

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0173b.3
(286 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
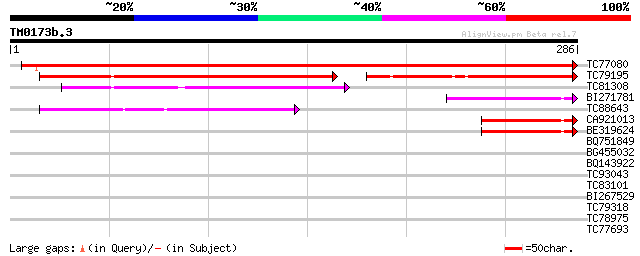
Score E
Sequences producing significant alignments: (bits) Value
TC77080 similar to GP|10177297|dbj|BAB10558. contains similarity... 444 e-125
TC79195 similar to GP|15529178|gb|AAK97683.1 AT5g67480/K9I9_4 {A... 131 2e-57
TC81308 similar to PIR|T01838|T01838 hypothetical protein T15F16... 65 3e-11
BI271781 homologue to GP|9954734|gb Unknown Protein {Arabidopsis... 58 4e-09
TC88643 weakly similar to GP|8778495|gb|AAF79503.1| F20N2.15 {Ar... 55 3e-08
CA921013 similar to GP|12597461|gb p300/CBP acetyltransferase-re... 47 1e-05
BE319624 47 1e-05
BQ751849 similar to GP|23491586|dbj calponin {Branchiostoma belc... 31 0.62
BG455032 weakly similar to GP|10440622|gb| putative NBS-LRR type... 29 1.8
BQ143922 29 2.4
TC93043 weakly similar to PIR|C84918|C84918 probable ATP-depende... 28 3.1
TC83101 similar to GP|15809844|gb|AAL06850.1 AT3g20870/MOE17_16 ... 28 4.0
BI267529 similar to GP|18460924|gb sialoglycoprotease GCP1 {Arab... 28 4.0
TC79318 similar to PIR|H84431|H84431 probable glutamate decarbox... 28 4.0
TC78975 similar to GP|21592656|gb|AAM64605.1 alcohol dehydrogena... 27 9.0
TC77693 weakly similar to GP|7108597|gb|AAF36490.1| K+ transport... 27 9.0
>TC77080 similar to GP|10177297|dbj|BAB10558. contains similarity to unknown
protein~gene_id:MDC12.13~pir||T06706 {Arabidopsis
thaliana}, partial (67%)
Length = 1539
Score = 444 bits (1141), Expect = e-125
Identities = 211/283 (74%), Positives = 245/283 (86%), Gaps = 3/283 (1%)
Frame = +2
Query: 7 QTTRVE--PEPDIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVP 64
QTT +E PE D+FIHTSDGTRIPAH+ ILAS+SPV E++ID+PRK RSSERIIQIHGVP
Sbjct: 149 QTTPMEVLPEYDVFIHTSDGTRIPAHTTILASVSPVFENIIDQPRKQRSSERIIQIHGVP 328
Query: 65 GDAVTAFLTFLYSRRCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDM 124
AVTAF+ FLYS CTEDEMD+YG+HLLALSHVY VP LKQRC KGL QR+ +ENVVD+
Sbjct: 329 STAVTAFVGFLYSSSCTEDEMDKYGIHLLALSHVYNVPQLKQRCIKGLVQRLTSENVVDV 508
Query: 125 LQLARLCDAPDLHLRCMNHLTRNFKTVEETEGWRFLTKHDPWLELEVLRFMDESETRREK 184
LQLARLCDAPDL +RCM LT +F V++TEGW+FLTKHDP LEL++LRFMDESETRRE
Sbjct: 509 LQLARLCDAPDLRVRCMKLLTNHFNAVQKTEGWKFLTKHDPVLELDILRFMDESETRREN 688
Query: 185 SRRSREVQGLYAQLSEAMECLDHICTEGCTLVGPHHVEVDRKSGPCNKFATCQGVQQLIR 244
SR+ RE QGLYA+LSEAMECL+HICTEGCT VGP+HVEV+++ PC+K++TCQG+Q LIR
Sbjct: 689 SRKHREEQGLYAELSEAMECLEHICTEGCTDVGPYHVEVNKEKKPCSKYSTCQGLQLLIR 868
Query: 245 HYATCTKRMGGGCLRCKRMWQLFRLHSYGCEHA-DSCNVPLCR 286
H+ATC KR+ GGC RCKRMWQLFRLHSY C+ DSCNVPLCR
Sbjct: 869 HFATCKKRVKGGCWRCKRMWQLFRLHSYVCQQTHDSCNVPLCR 997
>TC79195 similar to GP|15529178|gb|AAK97683.1 AT5g67480/K9I9_4 {Arabidopsis
thaliana}, partial (78%)
Length = 1519
Score = 131 bits (330), Expect(2) = 2e-57
Identities = 70/151 (46%), Positives = 98/151 (64%), Gaps = 1/151 (0%)
Frame = +2
Query: 16 DIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFL 75
D+ I+T +G I AHSNI+A SPVL+ M+ + + + R I I GVP DAV F+ +L
Sbjct: 329 DVCINTDNGGIIYAHSNIIAVASPVLKGMLKQANRS-NRWRSISIFGVPHDAVRVFIRYL 505
Query: 76 YSRRCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNT-ENVVDMLQLARLCDAP 134
YS ++EM + +HLL LSHVY VPHLK+ C + L + T +N+VD+ QLA LCDAP
Sbjct: 506 YSSCYEKEEMKEFVLHLLVLSHVYAVPHLKRECEQKLELGLLTMDNLVDVFQLALLCDAP 685
Query: 135 DLHLRCMNHLTRNFKTVEETEGWRFLTKHDP 165
L L C + +NF+TV E+EGW+ + + P
Sbjct: 686 RLSLICHRKILKNFRTVSESEGWKAMKQSHP 778
Score = 108 bits (270), Expect(2) = 2e-57
Identities = 51/106 (48%), Positives = 70/106 (65%)
Frame = +1
Query: 181 RREKSRRSREVQGLYAQLSEAMECLDHICTEGCTLVGPHHVEVDRKSGPCNKFATCQGVQ 240
RRE+ R+ E + +Y QL +AME L HIC +GC +GPH + + + PC ++ +C+G++
Sbjct: 829 RRERIRKMNERE-VYLQLYDAMEALVHICRDGCRTIGPHDKDF-KANQPC-RYTSCKGLE 999
Query: 241 QLIRHYATCTKRMGGGCLRCKRMWQLFRLHSYGCEHADSCNVPLCR 286
L+RH+A C R+ GGC CKRMWQL LHS C D C VPLCR
Sbjct: 1000LLVRHFAGCKLRVPGGCGHCKRMWQLLELHSRLCADPDFCRVPLCR 1137
>TC81308 similar to PIR|T01838|T01838 hypothetical protein T15F16.14 -
Arabidopsis thaliana, partial (57%)
Length = 1141
Score = 65.1 bits (157), Expect = 3e-11
Identities = 44/146 (30%), Positives = 71/146 (48%), Gaps = 1/146 (0%)
Frame = +2
Query: 27 IPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRR-CTEDEM 85
IPAH +L S SPV +M++ + S I+I V DA+ AF+ +LY+ C +++M
Sbjct: 485 IPAHKAVLVSRSPVFRAMLENDMEE-SRSGTIKIADVSYDALRAFVNYLYTAEACLDNQM 661
Query: 86 DRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNHLT 145
+LL L Y V HLK C K L ++N E + A +A L + +
Sbjct: 662 ---ACNLLVLGEKYQVKHLKAYCEKYLISKLNWEKAIVNFAFAHQHNANQLQDAALAVIM 832
Query: 146 RNFKTVEETEGWRFLTKHDPWLELEV 171
N + + E + L + +P L +E+
Sbjct: 833 ENMDNLTKNEDYTELVETNPRLVVEI 910
>BI271781 homologue to GP|9954734|gb Unknown Protein {Arabidopsis thaliana},
partial (7%)
Length = 655
Score = 58.2 bits (139), Expect = 4e-09
Identities = 25/66 (37%), Positives = 38/66 (56%)
Frame = +2
Query: 221 VEVDRKSGPCNKFATCQGVQQLIRHYATCTKRMGGGCLRCKRMWQLFRLHSYGCEHADSC 280
V + P ++ C+ V+ L RH C R GGC+ CK+MW L +LH+ C+ ++ C
Sbjct: 59 VHASQCRSPHCQYPNCRKVKGLFRHGMHCKTRASGGCVLCKKMWYLLQLHARACKESE-C 235
Query: 281 NVPLCR 286
+VP CR
Sbjct: 236 HVPRCR 253
>TC88643 weakly similar to GP|8778495|gb|AAF79503.1| F20N2.15 {Arabidopsis
thaliana}, partial (19%)
Length = 1072
Score = 55.1 bits (131), Expect = 3e-08
Identities = 37/131 (28%), Positives = 61/131 (46%)
Frame = +2
Query: 16 DIFIHTSDGTRIPAHSNILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFL 75
DI I+ + I AH +L+S S V SM K I I + + F+ FL
Sbjct: 680 DITINVNSEGNIRAHRAVLSSQSSVFRSMFSHNVKETDLSTI-NITDMSIQSFQTFINFL 856
Query: 76 YSRRCTEDEMDRYGMHLLALSHVYMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPD 135
Y ++E + + LL + Y + L++ C K L + ++ +NV++ LQ+A L
Sbjct: 857 YGN-VNDEEFLMHRLDLLHAADKYDIYELREACQKSLQEDIDPKNVLERLQIASLYQ*TT 1033
Query: 136 LHLRCMNHLTR 146
L CM +L +
Sbjct: 1034LKFSCMQYLVK 1066
>CA921013 similar to GP|12597461|gb p300/CBP acetyltransferase-related
protein 2 {Arabidopsis thaliana}, partial (1%)
Length = 780
Score = 46.6 bits (109), Expect = 1e-05
Identities = 17/48 (35%), Positives = 29/48 (60%)
Frame = -2
Query: 239 VQQLIRHYATCTKRMGGGCLRCKRMWQLFRLHSYGCEHADSCNVPLCR 286
+++L H + C R+ GGC CK++W + HS C+ ++ C +P CR
Sbjct: 506 IKKLFYHASKCKIRLNGGCQHCKKIWFILTSHSRNCKDSE-CRIPRCR 366
>BE319624
Length = 531
Score = 46.6 bits (109), Expect = 1e-05
Identities = 17/48 (35%), Positives = 29/48 (60%)
Frame = +2
Query: 239 VQQLIRHYATCTKRMGGGCLRCKRMWQLFRLHSYGCEHADSCNVPLCR 286
+++L H + C R+ GGC CK++W + HS C+ ++ C +P CR
Sbjct: 158 IKKLFYHASKCKIRLNGGCQHCKKIWFILTSHSRNCKDSE-CRIPRCR 298
>BQ751849 similar to GP|23491586|dbj calponin {Branchiostoma belcheri},
partial (16%)
Length = 684
Score = 30.8 bits (68), Expect = 0.62
Identities = 10/26 (38%), Positives = 18/26 (68%)
Frame = -1
Query: 248 TCTKRMGGGCLRCKRMWQLFRLHSYG 273
+C+ R GGCL+ +R W++F + + G
Sbjct: 309 SCSWRFRGGCLQARRKWEMFSIWTKG 232
>BG455032 weakly similar to GP|10440622|gb| putative NBS-LRR type resistance
protein 3' partial {Oryza sativa}, partial (3%)
Length = 666
Score = 29.3 bits (64), Expect = 1.8
Identities = 27/118 (22%), Positives = 52/118 (43%), Gaps = 2/118 (1%)
Frame = +1
Query: 32 NILASMSPVLESMIDRPRKHRSSERIIQIHGVPGDAVTAFLTFLYSRRCTEDEMDRYGMH 91
++ + P LES+ ++ + S R + I G G RC + + H
Sbjct: 304 SLFVTFLPQLESLPEQNWEGLQSLRALLIWGCRG------------LRCLPEGI----RH 435
Query: 92 LLALSHVYMV--PHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNHLTRN 147
L +L + ++ P LK+RC +G + + + ++L + +A L C+NH +N
Sbjct: 436 LTSLELLSIIDCPTLKERCKEGTGEDWDKIAHIPRIELIDV*EAETFTLICLNHHRKN 609
>BQ143922
Length = 999
Score = 28.9 bits (63), Expect = 2.4
Identities = 24/82 (29%), Positives = 41/82 (49%), Gaps = 5/82 (6%)
Frame = +2
Query: 30 HSNILASMSPVLESMIDRPRKHRSSERIIQIHGVP-GDAVTAFLT---FLYSRRCTEDEM 85
HS+ A +SP ES + R H++S +P + T ++T LY+ + DE
Sbjct: 755 HSHHTAPISPPAESQYNDERNHKASAS--HPKSLP*SNPRTPYITPY*PLYTLLGSRDEC 928
Query: 86 DRYGMHLLALSHVYM-VPHLKQ 106
DR G+ + L+ Y+ + +L Q
Sbjct: 929 DREGVRMRGLNRTYISIDYLSQ 994
>TC93043 weakly similar to PIR|C84918|C84918 probable ATP-dependent RNA
helicase A [imported] - Arabidopsis thaliana, partial
(14%)
Length = 735
Score = 28.5 bits (62), Expect = 3.1
Identities = 21/57 (36%), Positives = 28/57 (48%), Gaps = 4/57 (7%)
Frame = -3
Query: 209 CTEGCTL-VGPHHVEVDR---KSGPCNKFATCQGVQQLIRHYATCTKRMGGGCLRCK 261
C G TL VGPHHV++++ P K TC ++ A C K M CL C+
Sbjct: 175 CFYGPTLLVGPHHVKINKILHFGNPARKRRTC----TVVS*PAMCVK-MNMNCLCCR 20
>TC83101 similar to GP|15809844|gb|AAL06850.1 AT3g20870/MOE17_16
{Arabidopsis thaliana}, partial (53%)
Length = 516
Score = 28.1 bits (61), Expect = 4.0
Identities = 10/35 (28%), Positives = 19/35 (53%)
Frame = -2
Query: 248 TCTKRMGGGCLRCKRMWQLFRLHSYGCEHADSCNV 282
T TK + G + C W++ ++YGC ++ N+
Sbjct: 494 TNTKTLHGSKIHCHSFWKIM*TYTYGCNNSTE*NL 390
>BI267529 similar to GP|18460924|gb sialoglycoprotease GCP1 {Arabidopsis
thaliana}, partial (29%)
Length = 624
Score = 28.1 bits (61), Expect = 4.0
Identities = 12/37 (32%), Positives = 18/37 (48%)
Frame = -2
Query: 244 RHYATCTKRMGGGCLRCKRMWQLFRLHSYGCEHADSC 280
RH++ ++ RC + L R+ YGCE SC
Sbjct: 257 RHFSRLRQQCRRS*FRCPKRQYLHRVLEYGCERRRSC 147
>TC79318 similar to PIR|H84431|H84431 probable glutamate decarboxylase
[imported] - Arabidopsis thaliana, partial (77%)
Length = 1934
Score = 28.1 bits (61), Expect = 4.0
Identities = 15/50 (30%), Positives = 27/50 (54%)
Frame = +1
Query: 99 YMVPHLKQRCTKGLSQRVNTENVVDMLQLARLCDAPDLHLRCMNHLTRNF 148
++ +++ C K + + +N +N VDM + DLH RC+N + R F
Sbjct: 286 FVTTSMEEECNKLIMESIN-KNYVDMDEYPA---TTDLHNRCVNMIARLF 423
>TC78975 similar to GP|21592656|gb|AAM64605.1 alcohol dehydrogenase
putative {Arabidopsis thaliana}, partial (77%)
Length = 1691
Score = 26.9 bits (58), Expect = 9.0
Identities = 13/26 (50%), Positives = 18/26 (69%)
Frame = -1
Query: 91 HLLALSHVYMVPHLKQRCTKGLSQRV 116
HLL L H+ +VP L+QR L+QR+
Sbjct: 785 HLLFLMHLLLVPLLQQRGQLHLNQRL 708
>TC77693 weakly similar to GP|7108597|gb|AAF36490.1| K+ transporter HAK5
{Arabidopsis thaliana}, partial (67%)
Length = 2635
Score = 26.9 bits (58), Expect = 9.0
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = +3
Query: 47 RPRKHRSSERIIQIHGVPGDAV 68
+PR + SS RII HGV D++
Sbjct: 2124 KPRSNNSSGRIIPSHGVSSDSI 2189
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.136 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,638,786
Number of Sequences: 36976
Number of extensions: 168297
Number of successful extensions: 987
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 978
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 982
length of query: 286
length of database: 9,014,727
effective HSP length: 95
effective length of query: 191
effective length of database: 5,502,007
effective search space: 1050883337
effective search space used: 1050883337
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0173b.3


