

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
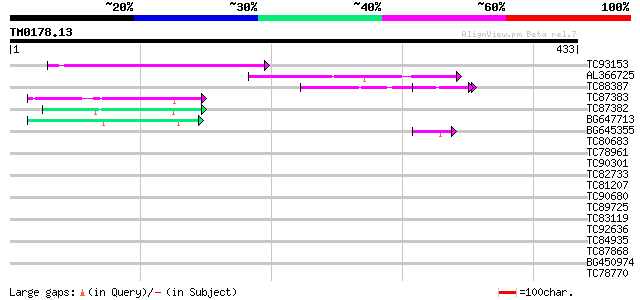
Query= TM0178.13
(433 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 115 3e-26
AL366725 55 5e-08
TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 51 8e-07
TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA PO... 50 1e-06
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 45 5e-05
BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent vir... 44 2e-04
BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.... 44 2e-04
TC80683 homologue to GP|8777424|dbj|BAA97014.1 gb|AAF56406.1~gen... 38 0.009
TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 36 0.025
TC90301 similar to GP|14194175|gb|AAK56282.1 AT5g58470/mqj2_60 {... 36 0.033
TC82733 similar to GP|10177404|dbj|BAB10535. gene_id:K24M7.12~pi... 34 0.13
TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-... 34 0.13
TC90680 similar to GP|12836314|dbj|BAB23601. data source:SPTR s... 34 0.13
TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.... 33 0.16
TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 33 0.21
TC92636 homologue to GP|15042313|gb|AAK82093.1 232R {Chilo iride... 32 0.36
TC84935 similar to PIR|G96631|G96631 probable RNA-binding protei... 32 0.48
TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 32 0.62
BG450974 similar to PIR|T05112|T05 splicing factor 9G8-like SR p... 32 0.62
TC78770 similar to GP|20259299|gb|AAM14385.1 putative cytochrome... 31 0.81
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 115 bits (289), Expect = 3e-26
Identities = 63/170 (37%), Positives = 92/170 (54%), Gaps = 1/170 (0%)
Frame = +2
Query: 30 AATAALVAAQNHAEENARRVQREEREQAAAQ-TRGLNDFKRQDPPKLSGGFDPEEADLWL 88
++ AALVAA E A+ VQ+ + + TR L F R PP G + P+ A WL
Sbjct: 11 SSDAALVAA---LEAVAQAVQQLPKVDTGSDGTRMLETFLRNHPPTFKGRYAPDGA*KWL 181
Query: 89 QELEKIFTFLRTTAEMKVDYATYLLTGEAEYWWRGARAMMEADHQAITWECFRGAFLDKY 148
+E+E+IF ++ KV + T++L EA+ WW ++E D +TW FR FL +Y
Sbjct: 182 KEIERIFRVMQCFETQKVQFGTHMLAEEADDWWISLLPVLEQDDAVVTWAMFRKEFLGRY 361
Query: 149 FPNSARAAKEAQFLRLRQGGMTVAEYASKLESLAKHFRYFRGQIDEGYMC 198
FP R KE +FL L+QG M+V EYA+K LA + ++ + E C
Sbjct: 362 FPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELATFYPHYSAETAEFSKC 511
>AL366725
Length = 485
Score = 55.1 bits (131), Expect = 5e-08
Identities = 37/167 (22%), Positives = 72/167 (42%), Gaps = 4/167 (2%)
Frame = +2
Query: 183 KHFRYFRGQIDEGYMCERFIDGLSYELQRAVQPLGLNRYQVLVEKTKGIEAIDNARGKFQ 242
K + ++ + E C +F +GL +++RA+ L + LV + E A K
Sbjct: 2 KFYPHYAAETAEFSKCIKFENGLRPDIKRAIGYQQLRVFPDLVNTCRIYEEDTKAHDKVV 181
Query: 243 SQNKPNQGNGGSAMTNQGRNDKGRHYQ---KKPYVRPQGQGTASGSFYPTGGNA-IAPRP 298
++ K +G DKG+ ++P + ++ G + + P+
Sbjct: 182 NERK-TKGQ*SRPKPYSAPADKGKQRMVDDRRPKKKDAPAEIVCFNYGEKGHKSNVCPKE 358
Query: 299 LAVSLDGVTCFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPK 345
+ C +C++KGH + C ++C+NCN+ GH+ C+ PK
Sbjct: 359 IK------KCVRCDKKGHIVADCKRNDIVCFNCNEEGHIGSQCKQPK 481
>TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (96%)
Length = 1286
Score = 51.2 bits (121), Expect = 8e-07
Identities = 36/135 (26%), Positives = 59/135 (43%), Gaps = 1/135 (0%)
Frame = +1
Query: 223 VLVEKTKGIEAIDNARGKFQSQNKPNQGNGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTA 282
V V K + ++ +R K +S++ ++ S + + R+++ H ++ PY R +G +
Sbjct: 235 VEVGKILKMSSVSRSRSKSRSRSPMDRSRSRSPVDRRIRSERFSH-REAPYRRDSRRGFS 411
Query: 283 SGSFYPTGGNAIAPRPLAVSLDGVT-CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDC 341
+ N P V C C+ GH AS C L CWNC +PGH+A C
Sbjct: 412 QDNLCK---NCKRPGHYVRECPNVAVCHNCSLPGHIASECSTKSL-CWNCKEPGHMASSC 579
Query: 342 RIPKVEDAANVAGAR 356
+ AG R
Sbjct: 580 PNEGICHTCGKAGHR 624
Score = 41.6 bits (96), Expect = 6e-04
Identities = 19/47 (40%), Positives = 25/47 (52%)
Frame = +1
Query: 308 CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKVEDAANVAG 354
C C ++GH A C C NC K GHLARDC + + N++G
Sbjct: 673 CNNCYKQGHIAVECTNEKA-CNNCRKTGHLARDCPNDPICNLCNISG 810
Score = 39.7 bits (91), Expect = 0.002
Identities = 18/37 (48%), Positives = 20/37 (53%), Gaps = 1/37 (2%)
Frame = +1
Query: 306 VTCFKCNQKGHYASHCPEGPLM-CWNCNKPGHLARDC 341
V C C Q GH + C GPLM C NC GH A +C
Sbjct: 907 VVCRSCQQFGHMSRDCMGGPLMICQNCGGRGHQAYEC 1017
Score = 39.7 bits (91), Expect = 0.002
Identities = 22/49 (44%), Positives = 26/49 (52%)
Frame = +1
Query: 308 CFKCNQKGHYASHCPEGPLMCWNCNKPGHLARDCRIPKVEDAANVAGAR 356
C C + GH A CP P+ C CN GH+AR C PK +NV G R
Sbjct: 730 CNNCRKTGHLARDCPNDPI-CNLCNISGHVARQC--PK----SNVIGDR 855
Score = 35.0 bits (79), Expect = 0.056
Identities = 17/40 (42%), Positives = 21/40 (52%), Gaps = 6/40 (15%)
Frame = +1
Query: 308 CFKCNQKGHYASHC-----PEGPL-MCWNCNKPGHLARDC 341
C C + GH A C P G L +C NC K GH+A +C
Sbjct: 595 CHTCGKAGHRARECTVPQKPPGDLRLCNNCYKQGHIAVEC 714
Score = 30.4 bits (67), Expect = 1.4
Identities = 15/56 (26%), Positives = 22/56 (38%), Gaps = 22/56 (39%)
Frame = +1
Query: 308 CFKCNQKGHYASHCPEG----------------------PLMCWNCNKPGHLARDC 341
C CN GH A CP+ ++C +C + GH++RDC
Sbjct: 787 CNLCNISGHVARQCPKSNVIGDRGGGGSLRGGYRDGGFRDVVCRSCQQFGHMSRDC 954
>TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA POLYMERASE II
{Encephalitozoon cuniculi}, partial (0%)
Length = 1247
Score = 50.4 bits (119), Expect = 1e-06
Identities = 39/139 (28%), Positives = 56/139 (40%), Gaps = 2/139 (1%)
Frame = -2
Query: 14 QQVTNSAAREEAEAQRAATAALVAAQNHAEENARRVQREEREQAAAQTRGLNDFKRQDPP 73
Q++ N A + EAQ+ V++ + + R QR E + +ND K D P
Sbjct: 871 QEIVN-AQQALLEAQQKRFKDHVSSSDSLSSRSSRSQRREFQ--------MNDIK*-DIP 722
Query: 74 KLSGGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLTGEAEYWWRGA--RAMMEAD 131
G P++ WLQ +E++F + E KV L A WW R E
Sbjct: 721 DFEGNLQPDDLLDWLQIMERLFKYKEVLEEQKVKIVAAKLKKLASIWWENVKRRRKREGK 542
Query: 132 HQAITWECFRGAFLDKYFP 150
+ TWE R KY P
Sbjct: 541 SKIKTWEKMRQKLTRKYLP 485
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 45.1 bits (105), Expect = 5e-05
Identities = 36/129 (27%), Positives = 47/129 (35%), Gaps = 4/129 (3%)
Frame = +2
Query: 26 EAQRAATAALVAAQNHAEENARRVQREEREQAAAQTRGL--NDFKRQDPPKLSGGFDPEE 83
E A A L A Q E + + +Q R L ND K D P G ++
Sbjct: 485 EIINAQQALLEAEQRRFEGDVSYSDSSSSRSSHSQRRQLQMNDIK-VDIPDFEGNLQLDD 661
Query: 84 ADLWLQELEKIFTFLRTTAEMKVDYATYLLTGEAEYWWRG--ARAMMEADHQAITWECFR 141
WLQ +E++F + E KV L A WW R E + TW+ R
Sbjct: 662 FLDWLQTIERVFEYKEVPEEQKVKIVAAKLKKHALIWWENLKRRRKREGKSKIKTWDKMR 841
Query: 142 GAFLDKYFP 150
KY P
Sbjct: 842 QKLTRKYLP 868
>BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent virus}, partial
(1%)
Length = 726
Score = 43.5 bits (101), Expect = 2e-04
Identities = 36/146 (24%), Positives = 54/146 (36%), Gaps = 11/146 (7%)
Frame = +2
Query: 14 QQVTNSAAREEAEAQRAATAALVAAQNHAEENARRVQREEREQA-AAQTRGLNDFKRQ-- 70
Q+ + EE Q V AQ E RR ++ + ++ +R +RQ
Sbjct: 143 QRPLHEIEMEEMRRQIQLLQETVNAQQALLEAQRRRNDDDGSGSDSSSSRSSRSHRRQTR 322
Query: 71 ------DPPKLSGGFDPEEADLWLQELEKIFTFLRTTAEMKVDYATYLLTGEAEYWWRGA 124
D P G P+E WLQ +E++F + E KV L A WW+
Sbjct: 323 MSKIKVDIPDF*GKLQPDEFVDWLQTIERVFKYKEVAEEQKVKIVAAKLKKHASIWWKNL 502
Query: 125 RAM--MEADHQAITWECFRGAFLDKY 148
+ E + TW+ R KY
Sbjct: 503 KRKRNCEGKSKIKTWDKMRQKLTRKY 580
>BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.5 [imported]
- Arabidopsis thaliana, partial (5%)
Length = 627
Score = 43.5 bits (101), Expect = 2e-04
Identities = 18/47 (38%), Positives = 24/47 (50%), Gaps = 13/47 (27%)
Frame = -1
Query: 308 CFKCNQKGHYASHCPEGPLM-------------CWNCNKPGHLARDC 341
C+KC Q GH+AS+CP C+ CN+PGH A +C
Sbjct: 501 CYKCQQPGHWASNCPSMSAANRVSGGSGGASGNCYKCNQPGHWANNC 361
Score = 33.1 bits (74), Expect = 0.21
Identities = 10/15 (66%), Positives = 14/15 (92%)
Frame = -1
Query: 308 CFKCNQKGHYASHCP 322
C+KCNQ GH+A++CP
Sbjct: 402 CYKCNQPGHWANNCP 358
>TC80683 homologue to GP|8777424|dbj|BAA97014.1
gb|AAF56406.1~gene_id:K9P8.7~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (16%)
Length = 1360
Score = 37.7 bits (86), Expect = 0.009
Identities = 23/105 (21%), Positives = 47/105 (43%), Gaps = 1/105 (0%)
Frame = +1
Query: 239 GKFQSQNKPNQGNGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGSFYPTGGNAIAPRP 298
G +++ N GG ++G+ K + ++K + + + + S + G I
Sbjct: 826 GDLHAKDAENLKTGGGKKISRGQKGKLKKMKEKYADQDEEERSIRMSLLASSGKPIKKE- 1002
Query: 299 LAVSLDGVTCFKCNQKGHYASHCP-EGPLMCWNCNKPGHLARDCR 342
+ + + + KG + P + P +C+ C K GHL+RDC+
Sbjct: 1003-----ETLPVIETSDKGKKSDSGPIDAPKICYKCKKVGHLSRDCK 1122
>TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (71%)
Length = 974
Score = 36.2 bits (82), Expect = 0.025
Identities = 13/37 (35%), Positives = 20/37 (53%), Gaps = 2/37 (5%)
Frame = +2
Query: 308 CFKCNQKGHYASHCPEGPL--MCWNCNKPGHLARDCR 342
CF C GH+A C G C+ C + GH+ ++C+
Sbjct: 359 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCK 469
Score = 29.6 bits (65), Expect = 2.4
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = +2
Query: 322 PEGPLMCWNCNKPGHLARDCR 342
P G C+NC GH ARDC+
Sbjct: 341 PPGSGRCFNCGIDGHWARDCK 403
>TC90301 similar to GP|14194175|gb|AAK56282.1 AT5g58470/mqj2_60 {Arabidopsis
thaliana}, partial (46%)
Length = 1222
Score = 35.8 bits (81), Expect = 0.033
Identities = 23/60 (38%), Positives = 27/60 (44%), Gaps = 1/60 (1%)
Frame = +1
Query: 235 DNARGKFQSQNKPNQGNGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASGSFYPT-GGNA 293
+ RG GGS NQGR D G + Q P Q G A G++ PT GGNA
Sbjct: 43 NGGRGNNNDGRSGGSNRGGSYGGNQGRED-GGYGQAPPVAAAQSYGGAGGNYPPTYGGNA 219
>TC82733 similar to GP|10177404|dbj|BAB10535.
gene_id:K24M7.12~pir||S42136~similar to unknown protein
{Arabidopsis thaliana}, partial (57%)
Length = 710
Score = 33.9 bits (76), Expect = 0.13
Identities = 15/45 (33%), Positives = 22/45 (48%), Gaps = 5/45 (11%)
Frame = +3
Query: 308 CFKCNQKGHYASHCPEGP-----LMCWNCNKPGHLARDCRIPKVE 347
C +C ++GH A +CP+G +NC GH +C P E
Sbjct: 357 CLRCRRRGHRAQNCPDGGSKEDFKY*YNCGDNGHSLANCPHPLQE 491
>TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-1 {Triticum
aestivum}, partial (39%)
Length = 630
Score = 33.9 bits (76), Expect = 0.13
Identities = 33/138 (23%), Positives = 47/138 (33%), Gaps = 20/138 (14%)
Frame = +3
Query: 227 KTKGIEAIDNARGKFQSQNKPNQGNGGSAMTNQG---RNDKGRHY---QKKPYVRPQGQG 280
KTK ++ + +G+ + N G GG +G RN G Y R +
Sbjct: 225 KTKAVD-VTGPKGEPLQVRQDNHGGGGGGRGFRGGERRNGGGGCYTCGDTGHIARDCDRS 401
Query: 281 TASGSFYPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEGP--------------L 326
+ +GG R A C+ C H+A C G
Sbjct: 402 DRNDRNDRSGGGGGGDRDRA-------CYTCGSFEHFARDCMRGGGNNNNGGGGYGGGGT 560
Query: 327 MCWNCNKPGHLARDCRIP 344
C+ C GH+ARDC P
Sbjct: 561 SCYRCGGVGHIARDCATP 614
>TC90680 similar to GP|12836314|dbj|BAB23601. data source:SPTR source
key:Q9H0W3 evidence:ISS~homolog to HYPOTHETICAL 64.6
KDA PROTEIN~putative, partial (5%)
Length = 1280
Score = 33.9 bits (76), Expect = 0.13
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = +2
Query: 328 CWNCNKPGHLARDCRI 343
CW C +PGHLA DC +
Sbjct: 977 CWKCQRPGHLAEDCLV 1024
>TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.150 -
Arabidopsis thaliana, partial (17%)
Length = 378
Score = 33.5 bits (75), Expect = 0.16
Identities = 16/57 (28%), Positives = 21/57 (36%), Gaps = 20/57 (35%)
Frame = +1
Query: 305 GVTCFKCNQKGHYASHCPEGPL--------------------MCWNCNKPGHLARDC 341
G C+ C + GH A C G C++C + GH ARDC
Sbjct: 7 GGGCYNCGESGHMARECTSGGGGGGGRYGGGGGGGGGGGGGGSCYSCGESGHFARDC 177
Score = 30.4 bits (67), Expect = 1.4
Identities = 9/20 (45%), Positives = 13/20 (65%)
Frame = +1
Query: 305 GVTCFKCNQKGHYASHCPEG 324
G +C+ C + GH+A CP G
Sbjct: 127 GGSCYSCGESGHFARDCPTG 186
>TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (16%)
Length = 421
Score = 33.1 bits (74), Expect = 0.21
Identities = 24/107 (22%), Positives = 45/107 (41%)
Frame = +3
Query: 225 VEKTKGIEAIDNARGKFQSQNKPNQGNGGSAMTNQGRNDKGRHYQKKPYVRPQGQGTASG 284
V+K K ++ ++R + +S+++ ++ S + R+D+ Y+ PY R +G +
Sbjct: 168 VKKEKNLKMSSDSRSRSRSRSR-SRSRSRSPRIRKIRSDR-HSYRDAPYRRDSSRGFSRD 341
Query: 285 SFYPTGGNAIAPRPLAVSLDGVTCFKCNQKGHYASHCPEGPLMCWNC 331
+ C C + GHYA CP +C NC
Sbjct: 342 NL---------------------CKNCKRPGHYARECP-NVAVCHNC 416
Score = 29.6 bits (65), Expect = 2.4
Identities = 9/15 (60%), Positives = 12/15 (80%)
Frame = +3
Query: 327 MCWNCNKPGHLARDC 341
+C NC +PGH AR+C
Sbjct: 345 LCKNCKRPGHYAREC 389
>TC92636 homologue to GP|15042313|gb|AAK82093.1 232R {Chilo iridescent
virus}, partial (1%)
Length = 772
Score = 32.3 bits (72), Expect = 0.36
Identities = 18/76 (23%), Positives = 33/76 (42%)
Frame = +3
Query: 22 REEAEAQRAATAALVAAQNHAEENARRVQREEREQAAAQTRGLNDFKRQDPPKLSGGFDP 81
+E E A A L A + +++ R + + + L + + D P G P
Sbjct: 285 QELQETVNAQQAILEAERRRVDDDGSSDSSSSRSSRSHRRKTLMNDIKVDIPDFEGELQP 464
Query: 82 EEADLWLQELEKIFTF 97
+E WLQ +E++F +
Sbjct: 465 DEFVDWLQAIERVFEY 512
>TC84935 similar to PIR|G96631|G96631 probable RNA-binding protein F8A5.17
[imported] - Arabidopsis thaliana, partial (41%)
Length = 552
Score = 32.0 bits (71), Expect = 0.48
Identities = 13/33 (39%), Positives = 17/33 (51%)
Frame = +2
Query: 309 FKCNQKGHYASHCPEGPLMCWNCNKPGHLARDC 341
F +G Y + G C+ C +PGH ARDC
Sbjct: 416 FSSGGRGSYGAGDRVGQDDCFKCGRPGHWARDC 514
Score = 28.9 bits (63), Expect = 4.0
Identities = 9/15 (60%), Positives = 11/15 (73%)
Frame = +2
Query: 308 CFKCNQKGHYASHCP 322
CFKC + GH+A CP
Sbjct: 473 CFKCGRPGHWARDCP 517
>TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (91%)
Length = 860
Score = 31.6 bits (70), Expect = 0.62
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +3
Query: 326 LMCWNCNKPGHLARDCR 342
L C+ C +PGH AR+CR
Sbjct: 330 LKCYECGEPGHFARECR 380
>BG450974 similar to PIR|T05112|T05 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (54%)
Length = 364
Score = 31.6 bits (70), Expect = 0.62
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +1
Query: 326 LMCWNCNKPGHLARDCR 342
L C+ C +PGH AR+CR
Sbjct: 313 LKCYECGEPGHFARECR 363
>TC78770 similar to GP|20259299|gb|AAM14385.1 putative cytochrome P450
protein {Arabidopsis thaliana}, partial (43%)
Length = 1007
Score = 31.2 bits (69), Expect = 0.81
Identities = 14/34 (41%), Positives = 21/34 (61%), Gaps = 2/34 (5%)
Frame = +3
Query: 395 KLWCYTFC--YRLSLCDVVKTRCDRITIRPDSIY 426
K+WCY +RLSLCD K+R +I ++ S +
Sbjct: 420 KVWCYVQVSHFRLSLCDDFKSRSSKICVKQISTF 521
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,147,295
Number of Sequences: 36976
Number of extensions: 183356
Number of successful extensions: 1106
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 1055
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1093
length of query: 433
length of database: 9,014,727
effective HSP length: 99
effective length of query: 334
effective length of database: 5,354,103
effective search space: 1788270402
effective search space used: 1788270402
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0178.13


