

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
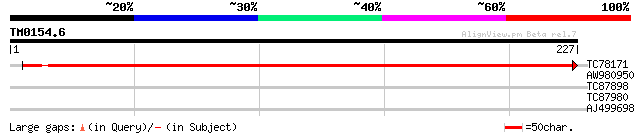
Query= TM0154.6
(227 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC78171 GP|6562649|emb|CAB62805.1 possible F-box protein FBL6 {L... 376 e-105
AW980950 28 2.9
TC87898 weakly similar to GP|1174199|gb|AAA86652.1| S25-PR6 {Nic... 27 4.9
TC87980 homologue to GP|4582789|emb|CAB40374.1 Starch synthase i... 27 8.4
AJ499698 27 8.4
>TC78171 GP|6562649|emb|CAB62805.1 possible F-box protein FBL6 {Leishmania
major}, partial (2%)
Length = 1110
Score = 376 bits (966), Expect = e-105
Identities = 182/222 (81%), Positives = 201/222 (89%)
Frame = +1
Query: 6 LVVILSFLHSASTGSAATNTQIKVTNNPADKLVAAINENRTAYKVSELYDNAGLACIALQ 65
L ILSFL G++ + TQIK+T+NPADKLVAAINENRTA+K S L+DN GLAC+ALQ
Sbjct: 208 LFFILSFL--LGLGASTSATQIKITDNPADKLVAAINENRTAHKDSSLFDNPGLACLALQ 381
Query: 66 YIKAYQGDCGAVGGPDAKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYVHAPEA 125
YIKAYQGDCGAVGG DAKKPPESQFAE FAPNCGVKAS+LA ITGRFLGCQTKYVHAPEA
Sbjct: 382 YIKAYQGDCGAVGGSDAKKPPESQFAEAFAPNCGVKASTLARITGRFLGCQTKYVHAPEA 561
Query: 126 FSEVLIQNQRSLEILHSKNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFEGGVAKL 185
FSE+LI+N++SL+IL+SKNHTQVGAAVTGTDGGSPYFWCVLFS GKPNSTFAFEGGVAK
Sbjct: 562 FSEILIRNEKSLDILYSKNHTQVGAAVTGTDGGSPYFWCVLFSNGKPNSTFAFEGGVAKD 741
Query: 186 TKPGCFSGANDECSGAHDWSPLSVMWLFAASVLIALGFAFPL 227
TKPGC+SGAND CSGAH WSP+S+MWLF ASV IA+GFAFPL
Sbjct: 742 TKPGCYSGANDVCSGAHVWSPVSLMWLFVASVSIAMGFAFPL 867
>AW980950
Length = 548
Score = 28.1 bits (61), Expect = 2.9
Identities = 14/44 (31%), Positives = 18/44 (40%), Gaps = 6/44 (13%)
Frame = +1
Query: 176 FAFEGGVAKLTKPG------CFSGANDECSGAHDWSPLSVMWLF 213
F+FEGG K + C G N C G PL +W +
Sbjct: 214 FSFEGGFGKCVRDADCVDEVCSPGCNKRCVGFECQCPL*CLWFY 345
>TC87898 weakly similar to GP|1174199|gb|AAA86652.1| S25-PR6 {Nicotiana
tabacum}, partial (25%)
Length = 1300
Score = 27.3 bits (59), Expect = 4.9
Identities = 20/55 (36%), Positives = 25/55 (45%), Gaps = 7/55 (12%)
Frame = +3
Query: 157 GGSPYFWCVLFSGGKPNSTFAF---EGGVAKLTKPGCFSGANDECSGA----HDW 204
GG +F+C+ + N F F GG+ L CF GAN C GA H W
Sbjct: 879 GGWGWFFCIF----RCNLYFVFLKYVGGIHNLIS*-CFGGANCNCLGAYVHQHQW 1028
>TC87980 homologue to GP|4582789|emb|CAB40374.1 Starch synthase isoform SS
III {Vigna unguiculata}, partial (30%)
Length = 1410
Score = 26.6 bits (57), Expect = 8.4
Identities = 10/22 (45%), Positives = 12/22 (54%)
Frame = +3
Query: 192 SGANDECSGAHDWSPLSVMWLF 213
+G N + HDWS V WLF
Sbjct: 48 NGFNPDIIHCHDWSSAPVAWLF 113
>AJ499698
Length = 258
Score = 26.6 bits (57), Expect = 8.4
Identities = 17/54 (31%), Positives = 25/54 (45%), Gaps = 2/54 (3%)
Frame = +3
Query: 111 RFLGCQTKYVHAPEAFSEVLIQNQRSLEILHS--KNHTQVGAAVTGTDGGSPYF 162
R+ +Y H P +F + IQN + +I HS KNHT + T +F
Sbjct: 27 RYSTVNQEYTH-P*SFPSLFIQNP*TFQIFHSKKKNHTHYLVVM*ATTSTQQHF 185
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,218,796
Number of Sequences: 36976
Number of extensions: 99057
Number of successful extensions: 402
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 402
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 402
length of query: 227
length of database: 9,014,727
effective HSP length: 93
effective length of query: 134
effective length of database: 5,575,959
effective search space: 747178506
effective search space used: 747178506
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0154.6


