

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
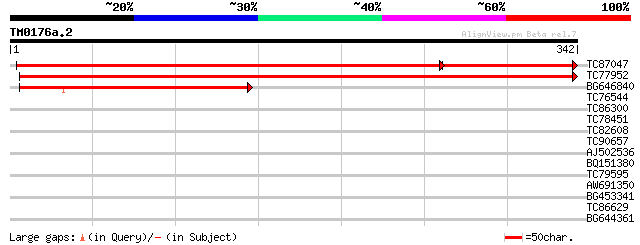
Query= TM0176a.2
(342 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC87047 similar to GP|9989062|gb|AAG10825.1| Unknown protein {Ar... 453 e-160
TC77952 similar to GP|9989062|gb|AAG10825.1| Unknown protein {Ar... 522 e-149
BG646840 similar to GP|15294204|gb AT5g62130/mtg10_150 {Arabidop... 192 2e-49
TC76544 homologue to GP|6581004|gb|AAF18411.1| putative integral... 33 0.12
TC86300 similar to GP|11762234|gb|AAG40395.1 At1g09340 {Arabidop... 32 0.35
TC78451 similar to GP|9279625|dbj|BAB01083.1 gb|AAF23830.1~gene_... 32 0.46
TC82608 30 1.3
TC90657 similar to GP|22775634|dbj|BAC15488. similar to RING zin... 30 1.3
AJ502536 weakly similar to PIR|A86363|A86 hypothetical protein A... 30 1.8
BQ151380 homologue to GP|1504034|dbj| KIAA0227 {Homo sapiens}, p... 29 3.0
TC79595 similar to GP|3850823|emb|CAA77136.1 U2 snRNP auxiliary ... 29 3.0
AW691350 similar to SP|Q9H461|FZD8_ Frizzled 8 precursor (Frizzl... 28 3.9
BG453341 weakly similar to GP|21684781|em beta-glucosidase {Cice... 28 5.1
TC86629 similar to GP|7008011|dbj|BAA90878.1 PsAD2 {Pisum sativu... 28 5.1
BG644361 similar to GP|19698833|gb 60S ribosomal protein L10A {A... 27 8.7
>TC87047 similar to GP|9989062|gb|AAG10825.1| Unknown protein {Arabidopsis
thaliana}, partial (79%)
Length = 1416
Score = 453 bits (1165), Expect(2) = e-160
Identities = 213/259 (82%), Positives = 227/259 (87%), Gaps = 1/259 (0%)
Frame = +1
Query: 5 YVFAFLLVLSW-SVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDR 63
+VF+F LVLS SV V+DAS GD P YR CIRQC+ETGCV CFP C F SDGE R
Sbjct: 175 FVFSFFLVLSCLSVIVVDASKGDAHPLYRSCIRQCEETGCVGPKCFPQCSFSSDGELVGR 354
Query: 64 PWYMQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAAS 123
PWY+QEPLYLQWKKWDC +DCRY+CMLDREKE++LLN PVKYHGKWPF RIYGMQE AS
Sbjct: 355 PWYIQEPLYLQWKKWDCLSDCRYYCMLDREKEKELLNHDPVKYHGKWPFKRIYGMQEPAS 534
Query: 124 VAFSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFH 183
VAFSALNLAMHFHGWVSFFI+LYYKLPLKDGK+AYYEYA LWHIY SLNSW WSAVFH
Sbjct: 535 VAFSALNLAMHFHGWVSFFIVLYYKLPLKDGKKAYYEYASLWHIYAFFSLNSWLWSAVFH 714
Query: 184 SRDVDLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFY 243
SRDVD+TE LDYSSAVILLGYSLIL ILRSFN+RDEATRVMVSAPLIAFVITHVMY+NFY
Sbjct: 715 SRDVDVTEKLDYSSAVILLGYSLILPILRSFNIRDEATRVMVSAPLIAFVITHVMYLNFY 894
Query: 244 KLDYGWNMIVCVVMAVVQL 262
KLDYGWNMIVCVVM V QL
Sbjct: 895 KLDYGWNMIVCVVMDVAQL 951
Score = 130 bits (326), Expect(2) = e-160
Identities = 63/83 (75%), Positives = 72/83 (85%), Gaps = 2/83 (2%)
Frame = +2
Query: 262 LVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGLLDAHALWHATTIPLTY- 320
L +WA+WAG+S HPSRWKLWLVVI+GGLAMLLEIYDFPPYEG LDAHA+WHATTIP+
Sbjct: 950 LPLWALWAGVSRHPSRWKLWLVVISGGLAMLLEIYDFPPYEGFLDAHAIWHATTIPINLR 1129
Query: 321 IWWSFI-RDDAEFRTSIRVKKAK 342
+ SFI RDDAEFRT+ +KKAK
Sbjct: 1130 LGRSFISRDDAEFRTARFLKKAK 1198
>TC77952 similar to GP|9989062|gb|AAG10825.1| Unknown protein {Arabidopsis
thaliana}, partial (81%)
Length = 1451
Score = 522 bits (1345), Expect = e-149
Identities = 234/336 (69%), Positives = 279/336 (82%)
Frame = +2
Query: 7 FAFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWY 66
F L+VL + +DAS GD D Y+ C+ QC+++GCV CF HCKF SDG+ D PWY
Sbjct: 227 FVVLVVLCSFLLSVDASDGDTDLIYKGCVEQCEKSGCVGDRCFQHCKFSSDGKPIDGPWY 406
Query: 67 MQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVAF 126
M EPLYL+WK+WDC+ DCRYHCML RE+ER L PVKYHGKWPF RIYG+QE +VA
Sbjct: 407 MHEPLYLEWKQWDCRTDCRYHCMLAREEERTKLGETPVKYHGKWPFRRIYGIQEPVAVAL 586
Query: 127 SALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRD 186
SALNLAM FHGWVSFFIL+YYKLPL+ K+AYYEY GLWHIYG+LS+N+W WSAVFHSR
Sbjct: 587 SALNLAMQFHGWVSFFILVYYKLPLRPDKKAYYEYTGLWHIYGILSMNAWLWSAVFHSRA 766
Query: 187 VDLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKLD 246
VDLTE L+YSSAV LLG+SLILAILR+FNVRDEATRVMVSAPL+AFV TH+MY+NFY+L+
Sbjct: 767 VDLTEKLNYSSAVALLGFSLILAILRAFNVRDEATRVMVSAPLVAFVTTHIMYLNFYELN 946
Query: 247 YGWNMIVCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGLLD 306
YG NM V ++MAVVQL+IWA+WAG+S HP+RWKLW VV+ G +AM+LE YDFPPY G +D
Sbjct: 947 YGLNMKVSMLMAVVQLLIWAIWAGVSSHPARWKLWTVVVGGVVAMILETYDFPPYMGYVD 1126
Query: 307 AHALWHATTIPLTYIWWSFIRDDAEFRTSIRVKKAK 342
AHA+W+A IPLT++WWS+IRDDAEFRTS +KK K
Sbjct: 1127AHAVWNAANIPLTFLWWSYIRDDAEFRTSALLKKVK 1234
>BG646840 similar to GP|15294204|gb AT5g62130/mtg10_150 {Arabidopsis
thaliana}, partial (31%)
Length = 613
Score = 192 bits (488), Expect = 2e-49
Identities = 88/147 (59%), Positives = 103/147 (69%), Gaps = 7/147 (4%)
Frame = +3
Query: 7 FAFLLVLSWSVEVIDASAGDVDPRY-------RDCIRQCQETGCVAQSCFPHCKFPSDGE 59
F L+VL + +DAS GD D Y R +C+++GCV CF HCKF SDG+
Sbjct: 171 FVVLVVLCSFLLSVDASDGDTDLIYNTDSSDRRSVWSRCEKSGCVGDRCFQHCKFSSDGK 350
Query: 60 HFDRPWYMQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQ 119
D PWYM EPLYL+WK+WDC+ DCRYHCML RE+ER L PVKYHGKWPF RIYG+Q
Sbjct: 351 PIDGPWYMHEPLYLEWKQWDCRTDCRYHCMLAREEERTKLGETPVKYHGKWPFRRIYGIQ 530
Query: 120 EAASVAFSALNLAMHFHGWVSFFILLY 146
E +VA SALNLAM FHGWV FFIL+Y
Sbjct: 531 EPVAVALSALNLAMQFHGWVFFFILIY 611
>TC76544 homologue to GP|6581004|gb|AAF18411.1| putative integral membrane
protein {Phaseolus vulgaris}, partial (48%)
Length = 844
Score = 33.5 bits (75), Expect = 0.12
Identities = 16/42 (38%), Positives = 23/42 (54%), Gaps = 6/42 (14%)
Frame = -2
Query: 56 SDGEHFDRPWYMQEPLYLQWKKW------DCQNDCRYHCMLD 91
SD HF + M+ PLYL+W+ W + + DCR ML+
Sbjct: 258 SDSFHFPNRFTMKFPLYLRWRSWEIK*RLETRADCRIMGMLE 133
>TC86300 similar to GP|11762234|gb|AAG40395.1 At1g09340 {Arabidopsis
thaliana}, partial (97%)
Length = 1328
Score = 32.0 bits (71), Expect = 0.35
Identities = 21/74 (28%), Positives = 36/74 (48%)
Frame = +2
Query: 194 DYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKLDYGWNMIV 253
D+S+ I+LG SL+ L S N+ L F I+H M + Y L WN+I+
Sbjct: 1091 DFSTDDIILGKSLVSV*LLSINL------------LSVFTISHRMNVFVYNLSCWWNVII 1234
Query: 254 CVVMAVVQLVIWAV 267
+ + +I+++
Sbjct: 1235 GLRIQTCTRIIFSL 1276
>TC78451 similar to GP|9279625|dbj|BAB01083.1
gb|AAF23830.1~gene_id:MOJ10.9~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (88%)
Length = 2452
Score = 31.6 bits (70), Expect = 0.46
Identities = 14/37 (37%), Positives = 22/37 (58%)
Frame = +3
Query: 106 YHGKWPFARIYGMQEAASVAFSALNLAMHFHGWVSFF 142
Y GKWPF+ + G Q S+A+ + ++ H W+S F
Sbjct: 1812 YSGKWPFSVLRGSQSFQSIAWKCVAVSSLQH-WLSIF 1919
>TC82608
Length = 798
Score = 30.0 bits (66), Expect = 1.3
Identities = 19/55 (34%), Positives = 26/55 (46%), Gaps = 8/55 (14%)
Frame = -2
Query: 262 LVIWAVW-AGLSGHPSR------WKLWLVVIAGGL-AMLLEIYDFPPYEGLLDAH 308
++ W W A L GHP R W +L+ + L L +I D+ PY LD H
Sbjct: 305 IIDWDPWTAYLDGHPKRVID*DLWVAYLLDLLSWLHGHLKQIIDWDPYTAYLDGH 141
>TC90657 similar to GP|22775634|dbj|BAC15488. similar to RING zinc finger
protein {Oryza sativa (japonica cultivar-group)},
partial (39%)
Length = 757
Score = 30.0 bits (66), Expect = 1.3
Identities = 15/37 (40%), Positives = 18/37 (48%)
Frame = +2
Query: 31 YRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYM 67
Y DC +QC V SC P C PS+ F+ YM
Sbjct: 344 YWDCFKQCS*ASRV--SCMPECYVPSNPSVFEWSHYM 448
>AJ502536 weakly similar to PIR|A86363|A86 hypothetical protein AAB72169.1
[imported] - Arabidopsis thaliana, partial (67%)
Length = 578
Score = 29.6 bits (65), Expect = 1.8
Identities = 18/65 (27%), Positives = 32/65 (48%), Gaps = 6/65 (9%)
Frame = +2
Query: 237 VMYINFYKLDYGWNMIVCVVMAVVQLVIWAVWAGLSGHP--SRWKLWLVV----IAGGLA 290
++ ++ KL + W + + + VI W G+ +P R +LW+VV + GG+
Sbjct: 350 MLQLHKVKLGFSWCLTLLSHKGSIMEVISVFWEGIRSNPP*ERCRLWVVVGCSDLQGGML 529
Query: 291 MLLEI 295
LL I
Sbjct: 530 RLLLI 544
>BQ151380 homologue to GP|1504034|dbj| KIAA0227 {Homo sapiens}, partial (4%)
Length = 1185
Score = 28.9 bits (63), Expect = 3.0
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = -1
Query: 164 LWHIYGLLSLNSWFWSAVFHSR 185
+W +GLL L SW WS V H R
Sbjct: 546 VWICWGLLLLASWGWSLVRHPR 481
>TC79595 similar to GP|3850823|emb|CAA77136.1 U2 snRNP auxiliary factor large
subunit {Nicotiana plumbaginifolia}, partial (77%)
Length = 1525
Score = 28.9 bits (63), Expect = 3.0
Identities = 12/31 (38%), Positives = 19/31 (60%), Gaps = 1/31 (3%)
Frame = +1
Query: 65 WYMQEPLYLQWKK-WDCQNDCRYHCMLDREK 94
W+ Q P + +WKK W ++C C+L RE+
Sbjct: 1255 WFYQSPXWTEWKKIWWKSSNC---CLLSREQ 1338
>AW691350 similar to SP|Q9H461|FZD8_ Frizzled 8 precursor (Frizzled-8) (Fz-8)
(hFz8). [Human] {Homo sapiens}, partial (2%)
Length = 1250
Score = 28.5 bits (62), Expect = 3.9
Identities = 15/46 (32%), Positives = 28/46 (60%), Gaps = 2/46 (4%)
Frame = -2
Query: 232 FVITHVMYINFYKLDYGWNMIVCVVMAVVQLV--IWAVWAGLSGHP 275
F+IT +YI YK +YG + + + +++++ + I A + LS HP
Sbjct: 640 FIITLHLYIYIYKSEYGLSPLSSLSLSILRRIYKIDAKYISLSCHP 503
>BG453341 weakly similar to GP|21684781|em beta-glucosidase {Cicer
arietinum}, partial (29%)
Length = 650
Score = 28.1 bits (61), Expect = 5.1
Identities = 21/75 (28%), Positives = 35/75 (46%), Gaps = 4/75 (5%)
Frame = +2
Query: 95 ERKLLNLVPVKYHGKWPF----ARIYGMQEAASVAFSALNLAMHFHGWVSFFILLYYKLP 150
E LL++ + YH + F A I G+ A+S L+ G+ F + + +
Sbjct: 230 EESLLDIYRIDYHYRHLFYLREAIIAGVNVKGYFAWSLLDNFEWHRGYTVRFGMTF--ID 403
Query: 151 LKDGKEAYYEYAGLW 165
KDG + Y + +GLW
Sbjct: 404 YKDGLKRYQKLSGLW 448
>TC86629 similar to GP|7008011|dbj|BAA90878.1 PsAD2 {Pisum sativum}, partial
(75%)
Length = 717
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/31 (35%), Positives = 23/31 (73%)
Frame = +2
Query: 237 VMYINFYKLDYGWNMIVCVVMAVVQLVIWAV 267
++Y++ Y L NMI+CV++ +++L ++AV
Sbjct: 368 IIYVSSYYLYAICNMILCVLLRIMKLELYAV 460
>BG644361 similar to GP|19698833|gb 60S ribosomal protein L10A {Arabidopsis
thaliana}, partial (79%)
Length = 752
Score = 27.3 bits (59), Expect = 8.7
Identities = 11/31 (35%), Positives = 18/31 (57%)
Frame = +2
Query: 79 DCQNDCRYHCMLDREKERKLLNLVPVKYHGK 109
+C N+C+ C + E+ K LV ++HGK
Sbjct: 461 ECANECQLSCFIAEEELAKC*VLVLEEHHGK 553
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.328 0.142 0.485
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,916,921
Number of Sequences: 36976
Number of extensions: 273442
Number of successful extensions: 1860
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 1839
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1859
length of query: 342
length of database: 9,014,727
effective HSP length: 97
effective length of query: 245
effective length of database: 5,428,055
effective search space: 1329873475
effective search space used: 1329873475
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0176a.2


