

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0162a.10
(265 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
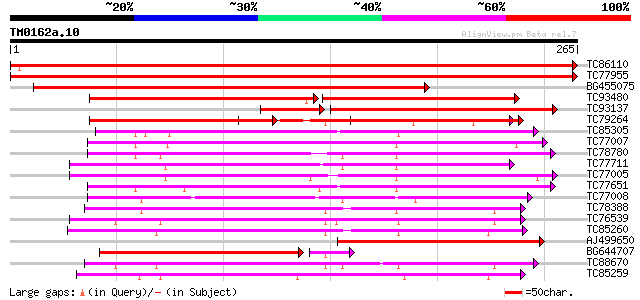
Score E
Sequences producing significant alignments: (bits) Value
TC86110 multifunctional aquaporin [Medicago truncatula] 392 e-110
TC77955 similar to GP|5139541|emb|CAB45652.1 nodulin26-like majo... 380 e-106
BG455075 similar to GP|5139541|emb nodulin26-like major intrinsi... 251 3e-67
TC93480 similar to PIR|B96840|B96840 hypothetical protein F23A5.... 122 3e-52
TC93137 weakly similar to PIR|JQ2285|JQ2285 nodulin-26 - soybean... 119 2e-30
TC79264 similar to PIR|B96840|B96840 hypothetical protein F23A5.... 119 2e-27
TC85305 aquaporin 108 2e-24
TC77007 similar to PIR|T10804|T10804 tonoplast intrinsic protein... 107 4e-24
TC78780 similar to PIR|JQ1106|JQ1106 tonoplast intrinsic protein... 107 4e-24
TC77711 similar to PIR|T46160|T46160 caffeic acid O-methyltransf... 107 6e-24
TC77005 similar to SP|P21653|TIP1_TOBAC Tonoplast intrinsic prot... 103 5e-23
TC77651 similar to PIR|JQ1106|JQ1106 tonoplast intrinsic protein... 101 3e-22
TC77008 similar to PIR|A84653|A84653 hypothetical protein At2g25... 100 6e-22
TC78388 homologue to GP|8071620|gb|AAF71816.1| putative aquapori... 97 8e-21
TC76539 aquaporin [Medicago truncatula] 96 2e-20
TC85260 homologue to GP|3158476|gb|AAC17529.1| aquaporin 2 {Sama... 95 3e-20
AJ499650 weakly similar to PIR|S01444|S014 nodulin-26 precursor ... 94 4e-20
BG644707 homologue to PIR|T04053|T04 nodulin-26 homolog F24G24.1... 94 8e-20
TC88670 similar to GP|535780|dbj|BAA05654.1| transmembrane prote... 92 2e-19
TC85259 homologue to PIR|T09260|T09260 aquaporin-like transmembr... 92 2e-19
>TC86110 multifunctional aquaporin [Medicago truncatula]
Length = 1411
Score = 392 bits (1008), Expect = e-110
Identities = 192/266 (72%), Positives = 228/266 (85%), Gaps = 1/266 (0%)
Frame = +3
Query: 1 MAN-NSASFETHDVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIV 59
MAN NSA ETH+VVL NKD+S T + S ++ SV F+QKL+AE VGT+FLIF GCASIV
Sbjct: 69 MANDNSARIETHEVVLDTNKDSSDTCKGSGSFVSVPFLQKLIAEMVGTYFLIFAGCASIV 248
Query: 60 VNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYI 119
VNK+NDNVVTLPGIA+VWGL L+VLIYS+GHISGAHFNPAVT AFATT+RFP +QV YI
Sbjct: 249 VNKDNDNVVTLPGIAIVWGLTLLVLIYSLGHISGAHFNPAVTIAFATTRRFPLLQVPAYI 428
Query: 120 ASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNR 179
++QLLGA LASG LK++FSG HD FSGT+PSG+NLQAFV+EFITTF LMF IS VATD R
Sbjct: 429 SAQLLGATLASGTLKLIFSGAHDHFSGTLPSGSNLQAFVLEFITTFYLMFTISGVATDTR 608
Query: 180 AIGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVA 239
AIGE+AGIAIGSTLLLN++I+GP+TGASMNP RTLGPA H++YR I +Y +S I GA+A
Sbjct: 609 AIGELAGIAIGSTLLLNVMIAGPVTGASMNPVRTLGPAFVHNEYRGIWIYLLSPILGAIA 788
Query: 240 GAWVFNILRYTDKPLHEITKGSSILK 265
GAWV+N +RYT+KPL EIT+ +S LK
Sbjct: 789 GAWVYNTVRYTNKPLREITQSASFLK 866
>TC77955 similar to GP|5139541|emb|CAB45652.1 nodulin26-like major intrinsic
protein {Pisum sativum}, partial (97%)
Length = 1272
Score = 380 bits (975), Expect = e-106
Identities = 179/265 (67%), Positives = 227/265 (85%)
Frame = +1
Query: 1 MANNSASFETHDVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVV 60
MA NSAS T+++VL+VNKD S E S ++ + F+QKLVAE +GT+FLIF GCAS++V
Sbjct: 145 MAENSASNATNEIVLNVNKDVSNKSEDSTSHATASFLQKLVAEVIGTYFLIFAGCASVLV 324
Query: 61 NKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIA 120
NKNN+NVVTLPGI++VWGL +MVL+YS+GHISGAHFNPAVT AFA+TKRFP QV Y+A
Sbjct: 325 NKNNENVVTLPGISIVWGLAVMVLVYSLGHISGAHFNPAVTIAFASTKRFPLKQVPAYVA 504
Query: 121 SQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRA 180
+Q+ G+ LASG L+++F+G H+QF GT+P+G++LQAFVIEFI TF LMF+IS VATDNRA
Sbjct: 505 AQVFGSTLASGTLRLIFTGKHNQFVGTLPAGSDLQAFVIEFIITFYLMFIISGVATDNRA 684
Query: 181 IGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAG 240
IGE+AGIA+GST+LLN++ +GPITGASMNPAR++GPA+ HS+YR I +Y VS I GAVAG
Sbjct: 685 IGELAGIAVGSTVLLNVMFAGPITGASMNPARSIGPALLHSEYRGIWIYLVSPILGAVAG 864
Query: 241 AWVFNILRYTDKPLHEITKGSSILK 265
AWV+N++RYTDKP+ EITK SS LK
Sbjct: 865 AWVYNVIRYTDKPVREITKSSSFLK 939
>BG455075 similar to GP|5139541|emb nodulin26-like major intrinsic protein
{Pisum sativum}, partial (59%)
Length = 652
Score = 251 bits (640), Expect = 3e-67
Identities = 119/185 (64%), Positives = 154/185 (82%)
Frame = +3
Query: 12 DVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLP 71
DVV+++N DA+K + + V +QKLVAE VGTFFLIF GCA++VVN NND VVTLP
Sbjct: 96 DVVMNINDDATKKCDDTTIDDHVPLLQKLVAEVVGTFFLIFAGCAAVVVNLNNDKVVTLP 275
Query: 72 GIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASG 131
GI++VWGL +MVL+YS+GHISGAHFNPAVT A TT RFP Q+ YI +Q++G+ LASG
Sbjct: 276 GISIVWGLAVMVLVYSIGHISGAHFNPAVTIAHTTTGRFPLKQLPAYIIAQVVGSTLASG 455
Query: 132 ILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGS 191
+LK++FSG +QF+GT+P+G++LQAFV+EFI TF LMF+IS VATDNRAIGE+AG+A+GS
Sbjct: 456 VLKLIFSGKENQFAGTLPAGSDLQAFVVEFIITFFLMFIISGVATDNRAIGELAGLAVGS 635
Query: 192 TLLLN 196
T++LN
Sbjct: 636 TVILN 650
Score = 27.3 bits (59), Expect = 6.2
Identities = 13/46 (28%), Positives = 26/46 (56%)
Frame = -1
Query: 91 ISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKML 136
+ +HF A +F F TT RFP + ++P ++ + L +G+ + +
Sbjct: 148 VVSSHFLVASSFIFITTSRFP-LLISPMFNNEYVRIKLRNGLYQRI 14
>TC93480 similar to PIR|B96840|B96840 hypothetical protein F23A5.11
[imported] - Arabidopsis thaliana, partial (78%)
Length = 902
Score = 122 bits (305), Expect(2) = 3e-52
Identities = 59/92 (64%), Positives = 77/92 (83%)
Frame = +1
Query: 147 TIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGA 206
T+PSG QAF ++FI +F LMFV++AVATD RA+GE+AGIA+G+T++LNILI+GPITGA
Sbjct: 625 TVPSGGYGQAFALKFIISFNLMFVVTAVATDTRAVGELAGIAVGATVMLNILIAGPITGA 804
Query: 207 SMNPARTLGPAIFHSKYRAIVVYFVSTIFGAV 238
SMNP RTLGPAI + Y+AI VY ++ I GA+
Sbjct: 805 SMNPVRTLGPAIAANNYKAIWVYLLAPILGAL 900
Score = 100 bits (250), Expect(2) = 3e-52
Identities = 51/112 (45%), Positives = 69/112 (61%), Gaps = 5/112 (4%)
Frame = +3
Query: 38 QKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFN 97
+K+ AEF+GT L+F G A+ +VN+ TL G A GL +M++I S GHISGAH N
Sbjct: 300 KKIGAEFIGTLILMFAGAATAIVNQKTQGSETLIGCATSTGLAVMIIILSTGHISGAHLN 479
Query: 98 PAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLF-----SGTHDQF 144
PAVT +FA K FPW V YI +Q+L ++ A+ LK +F G+H F
Sbjct: 480 PAVTISFAALKHFPWKHVPMYIGAQILASICAAFSLKAVFHPFMSGGSHRSF 635
Score = 39.3 bits (90), Expect = 0.002
Identities = 28/91 (30%), Positives = 48/91 (51%), Gaps = 8/91 (8%)
Frame = +3
Query: 160 EFITTFLLMFVISAVATDNR-------AIGEMAGIAIGSTLLLNILISGPITGASMNPAR 212
EFI T +LMF +A A N+ IG + G +++ IL +G I+GA +NPA
Sbjct: 315 EFIGTLILMFAGAATAIVNQKTQGSETLIG--CATSTGLAVMIIILSTGHISGAHLNPAV 488
Query: 213 TLG-PAIFHSKYRAIVVYFVSTIFGAVAGAW 242
T+ A+ H ++ + +Y + I ++ A+
Sbjct: 489 TISFAALKHFPWKHVPMYIGAQILASICAAF 581
>TC93137 weakly similar to PIR|JQ2285|JQ2285 nodulin-26 - soybean, partial
(47%)
Length = 633
Score = 119 bits (299), Expect(2) = 2e-30
Identities = 58/106 (54%), Positives = 77/106 (71%)
Frame = +2
Query: 151 GTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNP 210
G+N Q+ V+E I +FLLMFVISAV TD+RA+ + A IA+G TL LN+ I+GP++GASMNP
Sbjct: 110 GSNCQSLVLEIIISFLLMFVISAVTTDDRAVDDSASIAVGMTLTLNLFIAGPVSGASMNP 289
Query: 211 ARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHE 256
AR++GPAI Y+ + +Y V I GA+AGA +N LR KP E
Sbjct: 290 ARSIGPAIVIHIYKGLWIYIVGPIIGAIAGALAYNFLRSAYKPTSE 427
Score = 32.3 bits (72), Expect = 0.19
Identities = 27/108 (25%), Positives = 47/108 (43%), Gaps = 3/108 (2%)
Frame = +2
Query: 38 QKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFN 97
Q LV E + +F L+F ++ D+ ++ G+ L + ++ G +SGA N
Sbjct: 122 QSLVLEIIISFLLMF-----VISAVTTDDRAVDDSASIAVGMTLTLNLFIAGPVSGASMN 286
Query: 98 PAVTFAFATT---KRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHD 142
PA + A + WI + I + GA LA L+ + T +
Sbjct: 287 PARSIGPAIVIHIYKGLWIYIVGPIIGAIAGA-LAYNFLRSAYKPTSE 427
Score = 30.0 bits (66), Expect(2) = 2e-30
Identities = 14/30 (46%), Positives = 20/30 (66%)
Frame = +3
Query: 118 YIASQLLGAVLASGILKMLFSGTHDQFSGT 147
YI +QLLG+ LAS L ++F T + + GT
Sbjct: 9 YILAQLLGSTLASVTLSLMFDITPESYFGT 98
>TC79264 similar to PIR|B96840|B96840 hypothetical protein F23A5.11
[imported] - Arabidopsis thaliana, partial (82%)
Length = 861
Score = 119 bits (297), Expect = 2e-27
Identities = 67/135 (49%), Positives = 91/135 (66%), Gaps = 2/135 (1%)
Frame = +3
Query: 108 KRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG--TIPSGTNLQAFVIEFITTF 165
K + WI + ++AS + AS LK +F H SG T+PS QAF +EFI +F
Sbjct: 480 KMYLWILLHKFLAS-----ICASFTLKGVF---HPFMSGGVTVPSDEYGQAFALEFIISF 635
Query: 166 LLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRA 225
LMF ++AVATD RA+GE+AGIA+G+T++LNILI+GP TGASMN RTL PAI + Y+
Sbjct: 636 NLMFAVTAVATDTRAVGELAGIAVGATVMLNILIAGPATGASMNSVRTLRPAIAANNYKG 815
Query: 226 IVVYFVSTIFGAVAG 240
I +Y ++ I GA+ G
Sbjct: 816 IWLYLIAPILGALGG 860
Score = 91.3 bits (225), Expect = 4e-19
Identities = 45/88 (51%), Positives = 59/88 (66%)
Frame = +2
Query: 38 QKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFN 97
+K+ AEF+GT+ L+F G A+ +VN+ N TL G A GL +M++I S GHISGAH N
Sbjct: 254 KKVGAEFIGTYILMFAGIATAIVNQKIHNSETLIGCAGATGLAVMIIILSTGHISGAHLN 433
Query: 98 PAVTFAFATTKRFPWIQVAPYIASQLLG 125
PAVT +FA K FPW V IA+Q+ G
Sbjct: 434 PAVTISFAALKHFPWKNVPLDIAAQVFG 517
Score = 40.8 bits (94), Expect = 5e-04
Identities = 27/83 (32%), Positives = 45/83 (53%), Gaps = 6/83 (7%)
Frame = +2
Query: 160 EFITTFLLMFVISAVATDNRAIGEMAGI-----AIGSTLLLNILISGPITGASMNPARTL 214
EFI T++LMF A A N+ I + A G +++ IL +G I+GA +NPA T+
Sbjct: 269 EFIGTYILMFAGIATAIVNQKIHNSETLIGCAGATGLAVMIIILSTGHISGAHLNPAVTI 448
Query: 215 G-PAIFHSKYRAIVVYFVSTIFG 236
A+ H ++ + + + +FG
Sbjct: 449 SFAALKHFPWKNVPLDIAAQVFG 517
>TC85305 aquaporin
Length = 1221
Score = 108 bits (271), Expect = 2e-24
Identities = 74/214 (34%), Positives = 109/214 (50%), Gaps = 7/214 (3%)
Frame = +2
Query: 41 VAEFVGTFFLIFTGCAS-IVVNK-NNDNVVTLPGI---ALVWGLVLMVLIYSVGHISGAH 95
+AEF+ TF +F G S I NK ND T G+ ++ L V + +ISG H
Sbjct: 167 LAEFISTFIFVFAGSGSGIAYNKLTNDGAATPAGLISASIAHAFALFVAVSVGANISGGH 346
Query: 96 FNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQ 155
NPAVTF ++ YI +QLLG+++AS +L + + + F + G
Sbjct: 347 VNPAVTFGAFVGGNITLLRGIVYIIAQLLGSIVASALLVFVTASSVPAFGLSEGVGVG-P 523
Query: 156 AFVIEFITTFLLMFVISAVATDNRA--IGEMAGIAIGSTLLLNILISGPITGASMNPART 213
A V+E + TF L++ + A A D + IG +A IAIG + NIL+ G TGASMNPA +
Sbjct: 524 ALVLEIVMTFGLVYTVYATAVDPKKGNIGIIAPIAIGFIVGANILVGGAFTGASMNPAVS 703
Query: 214 LGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNIL 247
GPA+ + VY+ + G V+ +L
Sbjct: 704 FGPAVVSWSWSNHWVYWAGPLIGGGIAGLVYEVL 805
Score = 33.1 bits (74), Expect = 0.11
Identities = 27/121 (22%), Positives = 54/121 (44%), Gaps = 11/121 (9%)
Frame = +2
Query: 154 LQAFVIEFITTFLLMFVISA-------VATDNRAIGE---MAGIAIGSTLLLNILISGPI 203
L+A + EFI+TF+ +F S + D A A IA L + + + I
Sbjct: 155 LKAGLAEFISTFIFVFAGSGSGIAYNKLTNDGAATPAGLISASIAHAFALFVAVSVGANI 334
Query: 204 TGASMNPARTLGPAI-FHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSS 262
+G +NPA T G + + +VY ++ + G++ + + + + P +++G
Sbjct: 335 SGGHVNPAVTFGAFVGGNITLLRGIVYIIAQLLGSIVASALLVFVTASSVPAFGLSEGVG 514
Query: 263 I 263
+
Sbjct: 515 V 517
>TC77007 similar to PIR|T10804|T10804 tonoplast intrinsic protein delta
type - upland cotton, complete
Length = 1041
Score = 107 bits (268), Expect = 4e-24
Identities = 70/223 (31%), Positives = 109/223 (48%), Gaps = 8/223 (3%)
Frame = +2
Query: 37 VQKLVAEFVGTFFLIFTGCAS-IVVNKNNDNVVTLPG----IALVWGLVLMVLIYSVGHI 91
++ +AEF+ T +F G S I K + P +A+ G L V + +I
Sbjct: 128 IKAYIAEFISTLLFVFAGVGSAIAYGKLTSDAALDPAGLLAVAVCHGFALFVAVAVGANI 307
Query: 92 SGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSG 151
SG H NPAVTF A + + Y +QLLG+++A +L+ + G
Sbjct: 308 SGGHVNPAVTFGLAVGGQITILTGIFYWIAQLLGSIVACFLLQFVTGGLETPIHSVAAEV 487
Query: 152 TNLQAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLLNILISGPITGASMN 209
+ V E I TF L++ + A A D + +IG +A IAIG + NIL +GP +G SMN
Sbjct: 488 GPIGGVVTEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFIVGANILAAGPFSGGSMN 667
Query: 210 PARTLGPAIFHSKYRAIVVYFVSTIF-GAVAGAWVFNILRYTD 251
PAR+ GPA+ + +Y+ + G +AG N+ +T+
Sbjct: 668 PARSFGPAVVSGNFHDNWIYWAGPLIGGGLAGLIYGNVFMHTE 796
>TC78780 similar to PIR|JQ1106|JQ1106 tonoplast intrinsic protein alpha -
kidney bean, partial (95%)
Length = 999
Score = 107 bits (268), Expect = 4e-24
Identities = 72/233 (30%), Positives = 114/233 (48%), Gaps = 14/233 (6%)
Frame = +1
Query: 37 VQKLVAEFVGTFFLIFTGCAS--IVVNKNNDNVVT---LPGIALVWGLVLMVLIYSVGHI 91
++ +AEF TF +F G S +V D+ + L +AL L + S H+
Sbjct: 76 IRATIAEFASTFIFVFAGEGSGLALVKIYQDSAFSAGELLAVALAHAFALFAAVSSSMHV 255
Query: 92 SGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSG 151
SG H NPAVTF R ++ Y +QLLGA++A+ +L+++ + P G
Sbjct: 256 SGGHVNPAVTFGALIGGRISVLRAVYYWIAQLLGAIVAALLLRLVTNNMR-------PGG 414
Query: 152 TNL-------QAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLLNILISGP 202
+L ++E I TF LM+ + A A D + +IG +A +AIG + NIL+ GP
Sbjct: 415 FHLARGIGVGHGLILEIIMTFGLMYTVYATAIDPKRGSIGAIAPLAIGLIVGANILVGGP 594
Query: 203 ITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLH 255
GA MNPA GP++ ++ +++V GA A ++ L +P H
Sbjct: 595 FDGACMNPALAFGPSLVGWRWHYHWIFWVGPFIGAALAALIYEYLVIPTEPPH 753
>TC77711 similar to PIR|T46160|T46160 caffeic acid O-methyltransferase-like
protein - Arabidopsis thaliana, partial (86%)
Length = 2361
Score = 107 bits (266), Expect = 6e-24
Identities = 72/216 (33%), Positives = 107/216 (49%), Gaps = 8/216 (3%)
Frame = +2
Query: 29 DTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLP-----GIALVWGLVLMV 83
D SV ++ VAEF+ T +F G S + + L +A+ G L V
Sbjct: 98 DDSFSVSSIRAYVAEFISTLIFVFAGVGSAIAYAKLTSGAALDPAGLVAVAVCHGFALFV 277
Query: 84 LIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQ 143
+ +ISG H NPAVTF A + + Y +QLLG+++A +LK G
Sbjct: 278 AVSVGANISGGHVNPAVTFGLAIGGQITILTGIFYWIAQLLGSIVACFLLKYATGGLTIP 457
Query: 144 FSGTIPSGTNL-QAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLLNILIS 200
++ SG + V E I TF L++ + A A D + ++G +A IAIG + NIL +
Sbjct: 458 IH-SVASGVGAGEGVVTEIIITFGLVYTVYATAADPKKGSLGTIAPIAIGLIVGANILAA 634
Query: 201 GPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFG 236
GP +G SMNPAR+ GPA+ Y +Y+V + G
Sbjct: 635 GPFSGGSMNPARSFGPAVLSGDYHNNWIYWVGPLIG 742
>TC77005 similar to SP|P21653|TIP1_TOBAC Tonoplast intrinsic protein
root-specific RB7-5A (RT-TIP). [Common tobacco]
{Nicotiana tabacum}, partial (97%)
Length = 1055
Score = 103 bits (258), Expect = 5e-23
Identities = 74/241 (30%), Positives = 114/241 (46%), Gaps = 13/241 (5%)
Frame = +1
Query: 29 DTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLP-----GIALVWGLVLMV 83
D S ++ ++EF+ T +F G S + + + L +A+ L V
Sbjct: 139 DDSFSAASLKAYLSEFIATLIFVFAGVGSAIAYNDLTSDAALDPAGLVAVAVAHAFALFV 318
Query: 84 LIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSG---T 140
+ +ISG H NPAVTF A + Y +QLLG+++AS +L + + T
Sbjct: 319 GVAIAANISGGHLNPAVTFGLAIGGNITILTGLFYWIAQLLGSIVASLLLNYVTAKSVPT 498
Query: 141 HDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLLNIL 198
H +G P + V E I TF L++ + A A D + +IG +A IAIG + NIL
Sbjct: 499 HGVAAGLNP----IAGLVFEIIITFGLVYTVYATAADPKKGSIGTIAPIAIGFVVGANIL 666
Query: 199 ISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFN---ILRYTDKPLH 255
+GP +G SMNPAR+ GPA+ + +Y+V + G ++ I YT P
Sbjct: 667 AAGPFSGGSMNPARSFGPAVVSGNFADNWIYWVGPLIGGGLAGLIYGDVFIGSYTPAPAS 846
Query: 256 E 256
E
Sbjct: 847 E 849
>TC77651 similar to PIR|JQ1106|JQ1106 tonoplast intrinsic protein alpha -
kidney bean, partial (98%)
Length = 1154
Score = 101 bits (252), Expect = 3e-22
Identities = 73/227 (32%), Positives = 110/227 (48%), Gaps = 8/227 (3%)
Frame = +3
Query: 37 VQKLVAEFVGTFFLIFTGCAS-IVVNKNNDNVVTLPGIALVWGLV----LMVLIYSVGHI 91
++ VAEF T +F G S + + K + G LV + L I S H+
Sbjct: 183 IRATVAEFFSTCIFVFAGEGSALALRKIYKDAGASSGELLVLAIAHAFSLFAAISSSMHV 362
Query: 92 SGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQ-FSGTIPS 150
SG H NPAVTF R ++ Y +QLLG+ +A+ +L+++ + Q F+
Sbjct: 363 SGGHINPAVTFGALLGGRISVLRALYYWVAQLLGSFVAALLLRLVTNNMRPQAFNVAFGV 542
Query: 151 GTNLQAFVIEFITTFLLMFVISAVATDNR--AIGEMAGIAIGSTLLLNILISGPITGASM 208
G A V+E TF LM+V+ A A D + IG +A +AI + NIL GP GA M
Sbjct: 543 GAG-NALVLEIAMTFGLMYVVYATAIDPKRGTIGTIAPLAIALVVGANILAGGPFDGACM 719
Query: 209 NPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLH 255
NPAR GPA+ ++ +++V GA A ++ + +P H
Sbjct: 720 NPARAFGPALVGWRWHYHWIFWVGPFLGAAIAALLYEYIMVPVEPPH 860
>TC77008 similar to PIR|A84653|A84653 hypothetical protein At2g25810
[imported] - Arabidopsis thaliana, partial (97%)
Length = 967
Score = 100 bits (249), Expect = 6e-22
Identities = 69/214 (32%), Positives = 104/214 (48%), Gaps = 6/214 (2%)
Frame = +2
Query: 37 VQKLVAEFVGTFFLIFTGCASIVV--NKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGA 94
+Q L+ EF+ TF +F G S + + D +V L +A+ LV+ V+I S HISG
Sbjct: 74 IQALIVEFIATFLFVFAGVGSAMTADKLSGDALVGLFFVAIAHALVVAVMI-SAAHISGG 250
Query: 95 HFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNL 154
H NPAVT ++ Y QL+ + A +L L G + T+ SG
Sbjct: 251 HLNPAVTLGLLVGGHITIVRSILYWIDQLIASAAACYLLHYLSGGLTTP-AHTLASGVGY 427
Query: 155 -QAFVIEFITTFLLMFVISAVATDNRAIGEMAGIA---IGSTLLLNILISGPITGASMNP 210
Q V E + TF L+F + A D + G +AG+ +G + NIL G + ASMNP
Sbjct: 428 TQGVVWEIVLTFSLLFTVYATMVDPKK-GALAGLGPTLVGFVVGANILAGGAFSAASMNP 604
Query: 211 ARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVF 244
AR+ GPA+ + VY+V + G +++
Sbjct: 605 ARSFGPALVSGNWTDHWVYWVGPLIGGGLAGFIY 706
>TC78388 homologue to GP|8071620|gb|AAF71816.1| putative aquaporin PIP2-1
{Vitis berlandieri x Vitis rupestris}, partial (91%)
Length = 1120
Score = 96.7 bits (239), Expect = 8e-21
Identities = 66/230 (28%), Positives = 111/230 (47%), Gaps = 24/230 (10%)
Frame = +2
Query: 36 FVQKLVAEFVGTFFLIFTGCASIVV---------NKNNDNVVTLPGIALVWGLVLMVLIY 86
F + L+AEFV T ++ +++ N N + V + GIA +G ++ VL+Y
Sbjct: 140 FYRALIAEFVATLLFLYVTVLTVIGYNAQTDPAHNGTNCDGVGILGIAWAFGGMIFVLVY 319
Query: 87 SVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG 146
ISG H NPAVTF ++ I+ Y+ +Q LGA+ G++K G ++++ G
Sbjct: 320 CTAGISGGHINPAVTFGLFLARKVSLIRAVLYMVAQCLGAICGVGLVKAFQKGYYNRYKG 499
Query: 147 -------TIPSGTNLQAFVIEFITTFLLMFVISAVATDNR-----AIGEMAGIAIGSTLL 194
GT L A E I TF+L++ + + R + +A + IG +
Sbjct: 500 GANMLSAGYSKGTGLGA---EIIGTFVLVYTVFSATDPKRNARDSHVPVLAPLPIGFAVF 670
Query: 195 LNILISGPITGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGA 241
+ L + PITG +NPAR+ G A+ ++ +A +++V GA A
Sbjct: 671 MVHLATIPITGTGINPARSFGAAVIYNNEKAWDDQWIFWVGPFIGAAIAA 820
>TC76539 aquaporin [Medicago truncatula]
Length = 1423
Score = 95.5 bits (236), Expect = 2e-20
Identities = 67/235 (28%), Positives = 114/235 (48%), Gaps = 22/235 (9%)
Frame = +1
Query: 29 DTYTSVFFVQKLVAEFVGTF-FLIFTGCASIVVNKNNDNVVT---------LPGIALVWG 78
D T + ++AEFV T FL T I ++ +D + + GIA +G
Sbjct: 244 DELTKWSLYRAVIAEFVATLLFLYITVLTIIGYSRQSDTTIKGNTECDGVGVLGIAWAFG 423
Query: 79 LVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFS 138
++ +L+Y ISG H NPAVTF ++ I+ YI +Q LGA+ +G+ K +
Sbjct: 424 GMIFILVYCTAGISGGHINPAVTFGLFVGRKVSLIRAFLYIVAQCLGAICGAGLAKGFQT 603
Query: 139 GTHDQFSG---TIPSG-TNLQAFVIEFITTFLLMFVISAVATDNRA-----IGEMAGIAI 189
++++ G T+ G + A E I TF+L++ + + R+ + +A + I
Sbjct: 604 SYYNRYKGGVNTVSDGYSKGTALGAEIIGTFVLVYTVFSATDPKRSARDSHVPVLAPLPI 783
Query: 190 GSTLLLNILISGPITGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGA 241
G + + L + P+TG +NPAR+ GPA+ + +A +Y+V GA A
Sbjct: 784 GFAVFMVHLATIPVTGTGINPARSFGPAVIFNNEKAWDDQWIYWVGPFIGAAVAA 948
>TC85260 homologue to GP|3158476|gb|AAC17529.1| aquaporin 2 {Samanea saman},
complete
Length = 1282
Score = 94.7 bits (234), Expect = 3e-20
Identities = 64/239 (26%), Positives = 115/239 (47%), Gaps = 24/239 (10%)
Frame = +3
Query: 28 SDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNV---------VTLPGIALVWG 78
++ T F + L+AEF+ T ++ +++ +V V + GIA +G
Sbjct: 171 AEELTKWSFYRALIAEFIATLLFLYVTVLTVIGYSIQTDVKAGGDACGGVGILGIAWAFG 350
Query: 79 LVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFS 138
++ VL+Y ISG H NPAVTF ++ I+ Y+ +Q LGA+ G++K S
Sbjct: 351 GMIFVLVYCTAGISGGHINPAVTFGLFLARKVSLIRAIMYMVAQCLGAIAGVGLVKAFQS 530
Query: 139 GTHDQFSG-------TIPSGTNLQAFVIEFITTFLLMFVISAVATDNRA-----IGEMAG 186
D++ G +G L A E + TF+L++ + + R+ + +A
Sbjct: 531 AYFDRYGGGANFLHDGYSTGVGLGA---EIVGTFVLVYTVFSATDPKRSARDSHVPVLAP 701
Query: 187 IAIGSTLLLNILISGPITGASMNPARTLGPAIFHSK---YRAIVVYFVSTIFGAVAGAW 242
+ IG + + L + P+TG +NPAR+LG A+ ++K + +++V GA A+
Sbjct: 702 LPIGFAVFMVHLATIPVTGTGINPARSLGSAVIYNKDKPWDDHWIFWVGPFIGAAIAAF 878
>AJ499650 weakly similar to PIR|S01444|S014 nodulin-26 precursor - soybean,
partial (30%)
Length = 583
Score = 94.4 bits (233), Expect = 4e-20
Identities = 48/97 (49%), Positives = 65/97 (66%)
Frame = -2
Query: 154 LQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPART 213
L+A V E I TF+L+ I VATD+R ++AG+AIG ++L+NI+I+GP T ASMNPAR+
Sbjct: 501 LEALVWESIITFILVLTICGVATDHRGSKDLAGVAIGISVLINIIIAGPTTRASMNPARS 322
Query: 214 LGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYT 250
LGPAI Y+ I VY + GAV ++ LR T
Sbjct: 321 LGPAIVSGNYKNIWVYIIGPTIGAVFATVLYTFLRVT 211
>BG644707 homologue to PIR|T04053|T04 nodulin-26 homolog F24G24.180 -
Arabidopsis thaliana, partial (39%)
Length = 731
Score = 93.6 bits (231), Expect(2) = 8e-20
Identities = 45/95 (47%), Positives = 62/95 (64%)
Frame = +2
Query: 43 EFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTF 102
EFVGTF LIF A +VN+ + +L G A GL +M++I S GHISGAH NP++T
Sbjct: 371 EFVGTFILIFAATAGPIVNQKYNGAESLIGNAACSGLAVMIVILSTGHISGAHLNPSLTI 550
Query: 103 AFATTKRFPWIQVAPYIASQLLGAVLASGILKMLF 137
AFA + FPW+QV Y+A+Q+ ++ AS K+ F
Sbjct: 551 AFAALRHFPWVQVPAYVAAQVSASICASFASKVSF 655
Score = 35.4 bits (80), Expect = 0.023
Identities = 24/96 (25%), Positives = 48/96 (50%), Gaps = 8/96 (8%)
Frame = +2
Query: 159 IEFITTFLLMFVISAVATDNR-------AIGEMAGIAIGSTLLLNILISGPITGASMNPA 211
+EF+ TF+L+F +A N+ IG A G +++ IL +G I+GA +NP+
Sbjct: 368 LEFVGTFILIFAATAGPIVNQKYNGAESLIGNAA--CSGLAVMIVILSTGHISGAHLNPS 541
Query: 212 RTLG-PAIFHSKYRAIVVYFVSTIFGAVAGAWVFNI 246
T+ A+ H + + Y + + ++ ++ +
Sbjct: 542 LTIAFAALRHFPWVQVPAYVAAQVSASICASFASKV 649
Score = 20.4 bits (41), Expect(2) = 8e-20
Identities = 11/23 (47%), Positives = 13/23 (55%), Gaps = 2/23 (8%)
Frame = +1
Query: 141 HDQFSG--TIPSGTNLQAFVIEF 161
H SG T+PS QAF +EF
Sbjct: 655 HPFMSGGVTVPSVNTGQAFALEF 723
>TC88670 similar to GP|535780|dbj|BAA05654.1| transmembrane protein
{Arabidopsis thaliana}, complete
Length = 1106
Score = 92.4 bits (228), Expect = 2e-19
Identities = 70/228 (30%), Positives = 116/228 (50%), Gaps = 16/228 (7%)
Frame = +1
Query: 36 FVQKLVAEFVGTF-FLIFTGCASIVVNKNNDNV--VTLPGIALVWGLVLMVLIYSVGHIS 92
F + +AEFV TF FL T + VNK+++ V + GIA +G ++ L+Y IS
Sbjct: 202 FYRAGIAEFVATFLFLYITVLTVMGVNKSSNKCSSVGVQGIAWAFGGMIFALVYCTAGIS 381
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSG---TIP 149
G H NPAVTF ++ + YI Q LGA+ + ++K +++ G T+
Sbjct: 382 GGHINPAVTFGLFLARKLSLTRTLFYIIMQCLGAICGAAVVKGFQPHQYERLGGGANTLN 561
Query: 150 SGTNL-QAFVIEFITTFLLMFVISAVATDNRA------IGEMAGIAIGSTLLLNILISGP 202
G ++ E + TF+L++ + + ATD + + +A + IG + L L + P
Sbjct: 562 KGYSIGDGLGAEIVGTFVLVYTVFS-ATDAKRNARDSHVPILAPLPIGFAVFLVHLATIP 738
Query: 203 ITGASMNPARTLGPAIFHSKYRA---IVVYFVSTIFGAVAGAWVFNIL 247
ITG +NPAR+LG A+ ++K +A +++V GA A I+
Sbjct: 739 ITGTGINPARSLGAALIYNKDQAWDDHWIFWVGPFTGAALAALYHQIV 882
Score = 35.4 bits (80), Expect = 0.023
Identities = 26/109 (23%), Positives = 53/109 (47%), Gaps = 11/109 (10%)
Frame = +1
Query: 146 GTIPSGTNLQAFVIEFITTFLLMFV-ISAVATDNRAIGEMAGI-------AIGSTLLLNI 197
G + S + +A + EF+ TFL +++ + V N++ + + + A G + +
Sbjct: 181 GELSSWSFYRAGIAEFVATFLFLYITVLTVMGVNKSSNKCSSVGVQGIAWAFGGMIFALV 360
Query: 198 LISGPITGASMNPARTLGPAIFHSKYRAI---VVYFVSTIFGAVAGAWV 243
+ I+G +NPA T G +F ++ ++ + Y + GA+ GA V
Sbjct: 361 YCTAGISGGHINPAVTFG--LFLARKLSLTRTLFYIIMQCLGAICGAAV 501
>TC85259 homologue to PIR|T09260|T09260 aquaporin-like transmembrane channel
protein - alfalfa, complete
Length = 1502
Score = 92.0 bits (227), Expect = 2e-19
Identities = 64/226 (28%), Positives = 109/226 (47%), Gaps = 16/226 (7%)
Frame = +3
Query: 32 TSVFFVQKLVAEFVGTFFLIFTGCASIV-VNKNNDNVVT--LPGIALVWGLVLMVLIYSV 88
TS F + +AEFV TF ++ +++ VN++ T + GIA +G ++ L+Y
Sbjct: 225 TSWSFYRAGIAEFVATFLFLYITILTVMGVNRSESKCKTVGIQGIAWSFGGMIFALVYCT 404
Query: 89 GHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGIL-----KMLFSGTHDQ 143
ISG H NPAVTF ++ + Y+ Q LGA+ +G++ K ++ +
Sbjct: 405 AGISGGHINPAVTFGLFLARKLSLTRAVFYMVMQTLGAICGAGVVKGFEGKTFYTNKNGG 584
Query: 144 FSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGE-----MAGIAIGSTLLLNIL 198
+ P T E + TF+L++ + + R+ + +A + IG + L L
Sbjct: 585 ANFVAPGYTKGDGLGAEIVGTFVLVYTVFSATDAKRSARDSHVPILAPLPIGFAVFLVHL 764
Query: 199 ISGPITGASMNPARTLGPAIFHSK---YRAIVVYFVSTIFGAVAGA 241
+ PITG +NPAR+LG AI +K + +++V GA A
Sbjct: 765 ATIPITGTGINPARSLGAAIIFNKDLGWDDHWIFWVGPFIGAALAA 902
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.138 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,003,476
Number of Sequences: 36976
Number of extensions: 85298
Number of successful extensions: 599
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 558
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 571
length of query: 265
length of database: 9,014,727
effective HSP length: 94
effective length of query: 171
effective length of database: 5,538,983
effective search space: 947166093
effective search space used: 947166093
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0162a.10


