

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0157a.4
(594 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
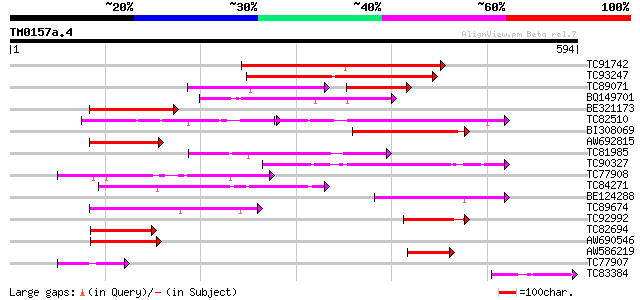
Score E
Sequences producing significant alignments: (bits) Value
TC91742 similar to GP|18481720|gb|AAL73542.1 hypothetical protei... 183 2e-46
TC93247 similar to GP|15810641|gb|AAL07245.1 unknown protein {Ar... 157 1e-38
TC89071 similar to PIR|G96701|G96701 unknown protein 76653-7450... 96 5e-34
BQ149701 similar to PIR|G86160|G86 protein F10O3.17 [imported] -... 141 8e-34
BE321173 weakly similar to GP|9758879|dbj| non-phototropic hypoc... 135 6e-32
TC82510 weakly similar to GP|14532868|gb|AAK64116.1 unknown prot... 102 1e-28
BI308069 similar to GP|15451228|gb photoreceptor-interacting pro... 122 4e-28
AW692815 weakly similar to GP|18481720|gb| hypothetical protein ... 117 1e-26
TC81985 similar to PIR|G96568|G96568 probable non-phototropic hy... 113 2e-25
TC90327 similar to GP|19310601|gb|AAL85031.1 unknown protein {Ar... 104 8e-23
TC77908 similar to GP|15810641|gb|AAL07245.1 unknown protein {Ar... 100 3e-21
TC84271 weakly similar to PIR|T02594|T02594 hypothetical protein... 98 1e-20
BE124288 similar to GP|6224712|gb| non-phototropic hypocotyl 3 {... 89 4e-18
TC89674 similar to PIR|T51779|T51779 non-phototropic hypocotyl 3... 79 6e-15
TC92992 similar to PIR|T02594|T02594 hypothetical protein At2g14... 63 3e-10
TC82694 similar to GP|6224712|gb|AAF05914.1| non-phototropic hyp... 58 9e-09
AW690546 similar to GP|6634762|gb| Contains a bZIP transcription... 55 6e-08
AW586219 52 5e-07
TC77907 similar to GP|15810641|gb|AAL07245.1 unknown protein {Ar... 47 3e-05
TC83384 similar to GP|15810641|gb|AAL07245.1 unknown protein {Ar... 46 4e-05
>TC91742 similar to GP|18481720|gb|AAL73542.1 hypothetical protein {Sorghum
bicolor}, partial (32%)
Length = 669
Score = 183 bits (464), Expect = 2e-46
Identities = 97/218 (44%), Positives = 142/218 (64%), Gaps = 5/218 (2%)
Frame = +1
Query: 244 PQHEHEKRVVVETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQ 303
P E ++++++E + LLP +K + L LLR A+ L +C LEKR+ MQL Q
Sbjct: 16 PPSEEDQKILLEEIDQLLPMQKGLVQTKFLFGLLRTAMILRVNPSCISSLEKRIGMQLDQ 195
Query: 304 AVLDDLLIPSYSFPGNTLFDVDTVQRIVVSYLEFE--IGNHSVSSADDE---YFSPSQRG 358
A L+DLL+P++S+ TL++VD VQRI+ +L + G S S D++ SPS
Sbjct: 196 AALEDLLMPNFSYSMETLYNVDCVQRILDHFLAMDQITGGASPCSIDEDGQLIGSPSLTP 375
Query: 359 IVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSE 418
I V KL++ YLAE+A D NL KF ++A +PE + P +DG YRAIDIYLK+HP+L E
Sbjct: 376 ITMVAKLIDGYLAEVAPDVNLKLPKFQALAAAVPEYARPLDDGLYRAIDIYLKSHPWLVE 555
Query: 419 MEKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVL 456
+++ +C +M CQKLS +A HAAQN+RLP++ ++QVL
Sbjct: 556 SDREQLCRLMDCQKLSLEACTHAAQNERLPIRIIVQVL 669
>TC93247 similar to GP|15810641|gb|AAL07245.1 unknown protein {Arabidopsis
thaliana}, partial (36%)
Length = 677
Score = 157 bits (396), Expect = 1e-38
Identities = 85/200 (42%), Positives = 127/200 (63%)
Frame = +1
Query: 249 EKRVVVETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDD 308
++R ++E +++L P EK A ++ L LLR AI+L + C+ +LEKR++ L +DD
Sbjct: 25 KQRSLLELIINLFPSEKAAFPINFLCCLLRCAIHLRASTVCKTELEKRISAILEHVTVDD 204
Query: 309 LLIPSYSFPGNTLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSPSQRGIVRVGKLMEN 368
LL+ S+S+ G LFD+++V+RI+ ++E E ++ E ++RV K ++
Sbjct: 205 LLVMSFSYDGEKLFDLESVRRIISVFVEKE--KNTAVFTPGELKQSFSVPMLRVAKTVDM 378
Query: 369 YLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVCSVM 428
YLAEIAA L SKF IA LIP+ + +D YRA+DIYLK HP L E+E++ VCSVM
Sbjct: 379 YLAEIAAYGELSISKFNGIAVLIPKHARKIDDDLYRAVDIYLKVHPKLDEIEREKVCSVM 558
Query: 429 HCQKLSRDARAHAAQNDRLP 448
KLS +AR HA+QN R+P
Sbjct: 559 DPLKLSYEARVHASQNKRVP 618
>TC89071 similar to PIR|G96701|G96701 unknown protein 76653-74502
[imported] - Arabidopsis thaliana, partial (42%)
Length = 947
Score = 95.5 bits (236), Expect(2) = 5e-34
Identities = 55/161 (34%), Positives = 91/161 (56%), Gaps = 12/161 (7%)
Frame = +2
Query: 187 WWGEDLTVLRIDIFQRVLIAMMGRGFKQFDL-GPIIMLYAQKSLEGLEIFGKGRKELEPQ 245
WW EDL L ID++ R +IA+ G +L G + +YA + L + G +K +
Sbjct: 140 WWAEDLAELSIDLYWRTMIAIKSGGKVPSNLIGDALKIYASRWLPNISKNGNRKKNVSLA 319
Query: 246 HEHEK-----------RVVVETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLE 294
R+++E++VSLLP EK A+S S L LL+A+ L + + +++L
Sbjct: 320 ESESNSDSASEITSKHRLLLESIVSLLPTEKGAVSCSFLLKLLKASNILNASSSSKMELA 499
Query: 295 KRVAMQLGQAVLDDLLIPSYSFPGNTLFDVDTVQRIVVSYL 335
+RV +QL +A ++DLLIPS S+ +TL+DV+ V I+ ++
Sbjct: 500 RRVGLQLEEATVNDLLIPSLSYTNDTLYDVELVMTILEQFM 622
Score = 67.8 bits (164), Expect(2) = 5e-34
Identities = 34/69 (49%), Positives = 47/69 (67%)
Frame = +3
Query: 353 SPSQRGIVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKA 412
S S ++V KL++ YL E+A D N SKFI++A++IPE + D YRAIDIYLKA
Sbjct: 741 SASHSSKLKVAKLVDRYLQEVARDVNFSLSKFIALADIIPEFARYDHDDLYRAIDIYLKA 920
Query: 413 HPFLSEMEK 421
HP L++ E+
Sbjct: 921 HPELNKSER 947
>BQ149701 similar to PIR|G86160|G86 protein F10O3.17 [imported] - Arabidopsis
thaliana, partial (26%)
Length = 649
Score = 141 bits (355), Expect = 8e-34
Identities = 88/221 (39%), Positives = 134/221 (59%), Gaps = 15/221 (6%)
Frame = +1
Query: 200 FQRVLIAMMGRGFKQFDLGPIIMLYAQKSLEGLEIFGKGRKELEPQHEHEKRVVVETLVS 259
F+RVL AM +G KQ + I++ YA SL+ + KG L+ + ++RV+VET+ S
Sbjct: 1 FKRVLSAMKSKGLKQDIISKILINYAHNSLQVV----KGNL-LDLEFLKKQRVLVETITS 165
Query: 260 LLPRE--KNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFP 317
LLP + K+ + ++ LS LL++AI + +CR DLE+R+ +QL QA+L+D+LIP+ +
Sbjct: 166 LLPTQSRKSQIPIAFLSSLLKSAIASSVSTSCRSDLERRIGLQLDQAILEDILIPTSLYQ 345
Query: 318 GN----TLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYF---------SPSQRGIVRVGK 364
N T++D D++ RI +L + + S DE SP Q I++V K
Sbjct: 346 NNNHSSTIYDTDSIVRIFSIFLNLDEEDEDDSRMRDETEMVYDFDSPGSPKQSSILKVSK 525
Query: 365 LMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRA 405
L++NYLAE+A D NL SKFIS+AEL+P+ + DG YRA
Sbjct: 526 LLDNYLAEVALDPNLLPSKFISLAELLPDHARIVSDGLYRA 648
>BE321173 weakly similar to GP|9758879|dbj| non-phototropic hypocotyl-like
protein {Arabidopsis thaliana}, partial (8%)
Length = 321
Score = 135 bits (339), Expect = 6e-32
Identities = 65/93 (69%), Positives = 81/93 (86%)
Frame = +2
Query: 84 MLRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVN 143
MLRCVAEYLEMTED S+GNLV R+D YLN+VAL+TI+G+VS+LH+ E+LL +AEK KLV+
Sbjct: 41 MLRCVAEYLEMTEDYSVGNLVARTDAYLNDVALKTIAGSVSILHISESLLPIAEKTKLVS 220
Query: 144 RCIDAIAYMASKQSQLCSPARSQCSSDGVMSSS 176
+CIDAIAY+A +SQLC+ RS+ SDGVMSSS
Sbjct: 221 KCIDAIAYIACNESQLCTSVRSESGSDGVMSSS 319
>TC82510 weakly similar to GP|14532868|gb|AAK64116.1 unknown protein
{Arabidopsis thaliana}, partial (31%)
Length = 1716
Score = 102 bits (253), Expect(2) = 1e-28
Identities = 73/257 (28%), Positives = 126/257 (48%), Gaps = 11/257 (4%)
Frame = +1
Query: 278 RAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFPGNTLFDVDTVQRIVVSYLEF 337
R + L + R LE + QL Q LD+LL+PS + N L+DV+ V R + ++L
Sbjct: 820 RVTLCLNISKCSRNKLEIMIGSQLDQVTLDNLLVPS-PYGINYLYDVNLVLRFLKAFLRQ 996
Query: 338 EIGNHSVSSADDEYFSPSQRGIVRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIP 397
G + + + +V L++ Y+AEIA D L SKF+++A +P+ +
Sbjct: 997 GTG------------AVAPIRMRKVASLIDLYIAEIAPDPCLKTSKFLALATALPDSARD 1140
Query: 398 TEDGKYRAIDIYLKAHPFLSEMEKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLS 457
+ D Y A+D+YL+ HP LS+ E+ +C ++ +KLS A H ++N + P ++ +Q L
Sbjct: 1141 SCDELYHAMDMYLEVHPQLSQDERIKICCGLNYEKLSPQACLHLSKNTKFPSKSAVQALI 1320
Query: 458 FQQKHLRETMNDGGVNWDGTSIPDKLNVYSAELNPVSKVISN-----------LRRENEE 506
QQ L+ + T D L S K SN ++ +NE+
Sbjct: 1321 SQQSKLKNLLLTTTTTPISTPYNDSLCGSSGSSQKGKKEKSNEQVVLYAGNFDIQPDNEK 1500
Query: 507 LKREIVKLKMKLQEIEK 523
+K + ++ ++ E+EK
Sbjct: 1501 MKEHLQGMQWRVMELEK 1551
Score = 43.1 bits (100), Expect(2) = 1e-28
Identities = 49/219 (22%), Positives = 96/219 (43%), Gaps = 10/219 (4%)
Frame = +3
Query: 76 NASFSLHKMLRCVAEYLEMTED-NSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQ 134
N S S + R AEY+EM E ++ NL+ +++ + ++ + S + L + LL
Sbjct: 228 NLSPSNLLLARYAAEYMEMNESVKNVSNLLDQTEKSVQGISYWSWSEILITLKQCQNLL- 404
Query: 135 VAEKAKLVNRCIDAIAYMASKQSQLCSPARSQCSSD--GVMSSSMASHRRPVVH------ 186
V + + ++ +C+DA+ S+ SP S CS+D G+ S + +
Sbjct: 405 VDDSSVMLEKCLDAVVGRLVSASE-ASPCPSSCSNDSSGIRFSCDSKSTESIKTNFTRST 581
Query: 187 WWGEDLTVLRIDIFQRVLIAMMGRGFKQFDLGPIIMLYAQKSLEGLEIFGKGRKELEPQH 246
WW EDL L + ++ +M+ + + +LY QK+ +
Sbjct: 582 WWFEDLLFLTPQMVIMLVKSMLSSKLDHYVISR-FLLYYQKA------------KFATAT 722
Query: 247 EHEKRVVVETLVSL-LPREKNAMSVSSLSMLLRAAIYLE 284
EK ++E ++ + ++N++S +L +L LE
Sbjct: 723 TDEKCKIIEMVIDMHYEIDRNSVSFKTLFGILSGDFVLE 839
>BI308069 similar to GP|15451228|gb photoreceptor-interacting protein-like
{Arabidopsis thaliana}, partial (19%)
Length = 522
Score = 122 bits (306), Expect = 4e-28
Identities = 60/122 (49%), Positives = 84/122 (68%)
Frame = +2
Query: 360 VRVGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEM 419
++V KL+++YLAEIA D NLP F++IA+L+ P+ DG YRAID+YLK HP +S+
Sbjct: 20 IKVAKLVDSYLAEIARDPNLPLLSFVNIADLVSSFPRPSHDGLYRAIDMYLKEHPGISKS 199
Query: 420 EKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETMNDGGVNWDGTSI 479
E+K +C +M C+KLS +A HA QN+RLP++ V+QVL F+Q T + G GTS
Sbjct: 200 ERKRICRLMDCRKLSAEACMHAVQNERLPLRVVVQVLFFEQMRASTTSSSG-----GTST 364
Query: 480 PD 481
PD
Sbjct: 365 PD 370
>AW692815 weakly similar to GP|18481720|gb| hypothetical protein {Sorghum
bicolor}, partial (14%)
Length = 660
Score = 117 bits (294), Expect = 1e-26
Identities = 57/78 (73%), Positives = 70/78 (89%)
Frame = +3
Query: 84 MLRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVN 143
MLRCVAEYLEMTED S+GNLVGR+D YLNEVAL+++SGA+S+LH ETLL +AEKAKLV+
Sbjct: 390 MLRCVAEYLEMTEDYSVGNLVGRTDSYLNEVALKSMSGAISILHTSETLLPIAEKAKLVS 569
Query: 144 RCIDAIAYMASKQSQLCS 161
RCIDAIA++AS++ Q S
Sbjct: 570 RCIDAIAFIASRRGQFGS 623
>TC81985 similar to PIR|G96568|G96568 probable non-phototropic hypocotyl
25081-26618 [imported] - Arabidopsis thaliana, partial
(42%)
Length = 745
Score = 113 bits (282), Expect = 2e-25
Identities = 74/219 (33%), Positives = 117/219 (52%), Gaps = 6/219 (2%)
Frame = +2
Query: 188 WGEDLTVLRIDIFQRVLIAMMGRGFKQFDLGPIIMLYAQKSLEGLEIFGKGRKELEPQHE 247
W +D T+L +D F + L + +G + +G II YA K L ++ KG + E E
Sbjct: 128 WFDDATILDMDYFVKTLSGIKAKGVRADLIGSIITHYASKWLP--DVAEKGITQFEESPE 301
Query: 248 H------EKRVVVETLVSLLPREKNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQL 301
+KR VETLV++LP EK+++ + L LLR A + + R +LEKR++ QL
Sbjct: 302 SVTTSWMKKRFFVETLVNVLPPEKDSIPCNFLLRLLRTANMVGVDGSYRQELEKRISWQL 481
Query: 302 GQAVLDDLLIPSYSFPGNTLFDVDTVQRIVVSYLEFEIGNHSVSSADDEYFSPSQRGIVR 361
QA L +L+IPS+S TL D + V R+V ++ D + S +V+
Sbjct: 482 DQACLKELVIPSFSHTCGTLLDFELVIRLVKRFVSL-----------DNEGAKSGAALVK 628
Query: 362 VGKLMENYLAEIAADRNLPASKFISIAELIPEQSIPTED 400
V KL+++YL E A D NL +F+++A +P + T+D
Sbjct: 629 VAKLVDSYLCEAAVDANLRLDEFVTLAGALPSHATATDD 745
>TC90327 similar to GP|19310601|gb|AAL85031.1 unknown protein {Arabidopsis
thaliana}, partial (22%)
Length = 798
Score = 104 bits (260), Expect = 8e-23
Identities = 81/259 (31%), Positives = 125/259 (47%)
Frame = +3
Query: 265 KNAMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFPGNTLFDV 324
K + L +LR + R +LEK + L QA LDDLL+ + +DV
Sbjct: 6 KKNFTCRGLFWVLRIVSRFGLSKGRRTELEKLIGGMLEQATLDDLLVSGHDM--GIYYDV 179
Query: 325 DTVQRIVVSYLEFEIGNHSVSSADDEYFSPSQRGIVRVGKLMENYLAEIAADRNLPASKF 384
+ V R+V +++ ++ D S + + RVG+L++ YL EI+ D+NL SKF
Sbjct: 180 NLVIRLVRQFVD--------TNGSD---GMSLQKMKRVGRLIDKYLREISPDQNLKISKF 326
Query: 385 ISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVCSVMHCQKLSRDARAHAAQN 444
+ +AE +P+ + DG Y+AIDIYL++H + E+ +C ++ KLS A A+N
Sbjct: 327 LGVAECLPDSARDCFDGVYKAIDIYLESHTNVPFEERSRLCKCLNYGKLSFGASKDLAKN 506
Query: 445 DRLPVQTVLQVLSFQQKHLRETMNDGGVNWDGTSIPDKLNVYSAELNPVSKVISNLRREN 504
R+P +Q L QQ + T N T P K N S I L +E
Sbjct: 507 PRIPPGVAMQALISQQTKI-PTSN------FVTESPRKSRSQIVLYNEAS--IDGLSQEK 659
Query: 505 EELKREIVKLKMKLQEIEK 523
+ +K + LK K E+EK
Sbjct: 660 KHMKLNLEVLKYKAVELEK 716
>TC77908 similar to GP|15810641|gb|AAL07245.1 unknown protein {Arabidopsis
thaliana}, partial (37%)
Length = 960
Score = 99.8 bits (247), Expect = 3e-21
Identities = 80/297 (26%), Positives = 127/297 (41%), Gaps = 70/297 (23%)
Frame = +2
Query: 51 SSAMKKTSEWNFPQEIPSDVTVQIDNASFSLHKMLR------------------------ 86
S AM++T +W F QEIPSDV V I A++SLHK +
Sbjct: 125 SLAMERTGQWIFSQEIPSDVIVTIGEATYSLHKFMLVAKSNYIRKLIMESNESELTRLDL 304
Query: 87 ----------------CVAEYLEMTEDNSI-----------------GNLVGRSDFYLNE 113
C E+T +N GNL GR++ +L++
Sbjct: 305 SDIPGGQEAFEKAAKFCYGVNFEITVNNVASLHCAAMFLQMTDEYCDGNLAGRTEGFLSQ 484
Query: 114 VALETISGAVSVLHMPETLLQVAEKAKLVNRCIDAIAYMASKQSQLCSPARSQCSSDGVM 173
VAL T+SGA VL L+ VA++ +V RC+DA++ S++C+ A
Sbjct: 485 VALTTLSGAAVVLKSCRQLIPVADELNIVKRCVDAVS------SKVCNEA---------- 616
Query: 174 SSSMASHRRPVVHWWGEDLTVLRIDIFQRVLIAMMGRGFKQFDLGPIIMLYAQKSLE--- 230
+ S P +WW E+L VL ++ F +V+ AM RG K L ++ Y +++L
Sbjct: 617 --NFPSRSPP--NWWTEELAVLDVESFGKVITAMKQRGAKYLTLAGALIAYTERTLRELV 784
Query: 231 ----------GLEIFGKGRKELEPQHEHEKRVVVETLVSLLPREKNAMSVSSLSMLL 277
G + + + + + E+R + ++V L P EK A ++ L LL
Sbjct: 785 RDQTGGGSGGGGKFRSSDSTDSDSETKSEQREIPTSIVPLFPSEKAAFPINFLCCLL 955
>TC84271 weakly similar to PIR|T02594|T02594 hypothetical protein At2g14820
[imported] - Arabidopsis thaliana, partial (35%)
Length = 786
Score = 97.8 bits (242), Expect = 1e-20
Identities = 76/251 (30%), Positives = 130/251 (51%), Gaps = 9/251 (3%)
Frame = +2
Query: 94 MTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNRCIDAIAYMA 153
M E+ GNL+ + D +L+ + ++ +L +++ + E+ K+VNRCI++IA A
Sbjct: 8 MHENIEKGNLLFKIDVFLSSSIFRSWKDSIILLQTTKSMSPLDEEQKVVNRCIESIANKA 187
Query: 154 ----SKQSQLCSPARSQCSSDGVMSSSMASHRRPVV--HWWGEDLTVLRIDIFQRVLIAM 207
SK + R + + + S+ R V WW EDL L +D+++ V+ +
Sbjct: 188 CVDVSKVDWSYTYNRKKLPEENGLESNQNGVRTRNVPKDWWVEDLCELEVDMYKSVITNI 367
Query: 208 MGRGFKQFD-LGPIIMLYAQKSLEGLEIFGKGRKELEPQHEHEKRVVVETLVSLLPREKN 266
+ + D +G + YA + L F KG E +H R++VET+V LLP +K
Sbjct: 368 KTKEIQSNDVIGEALKAYAYRKLPN---FSKGMIPCEDVSKH--RLIVETIVKLLPSDKG 532
Query: 267 AMSVSSLSMLLRAAIYLETTVACRLDLEKRVAMQLGQAVLDDLLIPSYSFPGN--TLFDV 324
++S L LL+A I++E+ R L KR+ QL +A ++D+LI + P T++DV
Sbjct: 533 SVSCRFLVKLLKAVIFVESEDRIRDVLVKRIGQQLEEASVNDILIKA---PDGEITMYDV 703
Query: 325 DTVQRIVVSYL 335
VQ+ V +L
Sbjct: 704 GIVQKNVREFL 736
>BE124288 similar to GP|6224712|gb| non-phototropic hypocotyl 3 {Arabidopsis
thaliana}, partial (25%)
Length = 621
Score = 89.4 bits (220), Expect = 4e-18
Identities = 51/153 (33%), Positives = 84/153 (54%), Gaps = 12/153 (7%)
Frame = +1
Query: 383 KFISIAELIPEQSIPTEDGKYRAIDIYLKAHPFLSEMEKKNVCSVMHCQKLSRDARAHAA 442
KF +AE +PE + ++DG YRA+D YL+AH L+E ++K +C VM CQKLS DA HAA
Sbjct: 1 KFQVVAEALPESARVSDDGLYRAMDSYLRAHATLTEHQRKRLCRVMDCQKLSIDACMHAA 180
Query: 443 QNDRLPVQTVLQVLSFQQKHLRETMNDGGVNWD------------GTSIPDKLNVYSAEL 490
QN+RLP++ V+QVL +Q + + + + T + +
Sbjct: 181 QNERLPLRVVVQVLFSEQVKISNALANSSLKGSVESQYQPMVTNRKTLLERTPQSFQEGW 360
Query: 491 NPVSKVISNLRRENEELKREIVKLKMKLQEIEK 523
K I+ L+ E E +K + ++L+ ++ ++K
Sbjct: 361 TTAKKDINTLKFELESVKTKYLELQHDMENLQK 459
>TC89674 similar to PIR|T51779|T51779 non-phototropic hypocotyl 3-like
protein - Arabidopsis thaliana, partial (35%)
Length = 1129
Score = 78.6 bits (192), Expect = 6e-15
Identities = 56/199 (28%), Positives = 95/199 (47%), Gaps = 18/199 (9%)
Frame = +1
Query: 84 MLRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVN 143
++ C AEYLEMT++ GNL+ +S+ + ++ L + L E +L AE LV+
Sbjct: 511 LVHCAAEYLEMTDEYEEGNLLTKSESFFHKNILRNWKDCILALQSSEPILSRAENLNLVD 690
Query: 144 RCIDAIAYMASKQSQLCSPARSQCSSDGVMSSSM--------ASHRRPVVHWWGEDLTVL 195
+C++A++ MA L S S+ A R WW ED++ L
Sbjct: 691 KCLNALSMMACTDPSLFGWPMMMYGSFQSPGGSILWNGINTGARIRTSESEWWFEDISYL 870
Query: 196 RIDIFQRVLIAMMGRGFKQFDLGPIIMLYAQKSLEGLEIF--GKGRK-------ELEPQH 246
+ +F+ ++ M RG + L IM Y++K L GL + G+G K L P
Sbjct: 871 SVRLFESLIKTMRSRGMRPESLAGAIMYYSRKHLPGLGRWQGGQGGKTRTVASFSLTPAS 1050
Query: 247 EH-EKRVVVETLVSLLPRE 264
++ V++E++ LLP++
Sbjct: 1051ATVDQXVLLESIEKLLPQK 1107
>TC92992 similar to PIR|T02594|T02594 hypothetical protein At2g14820
[imported] - Arabidopsis thaliana, partial (13%)
Length = 783
Score = 63.2 bits (152), Expect = 3e-10
Identities = 31/69 (44%), Positives = 43/69 (61%)
Frame = +2
Query: 413 HPFLSEMEKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSFQQKHLRETMNDGGV 472
HP +S+ EKK++C +M C+KLS DA HA QN+RLP + ++QVL F+Q +
Sbjct: 5 HPGISKGEKKSICQLMDCRKLSVDASLHAVQNERLPTRVIVQVLHFEQVRTAAS------ 166
Query: 473 NWDGTSIPD 481
GTS PD
Sbjct: 167 --SGTSTPD 187
>TC82694 similar to GP|6224712|gb|AAF05914.1| non-phototropic hypocotyl 3
{Arabidopsis thaliana}, partial (25%)
Length = 671
Score = 58.2 bits (139), Expect = 9e-09
Identities = 30/69 (43%), Positives = 44/69 (63%)
Frame = +1
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNR 144
LRC AEYLEMTED GNL+ +++ +L+ V L + ++ VL E L AE ++V R
Sbjct: 403 LRCAAEYLEMTEDLEEGNLIFKTEAFLSYVVLSSWRDSIVVLKSSEKLSPWAENLQIVRR 582
Query: 145 CIDAIAYMA 153
C ++IA+ A
Sbjct: 583 CSESIAWKA 609
>AW690546 similar to GP|6634762|gb| Contains a bZIP transcription factor
PF|00170 domain. ESTs gb|R30400 gb|AA650964
gb|AI994521, partial (22%)
Length = 536
Score = 55.5 bits (132), Expect = 6e-08
Identities = 27/75 (36%), Positives = 45/75 (60%)
Frame = +1
Query: 85 LRCVAEYLEMTEDNSIGNLVGRSDFYLNEVALETISGAVSVLHMPETLLQVAEKAKLVNR 144
L C AE+LEMTE+ GNL+ +++ + N+V L + ++ L + +L AE+ +V R
Sbjct: 253 LWCAAEHLEMTEEYGEGNLISQAETFFNQVVLRSWKDSMRALQTCDDVLDQAEELHIVKR 432
Query: 145 CIDAIAYMASKQSQL 159
CI+++A AS L
Sbjct: 433 CIESLAAKASTDPNL 477
>AW586219
Length = 704
Score = 52.4 bits (124), Expect = 5e-07
Identities = 23/51 (45%), Positives = 38/51 (74%), Gaps = 1/51 (1%)
Frame = +1
Query: 417 SEMEKKNVCSVMHCQKLSRDARAHAAQNDRLPVQTVLQVLSF-QQKHLRET 466
S+ EKK +C ++ CQ+LS + R HA +N+ LP++TV+Q+L + Q+K +ET
Sbjct: 1 SKAEKKRLCGILECQRLSPEVRVHAVKNEFLPLRTVVQLLYYEQEKDSKET 153
>TC77907 similar to GP|15810641|gb|AAL07245.1 unknown protein {Arabidopsis
thaliana}, partial (29%)
Length = 721
Score = 46.6 bits (109), Expect = 3e-05
Identities = 26/76 (34%), Positives = 41/76 (53%), Gaps = 1/76 (1%)
Frame = +1
Query: 51 SSAMKKTSEWNFPQEIPSDVTVQIDNASFSLHKMLRCVAEYLEMTEDNSIGNLVGRSD-F 109
S AM++T +W F QE+P+DV V++ A F LHK ++ + + N I L+ SD
Sbjct: 163 SLAMERTGQWVFSQEVPTDVIVEVGEARFCLHK-------FMLVAKSNYIRKLIMESDET 321
Query: 110 YLNEVALETISGAVSV 125
+L + L I G +
Sbjct: 322 HLTRIDLSDIPGGSGI 369
>TC83384 similar to GP|15810641|gb|AAL07245.1 unknown protein {Arabidopsis
thaliana}, partial (13%)
Length = 658
Score = 46.2 bits (108), Expect = 4e-05
Identities = 31/92 (33%), Positives = 50/92 (53%), Gaps = 2/92 (2%)
Frame = +3
Query: 505 EELKREIVKLKMKLQEIEKPAHESSAPSSPLISAYSPSVNKPPLPRKSFMKSVSRKLGRL 564
EEL+ E++++KM + +++K H + SS V K + + +F SVS+KLG+L
Sbjct: 81 EELRTELIRMKMYVSDLQKNGHGGTTSSS--------EVGKE-VKKSTFFSSVSKKLGKL 233
Query: 565 YPFSRAAADT--VTTPFKDRLKPEKKIRHSIS 594
PF + DT + D KP ++ R SIS
Sbjct: 234 NPFKNGSKDTSHIEDVGVDLTKPRRR-RFSIS 326
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.133 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,401,587
Number of Sequences: 36976
Number of extensions: 211363
Number of successful extensions: 1391
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 1346
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1382
length of query: 594
length of database: 9,014,727
effective HSP length: 102
effective length of query: 492
effective length of database: 5,243,175
effective search space: 2579642100
effective search space used: 2579642100
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0157a.4


