

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
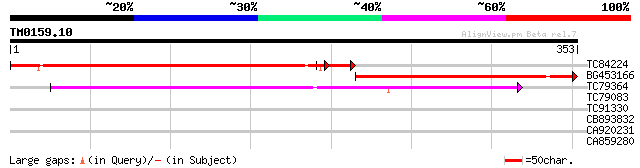
Query= TM0159.10
(353 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC84224 similar to GP|9758839|dbj|BAB09511.1 UDP-galactose trans... 324 2e-96
BG453166 similar to PIR|T49259|T492 hypothetical protein F12M12.... 243 8e-65
TC79364 similar to GP|16649145|gb|AAL24424.1 Unknown protein {Ar... 131 4e-31
TC79083 weakly similar to GP|10177510|dbj|BAB10904. nodulin-like... 32 0.28
TC91330 weakly similar to PIR|A84793|A84793 nodulin-like protein... 31 0.63
CB893832 weakly similar to GP|1813429|dbj| KIAA0160 gene product... 31 0.82
CA920231 25 4.7
CA859280 28 5.3
>TC84224 similar to GP|9758839|dbj|BAB09511.1 UDP-galactose transporter
related protein-like {Arabidopsis thaliana}, partial
(52%)
Length = 798
Score = 324 bits (831), Expect(2) = 2e-96
Identities = 167/204 (81%), Positives = 182/204 (88%), Gaps = 5/204 (2%)
Frame = +2
Query: 1 MAESPSTSSSSSPPDS---RDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYF 57
M+E P +SSSS+ + RDN LWKG FAVSGIM+TLVTYG+LQEKIMR+PYGA+KEYF
Sbjct: 146 MSEPPPSSSSSTTSTAVSFRDN-LWKGTFAVSGIMLTLVTYGLLQEKIMRIPYGAEKEYF 322
Query: 58 KHSLFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVS 117
KHSLFLVFCNRI TSAVSAGSLLASKKALDPVAPIYKY LVSV+NILTTTCQYEALKYVS
Sbjct: 323 KHSLFLVFCNRIMTSAVSAGSLLASKKALDPVAPIYKYSLVSVTNILTTTCQYEALKYVS 502
Query: 118 FPVQTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRG 177
FPVQTLAKCAKMIPVM+WGT+IMQKRY+GPDY+LAFLVTLGCSVFILYPAG DISPYSRG
Sbjct: 503 FPVQTLAKCAKMIPVMVWGTLIMQKRYKGPDYLLAFLVTLGCSVFILYPAGTDISPYSRG 682
Query: 178 RENTVWGVLLMTGYL--GCDGFTS 199
RENTVWG G+L G GFT+
Sbjct: 683 RENTVWG-CSSNGWLSWGLMGFTA 751
Score = 46.6 bits (109), Expect(2) = 2e-96
Identities = 20/24 (83%), Positives = 21/24 (87%)
Frame = +1
Query: 192 LGCDGFTSTFQDKLFRGYDMEIHN 215
LG DG STFQDK+FRGYDMEIHN
Sbjct: 727 LGFDGLYSTFQDKMFRGYDMEIHN 798
>BG453166 similar to PIR|T49259|T492 hypothetical protein F12M12.150 -
Arabidopsis thaliana, partial (28%)
Length = 663
Score = 243 bits (620), Expect = 8e-65
Identities = 120/138 (86%), Positives = 129/138 (92%)
Frame = +1
Query: 216 QIFYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDCFFDIALLSTVATASQFFISYTIRT 275
QIFYTTLCSC+LSLTGLI+QG +I A+EFVYRHHDCFFDIALLSTVAT SQFFISYTIRT
Sbjct: 1 QIFYTTLCSCLLSLTGLIVQGQMISAVEFVYRHHDCFFDIALLSTVATISQFFISYTIRT 180
Query: 276 FGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSLYAKSFTRKAPQKTTSS 335
FGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSLYAKSF +KAPQKTTS
Sbjct: 181 FGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSLYAKSFWKKAPQKTTS- 357
Query: 336 DSIPLVQSGDSNNLKDNP 353
S+ VQ+ +SN+LKDNP
Sbjct: 358 -SVAPVQNENSNDLKDNP 408
>TC79364 similar to GP|16649145|gb|AAL24424.1 Unknown protein {Arabidopsis
thaliana}, complete
Length = 1377
Score = 131 bits (329), Expect = 4e-31
Identities = 94/298 (31%), Positives = 139/298 (46%), Gaps = 4/298 (1%)
Frame = +3
Query: 26 FAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYFKHSLFLVFCNRITTSAVSAGSLLASKKA 85
F V+GI + G+LQE + +G E F+H FL + S +
Sbjct: 132 FCVAGIWSAYIYQGILQETLSTKRFGKDAERFEHLAFLNLAQNVVCLIWSYIMIKIWGGG 311
Query: 86 LDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMIWGTIIMQKRYQ 145
AP + Y ++N + EALKY+S+P Q LAK +KMIPVM+ G+++ RY
Sbjct: 312 NTGGAPWWSYWSAGITNTIGPAMGIEALKYISYPAQVLAKSSKMIPVMLMGSLVYGIRYT 491
Query: 146 GPDYMLAFLVTLGCSVF-ILYPAGEDISPYSRGRENTVWGVLLMTGYLGCDGFTSTFQDK 204
P+Y+ FLV G S F +L + + IS + +G+ + L DGFT+ QD
Sbjct: 492 IPEYLCTFLVAGGVSSFALLKTSSKTISKLAHPNAPLGYGLCFLN--LAFDGFTNATQDS 665
Query: 205 LFRGY-DMEIHNQIFYTTLCSCILSLTGLIL--QGHLIPAIEFVYRHHDCFFDIALLSTV 261
L Y + L I ++ + G A+ F +H + +DI L
Sbjct: 666 LKARYPKTSAWGIMLGMNLWGTIYNMIYMFAWPSGSGYEAVNFCKQHPEAAWDILLYCCC 845
Query: 262 ATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSL 319
Q FI TI FG+L TI TTR+ VSI++S + +PLS +QW +VF L
Sbjct: 846 GAVGQNFIFLTISRFGSLANTTITTTRKFVSIVVSSLLSGNPLSTKQWGCVTMVFSGL 1019
>TC79083 weakly similar to GP|10177510|dbj|BAB10904. nodulin-like protein
{Arabidopsis thaliana}, partial (31%)
Length = 1256
Score = 32.3 bits (72), Expect = 0.28
Identities = 53/317 (16%), Positives = 114/317 (35%), Gaps = 30/317 (9%)
Frame = +2
Query: 52 AQKEYFKHSLFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYE 111
A K+ +F+ + N + + +L +K P +C + + L T Q
Sbjct: 59 ATKDGMSIFIFIFYSNLLALCFLLPSTLFHHRKRAPPPISTSIFCRLLLLGCLKTATQTL 238
Query: 112 ALKYVSFPVQTLAKC-AKMIPVMIWGTIIMQ-------KRYQGPDYMLAFLVTLGCSVFI 163
+ LA ++P + I+ K++ ++ +V++ ++ +
Sbjct: 239 MTSGIKLSSPALASAIVNLVPAFTFILAIISRMEVLNMKKHSSQAKVIGTVVSIAGALVV 418
Query: 164 LYPAGEDI-------------SPYSRGRENTVWGVLLMTGYLGCDGFTSTFQDKLFRGYD 210
+ G + + G+ + + G L+ C Q + R Y
Sbjct: 419 TFYKGIPLINDAIKNIEMGASGIFLLGKSDWIIGAFLVAVGSFCLSLLFIVQTWIVRDYP 598
Query: 211 MEIHNQIFYTTLCSC----ILSLTGLILQGH-----LIPAIEFVYRHHDCFFDIALLSTV 261
E+ T+C C I ++ LI +G+ L P E + + F +L + +
Sbjct: 599 EEL----VINTICCCFVVIISTIVALIAEGNSNAWRLRPDKELLSVCYSATFVESLKNVI 766
Query: 262 ATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSLYA 321
T + R G + A R ++++ + ++ L IGA I+ YA
Sbjct: 767 QT-------WACRKKGPIYVAMFKPLRVVIALSMGVIFLGDNLYLGSMIGAAIIVIGFYA 925
Query: 322 KSFTRKAPQKTTSSDSI 338
+ + + TTS +++
Sbjct: 926 VIWAKAQEEHTTSENNL 976
>TC91330 weakly similar to PIR|A84793|A84793 nodulin-like protein [imported]
- Arabidopsis thaliana, partial (34%)
Length = 812
Score = 31.2 bits (69), Expect = 0.63
Identities = 27/97 (27%), Positives = 39/97 (39%), Gaps = 11/97 (11%)
Frame = +3
Query: 253 FDIALLSTVATASQFFIS-YTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIG 311
F A V + Q++I I+T G + R ++ L+C+ S L IG
Sbjct: 210 FAPAYAGIVTSGVQYYIQGLVIKTMGPVIVTAFNPVRMIIVTALACIILSEQLFLGSIIG 389
Query: 312 AVIVFGSLY----AKSFTRKA------PQKTTSSDSI 338
A++V LY KS KA P T DS+
Sbjct: 390 AIVVVLGLYLVVWGKSKEYKARNHVDMPPSPTKEDSL 500
>CB893832 weakly similar to GP|1813429|dbj| KIAA0160 gene product is novel.
{Homo sapiens}, partial (2%)
Length = 730
Score = 30.8 bits (68), Expect = 0.82
Identities = 14/51 (27%), Positives = 28/51 (54%)
Frame = -2
Query: 238 LIPAIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTR 288
L+P ++ + + FF++ L ST+A + Q I+ T ++T ++ TR
Sbjct: 210 LLPCLDLTFDSNSMFFNLPLKSTIAISLQNLITTTTTFPSSITIRQLLQTR 58
>CA920231
Length = 687
Score = 25.4 bits (54), Expect(2) = 4.7
Identities = 9/26 (34%), Positives = 14/26 (53%)
Frame = -3
Query: 214 HNQIFYTTLCSCILSLTGLILQGHLI 239
HN + Y C C+ S+ + L HL+
Sbjct: 370 HNVLEYVEFCGCVCSINVIQLATHLL 293
Score = 21.2 bits (43), Expect(2) = 4.7
Identities = 7/13 (53%), Positives = 10/13 (76%)
Frame = -1
Query: 176 RGRENTVWGVLLM 188
RGR+ +WG L+M
Sbjct: 402 RGRKANMWGFLIM 364
>CA859280
Length = 421
Score = 28.1 bits (61), Expect = 5.3
Identities = 18/57 (31%), Positives = 26/57 (45%), Gaps = 2/57 (3%)
Frame = +2
Query: 290 LVSIMLSCVWFSHPLSWEQWIGAVIVFGSLYAKSFTRKAPQ--KTTSSDSIPLVQSG 344
+ S++LS V + W WI V SL + T + P +T+S SI SG
Sbjct: 134 IFSLLLSTVGLIMKIKWASWISLVFAIISLTNEKATDQDPNSGRTSSITSILFATSG 304
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.137 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,553,783
Number of Sequences: 36976
Number of extensions: 187579
Number of successful extensions: 1392
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1371
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1389
length of query: 353
length of database: 9,014,727
effective HSP length: 97
effective length of query: 256
effective length of database: 5,428,055
effective search space: 1389582080
effective search space used: 1389582080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0159.10


