

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.5
(770 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
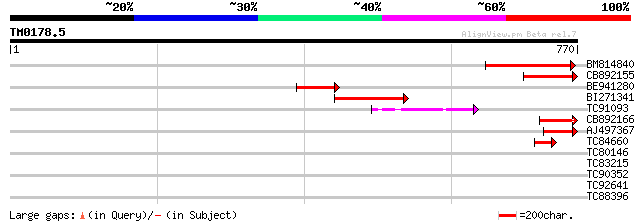
Score E
Sequences producing significant alignments: (bits) Value
BM814840 weakly similar to GP|15128241|db helicase-like protein ... 154 9e-38
CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292... 99 5e-21
BE941280 weakly similar to GP|20197614|g unknown protein {Arabid... 92 1e-18
BI271341 79 9e-15
TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein... 77 2e-14
CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis ... 58 1e-08
AJ497367 similar to GP|14140286|gb putative helicase {Oryza sati... 58 1e-08
TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG282... 44 3e-04
TC80146 40 0.004
TC83215 weakly similar to PIR|F96713|F96713 unknown protein T6L1... 30 3.5
TC90352 similar to SP|Q9LU93|MAD2_ARATH Mitotic spindle checkpoi... 30 4.6
TC92641 30 4.6
TC88396 weakly similar to GP|18616497|emb|CAD22847. unnamed prot... 29 7.8
>BM814840 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (6%)
Length = 733
Score = 154 bits (390), Expect = 9e-38
Identities = 75/122 (61%), Positives = 97/122 (79%)
Frame = +3
Query: 647 TRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAMTI 706
TR+I+ +L K +I A ++ G GE ++IPRM+L PS +++ F+R QFP+ L FAMTI
Sbjct: 33 TRLIIVSLGKNVICARVIGGTHAGEVSYIPRMNLIPSGANVSITFERCQFPLVLSFAMTI 212
Query: 707 NKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQE 766
NKSQGQ+L+ VGLYLPRPVFTHGQLYVA+SRVKSR GLK+LI D+ G S++T NV+YQE
Sbjct: 213 NKSQGQTLTSVGLYLPRPVFTHGQLYVAVSRVKSRSGLKILITDENGSPSSSTVNVVYQE 392
Query: 767 VF 768
VF
Sbjct: 393 VF 398
>CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (4%)
Length = 572
Score = 99.4 bits (246), Expect = 5e-21
Identities = 49/73 (67%), Positives = 59/73 (80%)
Frame = -2
Query: 698 VTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVVSN 757
V + FAMTINKSQGQSL H+G+YLP VF+HGQLYVALSRV SR+GLK+LI +D+G
Sbjct: 370 VEVYFAMTINKSQGQSLKHIGVYLPSSVFSHGQLYVALSRVTSREGLKILISNDDGEDDC 191
Query: 758 TTRNVMYQEVFDN 770
T NV+Y+EVF N
Sbjct: 190 VTSNVVYREVFHN 152
>BE941280 weakly similar to GP|20197614|g unknown protein {Arabidopsis
thaliana}, partial (8%)
Length = 403
Score = 91.7 bits (226), Expect = 1e-18
Identities = 43/59 (72%), Positives = 48/59 (80%)
Frame = -1
Query: 390 KASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVI 448
KA LIIWDE PM++KHCFEALDRSL D++KT DIPFGG VVVLGGDFRQIL V+
Sbjct: 178 KAKLIIWDEAPMMHKHCFEALDRSLRDVLKTVDERNKDIPFGGKVVVLGGDFRQILLVM 2
>BI271341
Length = 468
Score = 78.6 bits (192), Expect = 9e-15
Identities = 42/101 (41%), Positives = 63/101 (61%)
Frame = +1
Query: 441 FRQILPVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIKEFADWL 500
FR ILPVI +GSRS+I+ + INSS + HC+V++L NM LQ +S+ E ++F+ +
Sbjct: 1 FR*ILPVIPRGSRSDIIHATINSSCI*DHCQVVRLKKNMWLQQNGQSSNDPEFEQFSK*I 180
Query: 501 LQVGDGTVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAY 541
L+VGDG + ++ I+IPP LLI + L +V Y
Sbjct: 181 LKVGDGKIYEPNDSYADIDIPPELLISNYDDSLQTIVQSTY 303
>TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein T24M8.10 -
Arabidopsis thaliana, partial (11%)
Length = 701
Score = 77.4 bits (189), Expect = 2e-14
Identities = 48/145 (33%), Positives = 74/145 (50%)
Frame = +3
Query: 492 EIKEFADWLLQVGDGTVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKN 551
E K FA+ L G + ++ ++ PP +P+ +V YP L
Sbjct: 306 EFKTFAEILT----GKMSEPNDSYAEVDTPPG-------DPIDAIVQSTYPNLVSQYNNE 452
Query: 552 SFFQERAILAPTLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLN 611
F Q RAIL T E V++IN+++L +IPG+E S + +S+ + + EFL
Sbjct: 453 QFLQSRAILTSTDEVVDQINDYVLKLIPGEERVIYSAN---RSEVNDVQAFDAIPPEFLQ 623
Query: 612 DFKCSEIPNHAIKLKVGVPIMLIRN 636
K S++PNH + LKVG PIML+R+
Sbjct: 624 SLKTSDLPNHKLTLKVGTPIMLLRD 698
>CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 748
Score = 58.2 bits (139), Expect = 1e-08
Identities = 29/51 (56%), Positives = 38/51 (73%)
Frame = -1
Query: 720 YLPRPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQEVFDN 770
+ R VF+HGQLYVA+SRV SR GLK+L+ID+ G + T NV+Y +VF N
Sbjct: 289 FYDREVFSHGQLYVAISRVSSRNGLKILMIDENGDCIDNTTNVVY-KVFQN 140
>AJ497367 similar to GP|14140286|gb putative helicase {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 543
Score = 58.2 bits (139), Expect = 1e-08
Identities = 27/45 (60%), Positives = 37/45 (82%)
Frame = +1
Query: 726 FTHGQLYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQEVFDN 770
F++G+LYVA+SRV SRKGLK+L+ ++G NTT NV+Y+EVF N
Sbjct: 19 FSNGKLYVAVSRVTSRKGLKILLAHEDGNCMNTTSNVVYKEVFRN 153
>TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (1%)
Length = 1009
Score = 43.5 bits (101), Expect = 3e-04
Identities = 20/30 (66%), Positives = 25/30 (82%)
Frame = +2
Query: 713 SLSHVGLYLPRPVFTHGQLYVALSRVKSRK 742
S++ G+YLP+P+F HG LYVALSRV SRK
Sbjct: 68 SMATKGMYLPQPIF*HG*LYVALSRVTSRK 157
>TC80146
Length = 476
Score = 39.7 bits (91), Expect = 0.004
Identities = 16/35 (45%), Positives = 27/35 (76%)
Frame = -3
Query: 698 VTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLY 732
+ + + TINKS+ QSLS++ +YL RP+F+H ++Y
Sbjct: 471 IQIPYFKTINKSR*QSLSYMKIYLSRPIFSHEEMY 367
>TC83215 weakly similar to PIR|F96713|F96713 unknown protein T6L1.9
[imported] - Arabidopsis thaliana, partial (21%)
Length = 731
Score = 30.0 bits (66), Expect = 3.5
Identities = 21/72 (29%), Positives = 36/72 (49%), Gaps = 8/72 (11%)
Frame = +2
Query: 508 VKTIDEEETLIEIPPNLLIEQ---CKEPLLELVNFAYP-----KLAHNLQKNSFFQERAI 559
+K ++EE+ E+ N L E+ C E L+E+ NFA +A N +++ + +
Sbjct: 383 MKILEEEQKNFEVQINSLSEENEACHEQLIEIENFAESLKESIDIAXNRAESAEAKVTQL 562
Query: 560 LAPTLESVEEIN 571
LE EE+N
Sbjct: 563 TETNLELTEELN 598
>TC90352 similar to SP|Q9LU93|MAD2_ARATH Mitotic spindle checkpoint protein
MAD2. [Mouse-ear cress] {Arabidopsis thaliana}, partial
(97%)
Length = 1023
Score = 29.6 bits (65), Expect = 4.6
Identities = 10/15 (66%), Positives = 12/15 (79%)
Frame = -2
Query: 131 KHKNCKGCCLSHILR 145
KH++CKGCC HI R
Sbjct: 263 KHRDCKGCC*QHIRR 219
>TC92641
Length = 685
Score = 29.6 bits (65), Expect = 4.6
Identities = 18/55 (32%), Positives = 28/55 (50%), Gaps = 1/55 (1%)
Frame = +3
Query: 313 FFYMVLEELDKHLFGIHYLLLC-VQGALSS*MSHLAELHRYCYLEVELLIQDFLF 366
FF+ + + K L+ + LL + LSS ++ LH+Y YL V I +LF
Sbjct: 129 FFHCYYDLISKKLYYCYLLLYSSFEIQLSSLYKKISALHKYFYLSVFFAISSYLF 293
>TC88396 weakly similar to GP|18616497|emb|CAD22847. unnamed protein product
{Triticum aestivum}, partial (32%)
Length = 1399
Score = 28.9 bits (63), Expect = 7.8
Identities = 12/35 (34%), Positives = 23/35 (65%), Gaps = 1/35 (2%)
Frame = +3
Query: 297 FIRMFWMLFCL-IMVDSFFYMVLEELDKHLFGIHY 330
F+ + W++F L I+ D+ F+M+ ++ H F +HY
Sbjct: 288 FLHLDWLIFSLKILADTRFFMLPQDERIHPFWLHY 392
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.347 0.154 0.513
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,719,393
Number of Sequences: 36976
Number of extensions: 375606
Number of successful extensions: 3463
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1474
Number of HSP's successfully gapped in prelim test: 175
Number of HSP's that attempted gapping in prelim test: 1948
Number of HSP's gapped (non-prelim): 1736
length of query: 770
length of database: 9,014,727
effective HSP length: 104
effective length of query: 666
effective length of database: 5,169,223
effective search space: 3442702518
effective search space used: 3442702518
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0178.5


