

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0166.1
(916 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
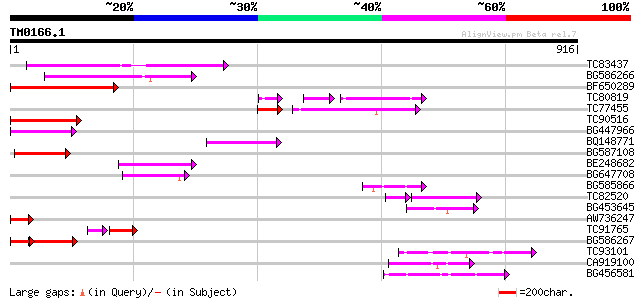
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 169 6e-42
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 139 6e-33
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 121 1e-27
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 74 1e-23
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 83 3e-22
TC90516 87 2e-17
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 83 4e-16
BQ148771 78 1e-14
BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - ... 76 5e-14
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 72 7e-13
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 63 6e-10
BG585866 61 2e-09
TC82520 43 2e-08
BG453645 52 8e-07
AW736247 weakly similar to PIR|T14619|T146 reverse transcriptase... 52 1e-06
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 42 1e-06
BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At... 51 2e-06
TC93101 50 3e-06
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 50 5e-06
BG456581 48 1e-05
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 169 bits (427), Expect = 6e-42
Identities = 114/327 (34%), Positives = 170/327 (51%), Gaps = 1/327 (0%)
Frame = +2
Query: 28 EFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYRLISLIGSIYKLLAKVLAARLQGVMPYL 87
EF+ KL KG+N+ FIALIPK P+ + D+R ISL+GS+YK+L K+LA RL+ V+ +
Sbjct: 17 EFHRNRKLFKGINSTFIALIPKVDNPQRLNDFRPISLVGSLYKILGKLLANRLRVVIGSV 196
Query: 88 LSENQFSFTKGRQIANCLLIGNEVVDTMLSSGRGGFLLKLDFAKAYDNIEWDFLMNMM-E 146
+S+ Q +F K RQI + + + L + R F K + + +
Sbjct: 197 ISDAQSAFVKNRQILEMVFL*QMRLWMRLRN*RKIFCCLRWILKRLITLSIGLIWILF*V 376
Query: 147 DLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFEIEKGLRQGDPLSPLLFNLCVQGLSVM 206
+ F WR WI CVSTAT +VLVNGSPT + K L Q + F
Sbjct: 377 GMSFLVLWRKWIKECVSTATTSVLVNGSPTNV--LMKSLVQTQLFTRYSFG--------- 523
Query: 207 WNRLAIRERPCGVVLGPGVCLNHLQFADDTLVFCDPECAQMEELFDVMTAFLWGSGLKLN 266
V+ P V ++HLQFA+DTL+ A + L + F SGLK+N
Sbjct: 524 -------------VVNP-VVVSHLQFANDTLLLETKNWANIRALRAALVIF*AMSGLKVN 661
Query: 267 YAKSELVGCNVDSAIVNQLAVELGVVVGTLPLPYLGSVLGGDPRRRGFWAPVIEKVRVAL 326
+ KS LV N+ + +++ A L VG +P YLG + G+ RR FW P++ +++ L
Sbjct: 662 FHKSGLVCVNIAPSWLSEAASVLSWKVGKVPFLYLGMPIEGNSRRLSFWEPIVNRIKARL 841
Query: 327 AS*GSNSVSLSGRLVVLRSVASAIPIY 353
S +S GRLV+L+SV +++ +Y
Sbjct: 842 TGWNSRFLSFGGRLVLLKSVLTSLSVY 922
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 139 bits (349), Expect = 6e-33
Identities = 89/256 (34%), Positives = 138/256 (53%), Gaps = 9/256 (3%)
Frame = -3
Query: 56 VTDYRLISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTM 115
V++YR I+ + YK++AK+L+ R+Q ++ ++S +Q +F GR I++ +LI ++++ +
Sbjct: 778 VSEYRTIAPCNTQYKIIAKILSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKILHYL 599
Query: 116 LSSGRG---GFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVN 172
SG +K D KAYD I W+FL ++ LGF W WI CVST + + L+N
Sbjct: 598 RQSGAKKHVSMAVKTDMTKAYDRIAWNFLREVLTRLGFHGIWISWIMECVSTVSYSFLIN 419
Query: 173 GSPTEFFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGPGVC-----L 227
G P +GLRQGDPLSP LF LC + LS + + A+R+ G + G V +
Sbjct: 418 GGPQGRVLPSRGLRQGDPLSPYLFILCTEVLSGLCQQ-ALRK---GTLPGVKVARNCPPI 251
Query: 228 NHLQFADDTLVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELV-GCNVDSAIVNQLA 286
NHL FADDT+ F + L +M + SG +N KS + AI++++
Sbjct: 250 NHLLFADDTMFFGKSNASSCAILLSIMDKYRAASGRCIN*TKSAITFSSKTSQAIIDRVK 71
Query: 287 VELGVVVGTLPLPYLG 302
EL + YLG
Sbjct: 70 GELKIAKEGGTGKYLG 23
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 121 bits (303), Expect = 1e-27
Identities = 59/177 (33%), Positives = 109/177 (61%), Gaps = 1/177 (0%)
Frame = +3
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PG DG+N FF+ W++I + ++ +F++ G + K +N ++ L+PK V ++R
Sbjct: 78 PGIDGYNVHFFKCSWNIIGDSVIDAILDFFKTGFMPKIINCTYVTLLPKEVNVTSVKNFR 257
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSG- 119
I+ IYK+++K+L +R+QGV+ ++SENQ +F KGR I + +++ +E+V + G
Sbjct: 258 PIACCSVIYKIISKILTSRMQGVLNSVVSENQSAFVKGRVIFDNIILSHELVKSYSRKGI 437
Query: 120 RGGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPT 176
++K+D KAYD+ EW F+ ++M +LGF ++ W+ + ++TA+ NG T
Sbjct: 438 SPRCMVKIDLXKAYDSXEWPFIKHLMLELGFPYKFVNWVMAXLTTASYTFNXNGDLT 608
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 74.3 bits (181), Expect(3) = 1e-23
Identities = 52/141 (36%), Positives = 64/141 (44%), Gaps = 2/141 (1%)
Frame = +2
Query: 535 WEWNLGFRRRFFGWEEQVYDELIARCNG-VVPSREKDSLIWLGDTNGRYTVKTFCMEVEL 593
WEW RR F WEE+ E N V+ D WL D Y+VK F +
Sbjct: 422 WEWT----RRLFVWEEECVRECCILLNNFVLQDNVNDKWRWLLDPVNGYSVKVFYRYITS 589
Query: 594 QLYGPPSWVVPSCVKKLAPQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEAVCVLCGQAE 653
+ +V K P KV LF+W+ NR+ NL +RGVL+ A CV CG +
Sbjct: 590 TGHISDRSLVDDVWHKHIPSKVSLFVWRLLRNRLPTKDNLVHRGVLLATNAACV-CGCVD 766
Query: 654 -ESVTHLILVCKFAWVLWSKV 673
ES THL L C LWS V
Sbjct: 767 SESTTHLFLHCNVFCSLWSLV 829
Score = 37.4 bits (85), Expect(3) = 1e-23
Identities = 19/39 (48%), Positives = 23/39 (58%)
Frame = +1
Query: 402 GLGVEFLKFRNKAMLLKWAWRFGQEPEALWRRVLCAKYG 440
GLGV N ++L KW WR + E LW RVL A+YG
Sbjct: 31 GLGVGAF---NLSLLGKWCWRLLVDKEGLWHRVLKARYG 138
Score = 37.0 bits (84), Expect(3) = 1e-23
Identities = 17/50 (34%), Positives = 24/50 (48%)
Frame = +3
Query: 475 LGKTFRDQCFCVVGDGLRISFWRDPWVASKTLAVLYPRFYAMASNQDIMV 524
+G F + VVGDG FW D W L + YPR +A +++ V
Sbjct: 228 VGNWFEENIRMVVGDGRDAFFWYDTWAGDVPLRLKYPRLLDLAMDKECKV 377
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 83.2 bits (204), Expect(2) = 3e-22
Identities = 65/215 (30%), Positives = 100/215 (46%), Gaps = 8/215 (3%)
Frame = -2
Query: 457 HPSRYMFDIVNVVLEDSVLGKTFRDQCFCVVGDGLRISFWRDPWVASKTLAVLYPRFYAM 516
+ SR+ D+++ LE+ + F + VGDG FW+D W +S L +PR +
Sbjct: 784 YASRWWKDLMS--LEEVGRVRWFPRELIRKVGDGRSSFFWKDAWDSSVPLRESFPRAFFP 611
Query: 517 ASNQDIMVSEAGVFSNLGWEWNLGFRR-RFFGWEEQVYDELIARCNGVVPSREKDSLIWL 575
+ + + G W L +RR F WE++ EL+ R GVV D +W
Sbjct: 610 YRLLKMGCGDLWDMNAEGVRWRLYWRRLELFEWEKERLLELLGRLEGVVLRYWADIWVWK 431
Query: 576 GDTNGRYTVKT--FCME----VELQLYGPPSWVVPSCVKKLAPQKVVLFLWQAAENRVAV 629
D G ++V + F ++ +E +L + K AP KV+ F W +R+
Sbjct: 430 PDKEGVFSVNSCYFLLQNLRLLEDRLSYEEEVIFRELWKSKAPAKVLAFSWTLFLDRIPT 251
Query: 630 MVNLHNRGVL-VDAEAVCVLCGQAEESVTHLILVC 663
MVNL R +L V+ CV CG +E+V HL L C
Sbjct: 250 MVNLGKRRLLRVEDSKRCVFCGCQDETVVHLFLHC 146
Score = 41.2 bits (95), Expect(2) = 3e-22
Identities = 18/40 (45%), Positives = 25/40 (62%)
Frame = -3
Query: 401 GGLGVEFLKFRNKAMLLKWAWRFGQEPEALWRRVLCAKYG 440
GGLGV ++ N ++L KW WR Q+ +LW+ VL YG
Sbjct: 951 GGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWKEVLEDIYG 832
>TC90516
Length = 983
Score = 87.4 bits (215), Expect = 2e-17
Identities = 46/115 (40%), Positives = 72/115 (62%)
Frame = -2
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
P PDGFN FF +++K +I MFE+FY L+K + F+ LI K + P ++ D+
Sbjct: 466 PRPDGFNLNFFIRLRNMLKADIEIMFEQFYTPANLLKIFS*YFLTLISKVEYPILLGDFS 287
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTM 115
L+S +GS+ KL+AKVLA RL +M ++ NQ +F +GRQ + ++ NE++ M
Sbjct: 286 LMSFLGSL*KLMAKVLALRLAHIMEKIIFVNQSTFVRGRQHVDGVVAINEIIVLM 122
Score = 31.2 bits (69), Expect = 1.9
Identities = 14/22 (63%), Positives = 18/22 (81%)
Frame = -2
Query: 168 AVLVNGSPTEFFEIEKGLRQGD 189
A+LVN SPT +I+KGL+QGD
Sbjct: 76 AILVNCSPT*EIDIQKGLKQGD 11
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 83.2 bits (204), Expect = 4e-16
Identities = 40/107 (37%), Positives = 62/107 (57%)
Frame = +2
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDG FF+++WH++ +E+ QM + + LN FI LIPK + P DYR
Sbjct: 311 PGPDGLPALFFQKYWHIVGKEVQQMVLQVLNNSMETEELNKTFIVLIPKGKNPNTPKDYR 490
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLI 107
ISL + K++ KV+A R++ +P ++ Q +F +GR I + LI
Sbjct: 491 PISLCNVVMKIITKVIANRVKQTLPDVIDVEQSAFVQGRLITDNALI 631
>BQ148771
Length = 680
Score = 78.2 bits (191), Expect = 1e-14
Identities = 44/122 (36%), Positives = 65/122 (53%)
Frame = -3
Query: 318 VIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHMAVYKAPLGVVADIEKLMRCFLW 377
V +V V LA+ +N +SL+ R+ + +SV A+P+Y M P + +I+KL R F+W
Sbjct: 597 VYYQVHVMLANWKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQKLQRKFVW 418
Query: 378 GRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAMLLKWAWRFGQEPEALWRRVLCA 437
G + RR V WE + K K I GLG+ L NKA ++K W +L V+
Sbjct: 417 GDTEVSRRYHAVGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSNSLCTEVMRG 238
Query: 438 KY 439
KY
Sbjct: 237 KY 232
>BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - Arabidopsis
thaliana, partial (17%)
Length = 677
Score = 76.3 bits (186), Expect = 5e-14
Identities = 38/90 (42%), Positives = 56/90 (62%)
Frame = +3
Query: 9 TFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYRLISLIGSI 68
+FF+ WH+IK ++++M F GKL LN I LIPK++ P +T+ R ISL
Sbjct: 408 SFFQHSWHIIKMDLLKMVNSFLASGKLDTRLNTTNICLIPKKKRPTRMTELRPISLCNVG 587
Query: 69 YKLLAKVLAARLQGVMPYLLSENQFSFTKG 98
YK+++KVL RL+ +P L+SE Q +F G
Sbjct: 588 YKIISKVLCQRLKVCLPSLISETQSAFVHG 677
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 72.4 bits (176), Expect = 7e-13
Identities = 45/127 (35%), Positives = 65/127 (50%), Gaps = 1/127 (0%)
Frame = +3
Query: 177 EFFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGV-VLGPGVCLNHLQFADD 235
E +++GL+QGDPL+P LF L +G+S + R G V G ++HLQ+ADD
Sbjct: 6 EEISVQRGLKQGDPLAPFLFLLVAEGISGLMKNAVNRNLFQGFDVKRGGTRVSHLQYADD 185
Query: 236 TLVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVGT 295
TL P + L ++ F SGLK+N+ KS L+G NV + L +
Sbjct: 186 TLCIGMPTVDNLWTLKALLQGFEMASGLKVNFHKSSLIGINVPRDFMEAACRFLNCREES 365
Query: 296 LPLPYLG 302
+P YLG
Sbjct: 366 IPFIYLG 386
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 62.8 bits (151), Expect = 6e-10
Identities = 41/115 (35%), Positives = 59/115 (50%), Gaps = 6/115 (5%)
Frame = +1
Query: 182 EKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGV-VLGPGVCLNHLQFADDTLVFC 240
EKGLRQGDPLSP LF LC LS + R ++ G+ V + HL FADD+L+F
Sbjct: 7 EKGLRQGDPLSPYLFILCANVLSGLLKREGNKQNLHGIQVARSDPKITHLLFADDSLLFA 186
Query: 241 DPECAQMEELFDVMTAFLWGSGLKLNYAKSEL-----VGCNVDSAIVNQLAVELG 290
+ + V+ ++ SG +N+ KSE+ V I Q+A++ G
Sbjct: 187 RANLTEAATIMQVLHSYQSASGQLVNFEKSEVSYSQNVPNQEKEMICQQIAIKTG 351
>BG585866
Length = 828
Score = 61.2 bits (147), Expect = 2e-09
Identities = 36/109 (33%), Positives = 57/109 (52%), Gaps = 5/109 (4%)
Frame = +3
Query: 570 DSLIWLGDTNGRYTVKT----FCMEVELQLYGPPSWVVPSCVKKLA-PQKVVLFLWQAAE 624
D+ IW ++NG YT K+ + E Y SW S + +L P+K FLW A
Sbjct: 348 DAYIWPHNSNGVYTAKSGYSWILSQTETVNYNNSSW---SWIWRLKIPEKYKFFLWLACH 518
Query: 625 NRVAVMVNLHNRGVLVDAEAVCVLCGQAEESVTHLILVCKFAWVLWSKV 673
N V + L++R ++ A+C CG+ EES H + C+F+ ++W K+
Sbjct: 519 NAVPTLSLLNHRNMV--NSAICSRCGEHEESFFHCVRDCRFSKIIWHKI 659
>TC82520
Length = 833
Score = 42.7 bits (99), Expect(2) = 2e-08
Identities = 30/116 (25%), Positives = 50/116 (42%), Gaps = 2/116 (1%)
Frame = +2
Query: 649 CGQAEESVTHLILVCKFAWVLWSKVLLREGVVWCVPASILSLLKEWDSLRKGSDLV--LW 706
CG +E + THL L C LWS VL ++ +PA I ++ S+
Sbjct: 317 CGDSE-TATHLFLHCDIFGSLWSHVLRWLHLLLVLPADIRQFFIQFTSMAGSPRFTHSFL 493
Query: 707 RLIPYALAWSVWMARNGLIFKGEEVSVETVWDIHITRMYWWIKAGKPHCSFSLCDF 762
+++ +A W +W RN +F+ T + + W+K + SFS D+
Sbjct: 494 QIMWFASVWVLWKKRNNRVFQNSLSDPSTFVEQVKMHSFLWLKFQQATFSFSYHDW 661
Score = 34.3 bits (77), Expect(2) = 2e-08
Identities = 15/40 (37%), Positives = 21/40 (52%)
Frame = +3
Query: 608 KKLAPQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEAVCV 647
+K P KV +F+W+ NR+ VNL R VL C+
Sbjct: 192 QKNIPSKVSMFVWRLFHNRLPTKVNLMQRHVLQQTHTACI 311
>BG453645
Length = 622
Score = 52.4 bits (124), Expect = 8e-07
Identities = 34/124 (27%), Positives = 58/124 (46%), Gaps = 7/124 (5%)
Frame = +3
Query: 641 DAEAVCVLCGQAEESVTHLILVCKFAWVLWSKVLLREGVVWCVPASILSLLKEWDSLRKG 700
+ + CVLC E+ TH+ L C A ++W +V+ V + +P +L W+ +G
Sbjct: 18 EVSSTCVLCNLKVETTTHIFLHCDVASLVWFRVMKWFDVNFLIPP---NLFVHWECWSEG 188
Query: 701 SDLVL-----WRLIPYALAWSVWMARNGLIFKGEE-VSVETVWDIHITRMYWWI-KAGKP 753
W L+ + W +W RN LIFKG V+ + + +I + W + ++ P
Sbjct: 189 GSANRVTKGHW-LVWHTTIWVLWAKRNDLIFKGLNCVAEDVIEEIKVLS*RWMLERSSTP 365
Query: 754 HCSF 757
C F
Sbjct: 366 SCFF 377
>AW736247 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (4%)
Length = 305
Score = 52.0 bits (123), Expect = 1e-06
Identities = 21/38 (55%), Positives = 28/38 (73%)
Frame = +1
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKG 38
PGPDG NFTF +E+W LIK + +++ EF+ GKL KG
Sbjct: 181 PGPDGVNFTFVKEYWELIKVDFLRVVMEFHTNGKLGKG 294
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 41.6 bits (96), Expect(2) = 1e-06
Identities = 20/46 (43%), Positives = 28/46 (60%)
Frame = +2
Query: 161 CVSTATLAVLVNGSPTEFFEIEKGLRQGDPLSPLLFNLCVQGLSVM 206
CV + VLVN + +GL+QGD LSP +F +CV+GLS +
Sbjct: 104 CVESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIICVEGLSFL 241
Score = 29.6 bits (65), Expect(2) = 1e-06
Identities = 11/32 (34%), Positives = 19/32 (59%)
Frame = +1
Query: 127 LDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWI 158
L +K Y+ ++ D+L +M +GF RW W+
Sbjct: 1 LYISKVYNRVD*DYLKEIMIKMGFNNRWIYWM 96
>BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (16%)
Length = 794
Score = 50.8 bits (120), Expect = 2e-06
Identities = 25/79 (31%), Positives = 49/79 (61%)
Frame = +1
Query: 31 EGGKLVKGLNAGFIALIPKRQTPEMVTDYRLISLIGSIYKLLAKVLAARLQGVMPYLLSE 90
E G L + L+PK+ + + ++R ISL YK+++KVL+ RL+ V+P++++E
Sbjct: 556 EQGNLKRESTKQTSGLVPKKLEAKRLVEFRPISLCNVAYKIVSKVLSKRLKSVLPWIITE 735
Query: 91 NQFSFTKGRQIANCLLIGN 109
Q +F + + I++ +LI +
Sbjct: 736 TQAAFGRRQLISDNILIAH 792
Score = 47.0 bits (110), Expect = 3e-05
Identities = 18/40 (45%), Positives = 27/40 (67%)
Frame = +3
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLN 40
PGPDG N FF++FW + +++ M +EF GKL +G+N
Sbjct: 465 PGPDGMNVFFFQQFWDTMGDDLTSMAQEFLRTGKLEEGIN 584
>TC93101
Length = 675
Score = 50.4 bits (119), Expect = 3e-06
Identities = 57/232 (24%), Positives = 100/232 (42%), Gaps = 9/232 (3%)
Frame = -1
Query: 629 VMVNLHNRGVLVDAEAVCVLCGQAEESVTHLILVCKFAWVLW--SKVLLREGVVWCVPAS 686
V L+ RGV + +C C E+ H+ + C+ +W S++ +R P +
Sbjct: 657 VRXELNKRGV--NCPPLCPRCYFNLETTNHIFMSCERTQRVWFGSQLSIR------FPDN 502
Query: 687 ILSLLKEW--DSLRKGSDLVLWRLIPYALAWSVWMARNGLIFKGEEVSVETV----WDIH 740
+W D++ ++ ++ ++ A+ +S+W ARN IF+ + VS +T+ +
Sbjct: 501 STINFSDWLFDAISNQTEEIIIKIS--AITYSIWHARNKAIFENQFVSEDTIIQ*AQNSI 328
Query: 741 ITRMYWWIKAGKPHCSFSLCDFLQNFGAVTVGQRVERCRQDVWTCPPEGILKWNVDGAAR 800
+ IK P+ S N V + W P ILK N D A
Sbjct: 327 LAYEQATIKPQNPNIVLSSLSATSNTNTTRRRSNV----RSRWQKPLNNILKANCD-ANL 163
Query: 801 GAPGMGGIGGVVRDATGEIRGFFSMSL-GVVFAYEAEVKAIRTALEFSKEFG 851
G G+G ++R+A GE + + + G A AE AI A+ +K++G
Sbjct: 162 QVQGRWGLGCIIRNADGEAKVTATWCINGFDCAATAETYAILAAMYLAKDYG 7
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 49.7 bits (117), Expect = 5e-06
Identities = 44/143 (30%), Positives = 59/143 (40%), Gaps = 4/143 (2%)
Frame = -2
Query: 612 PQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEAVCVLCGQAEESVTHLILVCKFAWVLWS 671
P KV +F W+ +R+ NL RGV+ +CV A ES HL L C + LWS
Sbjct: 566 PLKVSVFAWRLLRDRLPTKSNLIYRGVIPTEAGLCVSGCGALESAQHLFLSCSYFASLWS 387
Query: 672 KVLLREGVVWCVPASILS----LLKEWDSLRKGSDLVLWRLIPYALAWSVWMARNGLIFK 727
V G V V ++LS K S L +LI AW +W RN + F
Sbjct: 386 LVRDWIGFVG-VDTNVLSDHFVQFVHSTGGNKASQSFL-QLIWLLCAWVLWTERNNMCFN 213
Query: 728 GEEVSVETVWDIHITRMYWWIKA 750
+ + D W+KA
Sbjct: 212 DSITPLPRLLDKVKYLSLGWLKA 144
>BG456581
Length = 683
Score = 48.1 bits (113), Expect = 1e-05
Identities = 54/207 (26%), Positives = 86/207 (41%), Gaps = 4/207 (1%)
Frame = +2
Query: 605 SCVKKLAPQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEAVCVLCGQAEESVTHLILVCK 664
SC+ P W+ A +R+ NL +RG + ++C C + E+ HL L CK
Sbjct: 89 SCI----PPSHSFICWRLAHDRLPTDDNLSSRGCAL--VSMCSFCLEQVETSDHLFLRCK 250
Query: 665 FAWVLWSKVL--LREGVVWCVPASILSLLKEWDSLRKGSDLVLWRLIPYALAWSVWMARN 722
F LWS + LR G+ + ++LS L S + DL + ++ + S+W ARN
Sbjct: 251 FVVTLWSWLCSQLRVGLDFSSFKALLSSLPRHCS-SQVRDLYVAAVV--HMVHSIWWARN 421
Query: 723 GLIFKGEEVSVETVW-DIH-ITRMYWWIKAGKPHCSFSLCDFLQNFGAVTVGQRVERCRQ 780
+ F +VS V +H + + + GK C + L F + +
Sbjct: 422 NVRFSSAKVSAHAVQVRVHALIGLSGAVSTGK--CIAADAAILDVFRIPPHRRSMXEMVS 595
Query: 781 DVWTCPPEGILKWNVDGAARGAPGMGG 807
W P +K N DG+ G G
Sbjct: 596 VCWKPPSAPWVKGNTDGSXLNNSGAXG 676
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.142 0.462
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,379,197
Number of Sequences: 36976
Number of extensions: 571904
Number of successful extensions: 3426
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 2451
Number of HSP's successfully gapped in prelim test: 112
Number of HSP's that attempted gapping in prelim test: 910
Number of HSP's gapped (non-prelim): 2637
length of query: 916
length of database: 9,014,727
effective HSP length: 105
effective length of query: 811
effective length of database: 5,132,247
effective search space: 4162252317
effective search space used: 4162252317
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0166.1


