

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0154.8
(247 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
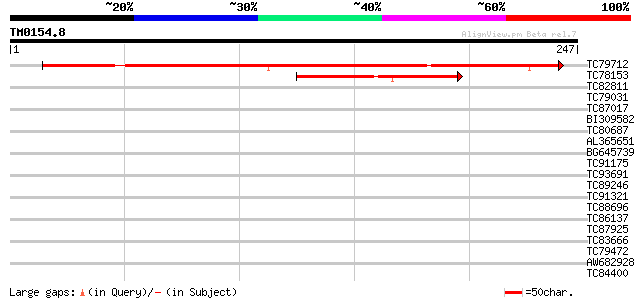
Sequences producing significant alignments: (bits) Value
TC79712 similar to GP|13374851|emb|CAC34485. putative protein {A... 325 8e-90
TC78153 homologue to GP|21617923|gb|AAM66973.1 unknown {Arabidop... 77 6e-15
TC82811 31 0.50
TC79031 weakly similar to GP|13492674|gb|AAK28303.1 phenylpropan... 30 0.66
TC87017 similar to PIR|A96564|A96564 unknown protein 23094-2177... 29 1.5
BI309582 similar to GP|9758127|dbj| SCARECROW gene regulator {Ar... 29 1.9
TC80687 similar to GP|3643082|gb|AAC36697.1| protein phosphatase... 28 2.5
AL365651 28 2.5
BG645739 similar to PIR|T45808|T4 helicase-like protein - Arabid... 28 4.3
TC91175 weakly similar to PIR|T05129|T05129 hypothetical protein... 28 4.3
TC93691 28 4.3
TC89246 homologue to PIR|T06594|T06594 heat shock protein dnaJ -... 28 4.3
TC91321 similar to GP|14486602|gb|AAK63206.1 C2H2 zinc-finger pr... 27 5.6
TC88696 weakly similar to PIR|T10620|T10620 probable calcium-bin... 27 5.6
TC86137 weakly similar to PIR|A96673|A96673 probable cytochrome ... 27 5.6
TC87925 similar to GP|13605495|gb|AAK32741.1 At1g65410/T8F5_19 {... 27 5.6
TC83666 similar to GP|15042832|gb|AAK82455.1 putative RNA bindin... 27 5.6
TC79472 similar to PIR|C96685|C96685 unknown protein F15E12.10 [... 27 5.6
AW682928 similar to GP|22507458|gb| Unknown (protein for MGC:303... 27 7.3
TC84400 similar to GP|2149820|gb|AAB58660.1| maturase {Nothofagu... 27 7.3
>TC79712 similar to GP|13374851|emb|CAC34485. putative protein {Arabidopsis
thaliana}, partial (39%)
Length = 926
Score = 325 bits (834), Expect = 8e-90
Identities = 164/229 (71%), Positives = 188/229 (81%), Gaps = 2/229 (0%)
Frame = +2
Query: 15 TETDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPELHKSFTTATDNCFRFLHSLA 74
T+T+ KK S CW N+Q+ A P LK PISN SPP+L S + A DNC + LHSLA
Sbjct: 65 TDTETKKNVSFCWGNSQSTLAFPFLKSPISNF----SPPQLPTSISNAVDNCSKLLHSLA 232
Query: 75 SQNPFLNKILSLRSEFDSLCFQIRCSNYRSRRWLSSH-NFAAVLPGDSVAGLVVGNGIQN 133
SQNPFLNK+LSL S+F C QIRCSNY + RW+S+H NFAAVLPGDSVAGLVV NG+ N
Sbjct: 233 SQNPFLNKLLSLSSQFHDTCVQIRCSNYSNSRWVSNHHNFAAVLPGDSVAGLVVANGLNN 412
Query: 134 FLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAFL 193
FL++YNTLLV RLVLTWFPN+PPAIV PLST+CDPYLN+FRGLIPPLGG LDLSPILAFL
Sbjct: 413 FLSLYNTLLVARLVLTWFPNAPPAIVAPLSTVCDPYLNVFRGLIPPLGG-LDLSPILAFL 589
Query: 194 VLNAFTSASAALPAELPVTQQSEQGVAATLQS-TDITSSQKKWMKRLQG 241
VLNAFTS +AALPAELPVT+QS+QG+ A LQ TD+TSSQ KWM+RLQG
Sbjct: 590 VLNAFTSTAAALPAELPVTEQSKQGLEARLQQPTDVTSSQNKWMRRLQG 736
>TC78153 homologue to GP|21617923|gb|AAM66973.1 unknown {Arabidopsis
thaliana}, partial (34%)
Length = 986
Score = 77.0 bits (188), Expect = 6e-15
Identities = 39/75 (52%), Positives = 56/75 (74%), Gaps = 3/75 (4%)
Frame = +3
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTI---CDPYLNLFRGLIPPLGG 182
VV G++ +L+IY+ +L+VR++L+WFPN P PLS I CDPYL+LFR ++PP+
Sbjct: 447 VVAAGLKKWLDIYSGVLLVRVLLSWFPNIPWERQ-PLSAIRDLCDPYLSLFRNIVPPVFD 623
Query: 183 TLDLSPILAFLVLNA 197
TLD+SP+LAF VL +
Sbjct: 624 TLDISPLLAFAVLGS 668
>TC82811
Length = 961
Score = 30.8 bits (68), Expect = 0.50
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = -3
Query: 55 LHKSFTTATDNCFRFLHSLASQNPFLNKILSLRSEFDSLCF 95
+H T NCF ++L + PFLN +L+L + +L F
Sbjct: 935 IHTYKITQIINCFNTFYNLTTYKPFLNTLLNLDDKHTNLFF 813
>TC79031 weakly similar to GP|13492674|gb|AAK28303.1
phenylpropanoid:glucosyltransferase 1 {Nicotiana
tabacum}, partial (32%)
Length = 1385
Score = 30.4 bits (67), Expect = 0.66
Identities = 20/57 (35%), Positives = 28/57 (49%), Gaps = 13/57 (22%)
Frame = -1
Query: 31 QTPFALPILKPPISNLTSFL-----SPPEL--------HKSFTTATDNCFRFLHSLA 74
+ P + L +++LT FL SPP + FTT+ N F+FLHSLA
Sbjct: 1151 EPPTLIAFLASTLNSLTLFLIATSSSPPSIINLIAFPISSLFTTSLPNSFQFLHSLA 981
>TC87017 similar to PIR|A96564|A96564 unknown protein 23094-21772
[imported] - Arabidopsis thaliana, partial (30%)
Length = 1701
Score = 29.3 bits (64), Expect = 1.5
Identities = 15/42 (35%), Positives = 24/42 (56%)
Frame = -3
Query: 168 PYLNLFRGLIPPLGGTLDLSPILAFLVLNAFTSASAALPAEL 209
P+L+LF +GG+L +SP L+ + +S S LP+ L
Sbjct: 673 PFLSLFSVFFSSVGGSLLISPALSLSATSFLSSFSILLPSLL 548
>BI309582 similar to GP|9758127|dbj| SCARECROW gene regulator {Arabidopsis
thaliana}, partial (9%)
Length = 692
Score = 28.9 bits (63), Expect = 1.9
Identities = 17/53 (32%), Positives = 22/53 (41%)
Frame = +1
Query: 17 TDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPELHKSFTTATDNCFRF 69
T+P VS H W N P + PP+S + + P S TD F F
Sbjct: 271 TNPLSVSLHPWHNNNIPVSSSTSLPPLSFPPTDFTDPFQVGSGPENTDPAFNF 429
>TC80687 similar to GP|3643082|gb|AAC36697.1| protein phosphatase-2C; PP2C
{Mesembryanthemum crystallinum}, partial (43%)
Length = 861
Score = 28.5 bits (62), Expect = 2.5
Identities = 16/48 (33%), Positives = 26/48 (53%), Gaps = 4/48 (8%)
Frame = -1
Query: 35 ALPILKPPISNLTSFL----SPPELHKSFTTATDNCFRFLHSLASQNP 78
+L +L P++N S L +PP + S ++A C RF +LA+ P
Sbjct: 657 SLSVLSNPLANTASHLFCHSAPPAISCSHSSAITLCIRFAQNLAT*AP 514
>AL365651
Length = 524
Score = 28.5 bits (62), Expect = 2.5
Identities = 17/36 (47%), Positives = 23/36 (63%), Gaps = 3/36 (8%)
Frame = -3
Query: 42 PISNLTSFLSPPEL--HKSFTTATD-NCFRFLHSLA 74
P + L+ SP + +KS T TD NCFRFLHS++
Sbjct: 249 PTTTLSQE*SPRSIVRYKS*TKHTDWNCFRFLHSIS 142
>BG645739 similar to PIR|T45808|T4 helicase-like protein - Arabidopsis
thaliana, partial (17%)
Length = 818
Score = 27.7 bits (60), Expect = 4.3
Identities = 20/70 (28%), Positives = 33/70 (46%)
Frame = -2
Query: 137 IYNTLLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAFLVLN 196
+YN + ++LVL S P L++ P + LF + + SP L+ ++LN
Sbjct: 604 LYN*MQSIKLVLI---QSTPMFCMVLNSF*KPLIELFMAIK*SWHNEVKKSP*LSHIILN 434
Query: 197 AFTSASAALP 206
TS A+P
Sbjct: 433 GCTSQQ*AIP 404
>TC91175 weakly similar to PIR|T05129|T05129 hypothetical protein F7H19.160
- Arabidopsis thaliana, partial (18%)
Length = 981
Score = 27.7 bits (60), Expect = 4.3
Identities = 18/44 (40%), Positives = 23/44 (51%)
Frame = +3
Query: 17 TDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPELHKSFT 60
T P+ SSHC R T + I P + T+ S P LH+SFT
Sbjct: 99 TKPRNSSSHC-RQTFASTSSSIF--PNTKFTNHESLPSLHESFT 221
>TC93691
Length = 482
Score = 27.7 bits (60), Expect = 4.3
Identities = 18/61 (29%), Positives = 28/61 (45%)
Frame = -2
Query: 84 LSLRSEFDSLCFQIRCSNYRSRRWLSSHNFAAVLPGDSVAGLVVGNGIQNFLNIYNTLLV 143
LS RS + CF N RRWLS +LP S+ + ++N +N+ +L
Sbjct: 211 LSSRSVLERFCFIRSTINSCKRRWLS-----ILLPPSSLRA*GLMKKVENIINMIRPILE 47
Query: 144 V 144
+
Sbjct: 46 I 44
>TC89246 homologue to PIR|T06594|T06594 heat shock protein dnaJ - garden
pea, partial (50%)
Length = 1119
Score = 27.7 bits (60), Expect = 4.3
Identities = 10/25 (40%), Positives = 18/25 (72%)
Frame = +2
Query: 132 QNFLNIYNTLLVVRLVLTWFPNSPP 156
QN +++ LL++ ++L+ FPN PP
Sbjct: 434 QNAFTLFSPLLLIIILLSVFPNPPP 508
>TC91321 similar to GP|14486602|gb|AAK63206.1 C2H2 zinc-finger protein
SERRATE {Arabidopsis thaliana}, partial (23%)
Length = 1069
Score = 27.3 bits (59), Expect = 5.6
Identities = 17/61 (27%), Positives = 24/61 (38%)
Frame = -1
Query: 43 ISNLTSFLSPPELHKSFTTATDNCFRFLHSLASQNPFLNKILSLRSEFDSLCFQIRCSNY 102
IS L PP ++ NC + L+ +PF+ +LS F F SN
Sbjct: 568 ISELNDTAGPPYASNIVSSRFPNCLSWDGELSGLSPFVLSLLSWSRRFPRSSFNRPSSNL 389
Query: 103 R 103
R
Sbjct: 388 R 386
>TC88696 weakly similar to PIR|T10620|T10620 probable calcium-binding
protein F21C20.130 - Arabidopsis thaliana, partial
(17%)
Length = 648
Score = 27.3 bits (59), Expect = 5.6
Identities = 16/43 (37%), Positives = 22/43 (50%)
Frame = +1
Query: 48 SFLSPPELHKSFTTATDNCFRFLHSLASQNPFLNKILSLRSEF 90
S LS +LH+ F NC F+ SL N L +I + S+F
Sbjct: 49 SILSSTDLHRIFEKLDTNCDGFV-SLEELNSVLQRICNTSSQF 174
>TC86137 weakly similar to PIR|A96673|A96673 probable cytochrome P450
F13O11.25 [imported] - Arabidopsis thaliana, partial
(69%)
Length = 1695
Score = 27.3 bits (59), Expect = 5.6
Identities = 28/96 (29%), Positives = 39/96 (40%), Gaps = 4/96 (4%)
Frame = -3
Query: 15 TETDPKKVSSHCWRNTQTPFALPILKPPISNLTSFLSPPELHKSFTTATDNCFRFLH--- 71
++T K S C ++ T P PPI++ S S P FTT F +H
Sbjct: 1168 SKTTHFKYRSFCKSSSFTSSFSPFSSPPITSRISSTSLPCTSGYFTT-----FAIIHCKL 1004
Query: 72 -SLASQNPFLNKILSLRSEFDSLCFQIRCSNYRSRR 106
+ S PF N L F SL F+ + S+R
Sbjct: 1003 VEVVSVPPFKNSEQRLTISF-SLSFRFSSGHSNSKR 899
>TC87925 similar to GP|13605495|gb|AAK32741.1 At1g65410/T8F5_19 {Arabidopsis
thaliana}, partial (82%)
Length = 1699
Score = 27.3 bits (59), Expect = 5.6
Identities = 12/24 (50%), Positives = 13/24 (54%)
Frame = +2
Query: 38 ILKPPISNLTSFLSPPELHKSFTT 61
IL PP S L L PP H+S T
Sbjct: 80 ILHPPFSTLLKILHPPSPHESHAT 151
>TC83666 similar to GP|15042832|gb|AAK82455.1 putative RNA binding protein
{Oryza sativa}, partial (52%)
Length = 896
Score = 27.3 bits (59), Expect = 5.6
Identities = 15/34 (44%), Positives = 19/34 (55%)
Frame = -2
Query: 53 PELHKSFTTATDNCFRFLHSLASQNPFLNKILSL 86
PELH + D CF LH+L QNP + + L L
Sbjct: 466 PELHSDYRQQ-D*CFFDLHTL*MQNPLICQYLCL 368
>TC79472 similar to PIR|C96685|C96685 unknown protein F15E12.10 [imported] -
Arabidopsis thaliana, partial (64%)
Length = 1001
Score = 27.3 bits (59), Expect = 5.6
Identities = 28/81 (34%), Positives = 36/81 (43%), Gaps = 4/81 (4%)
Frame = -1
Query: 33 PFALPILKPPISNLTSFLSPPELHKSFTTATDNCFRFLHSLASQNPFLNKILS----LRS 88
P ALP +PP+S+L P E K T+ FL S+ KI +R
Sbjct: 263 PIALPPNRPPLSSLDQEKKP*E--KRVFLITNLWTVFLADCGSRRGTPVKITGEMN*VRI 90
Query: 89 EFDSLCFQIRCSNYRSRRWLS 109
E DS+ + R S YRS W S
Sbjct: 89 EVDSI-EEKRNSGYRSEIWCS 30
>AW682928 similar to GP|22507458|gb| Unknown (protein for MGC:30371) {Mus
musculus}, partial (13%)
Length = 641
Score = 26.9 bits (58), Expect = 7.3
Identities = 21/68 (30%), Positives = 30/68 (43%), Gaps = 11/68 (16%)
Frame = +3
Query: 152 PNSPPAIVGPLSTIC-------DPYLNLFRGL----IPPLGGTLDLSPILAFLVLNAFTS 200
P PP+ P+ST+ P L L R L +PP +SP+ L+LNA
Sbjct: 153 PLLPPSATPPISTLLLLELLLESPLLILGRMLTPPPLPPSPLLQQMSPLPLHLILNALLM 332
Query: 201 ASAALPAE 208
+P+E
Sbjct: 333 VPNIVPSE 356
>TC84400 similar to GP|2149820|gb|AAB58660.1| maturase {Nothofagus
cunninghamii}, partial (12%)
Length = 671
Score = 26.9 bits (58), Expect = 7.3
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = -3
Query: 32 TPFALPILKPPISNLTSFLSP 52
TPF LP KP + + SF+SP
Sbjct: 573 TPFLLPHKKPIVGSCNSFISP 511
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,108,988
Number of Sequences: 36976
Number of extensions: 150421
Number of successful extensions: 927
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 915
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 922
length of query: 247
length of database: 9,014,727
effective HSP length: 94
effective length of query: 153
effective length of database: 5,538,983
effective search space: 847464399
effective search space used: 847464399
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0154.8


