKVNTDANLQN-HGKWGLGIIIRDEV 722
Query: 1678 GAWLGGFMSHTSIGNPFL-AEVLALRDGLCLAWVQGHRRIIREVD 1721
G + T + L AE AL G+ A G ++ E D
Sbjct: 723 GLVMAASTWETDGNDRALEAEAYALLTGMRFAKDCGFXKVXFEGD 857
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 57.4 bits (137), Expect = 5e-08
Identities = 49/214 (22%), Positives = 89/214 (40%), Gaps = 18/214 (8%)
Frame = -3
Query: 1361 ASPIWQGILKARDQLQEGFQFRMGDG-SSSVWYHDWTGKGKLAESLDFVHISDT------ 1413
AS W+ I A+ +++G + +G+G ++++W + K + H S
Sbjct: 738 ASYAWRSIHSAQHLIKQGAKVIIGNGENTNIWEREMAWKLTCVTNHPNKHSSRAY*APTL 559
Query: 1414 ---KLCLHELVDNNQWNLNRCMTVFPLHIRDWFARVEPRVVANGIDRWSWKDNSDGISSV 1470
+ C + + N N ++FP R + P+ G D +SW+ + G SV
Sbjct: 558 YGYEGCRSDDPMRRERNANLINSIFPEGTRRKILSIHPQGPI-GEDSYSWEYSKSGHYSV 382
Query: 1471 HDGY----QWLREKKMAGVVH----GDSWLWI*KLHAPEKL*VFIWLVLHDALPTNANRY 1522
GY + G V D + + K + K+ F+W + ++LPT AN
Sbjct: 381 KSGYYVQTNIIAAANQRGTVDQPSLDDLYQRVWKYNTSPKVRHFLWRCISNSLPTAANMR 202
Query: 1523 RCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELW 1556
++ C+RC E H L CP+++ +W
Sbjct: 201 SRHISKDGSCSRCGMESETVNHILFQCPYARLIW 100
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 51.6 bits (122), Expect = 3e-06
Identities = 48/190 (25%), Positives = 76/190 (39%), Gaps = 22/190 (11%)
Frame = -2
Query: 1382 RMGDGSSSVWYHD-WTGKGKLAESLD--FVHISDTKLCLHELVDNNQWNLNRCMTVFPLH 1438
++GDG SS ++ D W L ES F K+ +L D N + + L
Sbjct: 703 KVGDGRSSFFWKDAWDSSVPLRESFPRAFFPYRLLKMGCGDLWDMNAEGVRWRLYWRRLE 524
Query: 1439 IRDW--------FARVEPRVVANGIDRWSWKDNSDGISSVHDGYQWLREKKM-------- 1482
+ +W R+E V+ D W WK + +G+ SV+ Y L+ ++
Sbjct: 523 LFEWEKERLLELLGRLEGVVLRYWADIWVWKPDKEGVFSVNSCYFLLQNLRLLEDRLSYE 344
Query: 1483 AGVVHGDSWLWI*KLHAPEKL*VFIWLVLHDALPTNAN---RYRCKLAVSPGCTRCSSPE 1539
V+ + W K AP K+ F W + D +PT N R ++ S C C +
Sbjct: 343 EEVIFRELW----KSKAPAKVLAFSWTLFLDRIPTMVNLGKRRLLRVEDSKRCVFCGCQD 176
Query: 1540 EDTIHCLRDC 1549
E +H C
Sbjct: 175 ETVVHLFLHC 146
Score = 37.0 bits (84), Expect = 0.070
Identities = 18/43 (41%), Positives = 24/43 (54%)
Frame = -3
Query: 1304 PKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKY 1346
P+ GGL ++D+ N SLL K W L + S LW +VL Y
Sbjct: 963 PRCKGGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWKEVLEDIY 835
>TC83479 similar to GP|23476992|emb|CAD48949. hypothetical protein {Plasmodium
falciparum 3D7}, partial (0%)
Length = 1222
Score = 48.9 bits (115), Expect = 2e-05
Identities = 31/78 (39%), Positives = 42/78 (53%)
Frame = +2
Query: 1644 WKPPPPEFVKLNTDGSFQDTASSMGGGGLIRDAKGAWLGGFMSHTSIGNPFLAEVLALRD 1703
WK P + KLNTDGS + G GGL+RD +G + F+S G+ FL E+ A+
Sbjct: 977 WKKPEIGWTKLNTDGSVNKETA--GFGGLLRDYRGEPICAFVSKAPQGDTFLVELWAIWR 1150
Query: 1704 GLCLAWVQGHRRIIREVD 1721
GL L+ G + I E D
Sbjct: 1151 GLVLSLGLGIKSIWVESD 1204
>BG584442
Length = 775
Score = 48.5 bits (114), Expect = 2e-05
Identities = 31/116 (26%), Positives = 57/116 (48%), Gaps = 1/116 (0%)
Frame = +1
Query: 1229 FKKKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLK-N 1287
F KK++ + L+ V + K + +I +Y M +F L S +I + M F W+
Sbjct: 373 FDKKINF*RNKCLSKVM*EVMIKYALQSISSYVMSIFLLLNSQVDEIEKIMNTFSWVHVG 552
Query: 1288 SGQRGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLA 1343
++G H ++ + L K +GG+ D FN +LGK V N + L+++ ++
Sbjct: 553 ENRKGMHWMS*EKLFVHKNYGGMGFTDFTTFNIPMLGKQV*SFLLNRTTLFLEKIS 720
>TC88926 similar to GP|7290507|gb|AAF45960.1| peb gene product {Drosophila
melanogaster}, partial (1%)
Length = 1073
Score = 46.2 bits (108), Expect = 1e-04
Identities = 44/155 (28%), Positives = 69/155 (44%), Gaps = 7/155 (4%)
Frame = +3
Query: 1645 KPPPPEFVKLNTDGSF--QDTASSMGGGGLIRDAKGAWLGGFMSHTSIGNPFL----AEV 1698
+P E V LN DGS + S G GG++ D+ G WL GF NP L E
Sbjct: 321 RPDYLEEVILNVDGSLLREREVPSAGCGGVLSDSSGKWLCGFAQKL---NPNLKVDETEK 491
Query: 1699 LALRDGLCLAWVQGHRRIIREVDCAGLSDILRFEDRCRLHDHAGVLQEIRELMERP-WRC 1757
A+ GL +G R+I+ + D G+ + C + ++ IR+L+ P W
Sbjct: 492 EAILRGLLWVKEKGKRKILVKSDNEGVV----YSVNCGGRSNDPLVCGIRDLLNSPHWEA 659
Query: 1758 SVHWTNRDSNASADFLARRGGFLSTIGVVELMVAP 1792
++ + SNA AD LA + ++ + + P
Sbjct: 660 TLTCIHGRSNAVADRLAHKAHSFTSFDLCQFDYPP 764
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 40.4 bits (93), Expect = 0.006
Identities = 16/39 (41%), Positives = 25/39 (64%)
Frame = +2
Query: 1067 CVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSPYLFVLC 1105
CV ++ +L N + P RGL++GD LSPY+F++C
Sbjct: 104 CVESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIIC 220
>BQ122106 similar to GP|9366656|emb|C probable similar to ring-h2 finger
protein rha1a. {Trypanosoma brucei}, partial (18%)
Length = 693
Score = 40.4 bits (93), Expect = 0.006
Identities = 42/151 (27%), Positives = 68/151 (44%), Gaps = 12/151 (7%)
Frame = -2
Query: 1654 LNTDGSFQDTASSMGGGGLIRDAKGAWLGGFMSH-TSIGNPFLAEVLALRDGLCLAWVQG 1712
LN DGS G GLIR+ G + GF + T+ + LAE+ A+ GL
Sbjct: 539 LNVDGSCLGNP*PTGFNGLIRNIAGLFNSGFPGNITNTSDILLAELHAIFQGL------- 381
Query: 1713 HRRIIREVDCAGLSDILRFED-----------RCRLHDHAGVLQEIRELMERPWRCSVHW 1761
R + G+SD + + D + H +A ++Q+I++L+ + SV
Sbjct: 380 -----RMISDMGISDFVCYFDSLHYVSLINGPSMKFHVYATLIQDIKDLVITS-KASVFH 219
Query: 1762 TNRDSNASADFLARRGGFLSTIGVVELMVAP 1792
T + N ADFL G ++ V+ + V+P
Sbjct: 218 TLCEGNYCADFLEMLGA--ASDSVLTIHVSP 132
>AW686588
Length = 567
Score = 40.0 bits (92), Expect = 0.008
Identities = 39/151 (25%), Positives = 57/151 (36%), Gaps = 12/151 (7%)
Frame = +1
Query: 1470 VHDGYQWLREKKMAGVVHGDSWLWI*KLHAPEKL*VFIWLVLHDALPTNAN--RYRCKLA 1527
+ G W+ K + G W+* P K+ + W ++ D LPT AN R RC
Sbjct: 67 IESGIPWVHP*KEL-LATGAQTTWL*---VPLKVSILAWRLIRDRLPTKANLVRRRCLAV 234
Query: 1528 VSPGCTRCSSPEEDTIHCLRDCPHSQELWGKLGAW-GWSTFRLPDLKA*VKTQLQGSNNI 1586
+ GC E H C +W + AW G S DL + + +
Sbjct: 235 EAAGCVVGCGIAETANHLFLHCATFGAVWQHIRAWIGVSGADPHDLSDHFIQFITCTGHT 414
Query: 1587 R---------FLAGLWGVWKWRNNVVLDPLP 1608
R +L +W VW RNN + + P
Sbjct: 415 RARRSFMQLIWLLCVWMVWNERNNRLFN*YP 507
>AW690000
Length = 652
Score = 40.0 bits (92), Expect = 0.008
Identities = 22/57 (38%), Positives = 33/57 (57%)
Frame = +3
Query: 1643 LWKPPPPEFVKLNTDGSFQDTASSMGGGGLIRDAKGAWLGGFMSHTSIGNPFLAEVL 1699
+W+PP P ++K NTDGS + +S+ GG+ R+ L F +T N F AE+L
Sbjct: 408 IWRPPIPHWIKCNTDGSSRSHSSAC--GGIFRNHDTDLLLCFAENTGECNAFHAELL 572
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.333 0.144 0.485
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 66,928,983
Number of Sequences: 36976
Number of extensions: 1105823
Number of successful extensions: 6342
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 4609
Number of HSP's successfully gapped in prelim test: 219
Number of HSP's that attempted gapping in prelim test: 1465
Number of HSP's gapped (non-prelim): 5183
length of query: 1813
length of database: 9,014,727
effective HSP length: 110
effective length of query: 1703
effective length of database: 4,947,367
effective search space: 8425366001
effective search space used: 8425366001
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0178.7


