

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
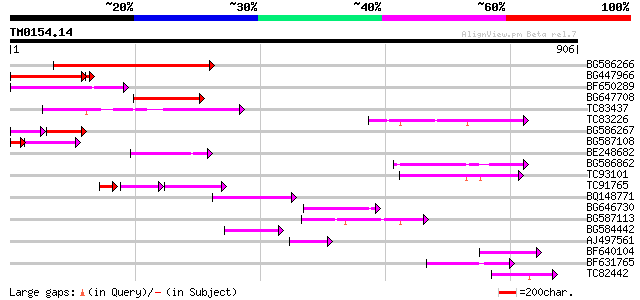
Query= TM0154.14
(906 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 220 2e-57
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 148 6e-36
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 134 2e-31
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 125 1e-28
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 106 3e-23
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 99 1e-20
BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At... 62 4e-18
BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - ... 75 3e-17
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 87 4e-17
BG586862 85 1e-16
TC93101 79 6e-15
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 69 6e-12
BQ148771 69 1e-11
BG646730 66 5e-11
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 66 7e-11
BG584442 57 3e-08
AJ497561 weakly similar to GP|10140689|gb putative non-LTR retro... 50 3e-06
BF640104 50 3e-06
BF631765 47 3e-05
TC82442 47 3e-05
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 220 bits (561), Expect = 2e-57
Identities = 111/256 (43%), Positives = 159/256 (61%)
Frame = -3
Query: 71 NQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNALVAFECFHYMK 130
+++R I+ CN +KII K L+ R++ +L +I SQ+AFVPGR I+DN L+ + HY++
Sbjct: 775 SEYRTIAPCNTQYKIIAKILSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKILHYLR 596
Query: 131 KKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVSTVDFSIMLNG 190
+ + + MA+K DM+KAYDR+ W FL VL ++GF W+S IM CVSTV +S ++NG
Sbjct: 595 QSGAKKHVSMAVKTDMTKAYDRIAWNFLREVLTRLGFHGIWISWIMECVSTVSYSFLING 416
Query: 191 APQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIARTAPVISHLLF 250
PQ P RGLRQGDPLSPYLFILC EV S L +A+ +L G+K+AR P I+HLLF
Sbjct: 415 GPQGRVLPSRGLRQGDPLSPYLFILCTEVLSGLCQQALRKGTLPGVKVARNCPPINHLLF 236
Query: 251 ADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDCFHELRQLLTV 310
ADD++ F ++ + +I+ Y ASG+ IN KS ++ S ++ L +
Sbjct: 235 ADDTMFFGKSNASSCAILLSIMDKYRAASGRCIN*TKSAITFSSKTSQAIIDRVKGELKI 56
Query: 311 KAVESYDKYLGLPTII 326
KYLG I+
Sbjct: 55 AKEGGTGKYLGYRNIL 8
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 148 bits (373), Expect(2) = 6e-36
Identities = 73/123 (59%), Positives = 91/123 (73%)
Frame = +2
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V EA+ QMHP KAPG DGLPALF+QK+WHI+G +V LQVL+ + +NKT +VL
Sbjct: 269 EVLEAINQMHPVKAPGPDGLPALFFQKYWHIVGKEVQQMVLQVLNNSMETEELNKTFIVL 448
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK K P +R ISLCNV+ KIITK +ANR+K LPD+I Q+AFV GRLITDNAL
Sbjct: 449 IPKGKNPNTPKDYRPISLCNVVMKIITKVIANRVKQTLPDVIDVEQSAFVQGRLITDNAL 628
Query: 121 VAF 123
+A+
Sbjct: 629 IAW 637
Score = 21.9 bits (45), Expect(2) = 6e-36
Identities = 7/14 (50%), Positives = 10/14 (71%)
Frame = +1
Query: 122 AFECFHYMKKKISG 135
+ ECFH++K K G
Sbjct: 631 SMECFHWLKXKKKG 672
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 134 bits (337), Expect = 2e-31
Identities = 70/190 (36%), Positives = 111/190 (57%)
Frame = +3
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V+ ALF M +KAPG DG F++ W+IIGD V L IIN T + L
Sbjct: 36 EVKNALFSMDSSKAPGIDGYNVHFFKCSWNIIGDSVIDAILDFFKTGFMPKIINCTYVTL 215
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
+PK FR I+ C+VI+KII+K L +R++ +L ++ E+Q+AFV GR+I DN +
Sbjct: 216 LPKEVNVTSVKNFRPIACCSVIYKIISKILTSRMQGVLNSVVSENQSAFVKGRVIFDNII 395
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
++ E +K G + +K+D+ KAYD EWPF+ ++ ++GFP +V+ +M ++
Sbjct: 396 LSHELVKSYSRK--GISPRCMVKIDLXKAYDSXEWPFIKHLMLELGFPYKFVNWVMAXLT 569
Query: 181 TVDFSIMLNG 190
T ++ NG
Sbjct: 570 TASYTFNXNG 599
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 125 bits (313), Expect = 1e-28
Identities = 60/114 (52%), Positives = 84/114 (73%)
Frame = +1
Query: 198 PHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIARTAPVISHLLFADDSVIF 257
P +GLRQGDPLSPYLFILC V S L+ + +LHGI++AR+ P I+HLLFADDS++F
Sbjct: 4 PEKGLRQGDPLSPYLFILCANVLSGLLKREGNKQNLHGIQVARSDPKITHLLFADDSLLF 183
Query: 258 ARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDCFHELRQLLTVK 311
ARA + EA +I +L SY+ ASGQ++N +KS++S S+NVP+ + Q + +K
Sbjct: 184 ARANLTEAATIMQVLHSYQSASGQLVNFEKSEVSYSQNVPNQEKEMICQQIAIK 345
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 106 bits (265), Expect = 3e-23
Identities = 88/337 (26%), Positives = 143/337 (42%), Gaps = 15/337 (4%)
Frame = +2
Query: 53 INKTLLVLIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPG 112
IN T + LIPKV P N FR ISL ++KI+ K LANRL++++ +I ++Q+AFV
Sbjct: 50 INSTFIALIPKVDNPQRLNDFRPISLVGSLYKILGKLLANRLRVVIGSVISDAQSAFVKN 229
Query: 113 RLITDNALV--------------AFECFHYMKKKISGCN-GVMALKLDMSKAYDRVEWPF 157
R I + + F C ++ K++ + G++ + +
Sbjct: 230 RQILEMVFL*QMRLWMRLRN*RKIFCCLRWILKRLITLSIGLIWILF*VG---------- 379
Query: 158 LASVLQQMGFPASWVSLIMTCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCG 217
M F W I CVST S+++NG+P + L Q + Y F +
Sbjct: 380 -------MSFLVLWRKWIKECVSTATTSVLVNGSPTNVLM--KSLVQTQLFTRYSFGVVN 532
Query: 218 EVFSALINKAVLASSLHGIKIARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQ 277
V V+SHL FA+D+++ +++A L +
Sbjct: 533 PV------------------------VVSHLQFANDTLLLETKNWANIRALRAALVIF*A 640
Query: 278 ASGQVINLDKSKLSCSRNVPDDCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFV 337
SG +N KS L C N+ E +L+ K + YLG+P + + +
Sbjct: 641 MSGLKVNFHKSGLVCV-NIAPSWLSEAASVLSWKVGKVPFLYLGMPIEGNSRRLSFWEPI 817
Query: 338 KERVWKKLKGWKERSLYRAGREVLVKAVAHAIPSYVM 374
R+ +L GW R L GR VL+K+V ++ Y +
Sbjct: 818 VNRIKARLTGWNSRFLSFGGRLVLLKSVLTSLSVYAL 928
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 98.6 bits (244), Expect = 1e-20
Identities = 85/287 (29%), Positives = 122/287 (41%), Gaps = 32/287 (11%)
Frame = +3
Query: 574 WPFSSDGHYSTKSGYQFLRHGAEAVVASSSQVPSLPRADWRKLWKADAL----------- 622
W + G YS KSGY LR + ++S S W+K+W +
Sbjct: 6 WMHNPTGIYSVKSGYNTLRTWQTQQINNTS-TSSDETLIWKKIWSLHTIPRHKVLLWRIL 182
Query: 623 ---LPVRAYLHARGLVTDPTCPRCLVALETVQHALLFCPMVQPIWFASSLGFRLAHECRV 679
LPVR+ L RG+ P CPRC ET+ H + CP+ + +WF S+L +
Sbjct: 183 NDSLPVRSSLRKRGIQCYPLCPRCHSKTETITHLFMSCPLSKRVWFGSNLCINFDNLPNP 362
Query: 680 HEFVSDFMQV---ADMSSLGAFLTLLYAIWMARNELCFKEKASTLEQILTHAAS------ 730
+ F++ + D ++Y +W ARN L E + LE + AS
Sbjct: 363 N-FIN*LYEAIL*KDECITI*IAAIIYNLWHARN-LSVLEDQTILEMDIIQRASNCISDY 536
Query: 731 ---LTPLPP---LEGPMVPQVHRP--NT-WSRPHPGTFKINFDAAMAPTGDAGFGLVARN 781
T PP G HRP NT W RP+ G K+N DA + G G G++ R+
Sbjct: 537 KQANTQAPPSMARTGYDPRSQHRPAKNTKWKRPNLGLVKVNTDANLQNHGKWGLGIIIRD 716
Query: 782 SRGEVLAAACSSHGPAASSLLAEGLSFRWALGLAMDLGFRRVVFEID 828
G V+AA+ +L AE + + A D GF +V FE D
Sbjct: 717 EVGLVMAASTWETDGNDRALEAEAYALLTGMRFAKDCGFXKVXFEGD 857
>BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (16%)
Length = 794
Score = 62.0 bits (149), Expect(2) = 4e-18
Identities = 31/63 (49%), Positives = 44/63 (69%)
Frame = +1
Query: 60 LIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNA 119
L+PK + +FR ISLCNV +KI++K L+ RLK +LP +I E+Q AF +LI+DN
Sbjct: 601 LVPKKLEAKRLVEFRPISLCNVAYKIVSKVLSKRLKSVLPWIITETQAAFGRRQLISDNI 780
Query: 120 LVA 122
L+A
Sbjct: 781 LIA 789
Score = 48.1 bits (113), Expect(2) = 4e-18
Identities = 22/56 (39%), Positives = 33/56 (58%)
Frame = +3
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKT 56
+V EA+F ++P K PG DG+ F+Q+FW +GDD++ + L INKT
Sbjct: 423 EVREAVFDINPHKCPGPDGMNVFFFQQFWDTMGDDLTSMAQEFLRTGKLEEGINKT 590
>BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - Arabidopsis
thaliana, partial (17%)
Length = 677
Score = 75.1 bits (183), Expect(2) = 3e-17
Identities = 41/89 (46%), Positives = 53/89 (59%)
Frame = +3
Query: 24 FYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVLIPKVKKPMHANQFRLISLCNVIF 83
F+Q WHII D+ L + +N T + LIPK K+P + R ISLCNV +
Sbjct: 411 FFQHSWHIIKMDLLKMVNSFLASGKLDTRLNTTNICLIPKKKRPTRMTELRPISLCNVGY 590
Query: 84 KIITKTLANRLKIILPDLICESQTAFVPG 112
KII+K L RLK+ LP LI E+Q+AFV G
Sbjct: 591 KIISKVLCQRLKVCLPSLISETQSAFVHG 677
Score = 32.3 bits (72), Expect(2) = 3e-17
Identities = 14/25 (56%), Positives = 17/25 (68%)
Frame = +2
Query: 1 DVEEALFQMHPTKAPGCDGLPALFY 25
+V ALF MHP KAPG DG+ L +
Sbjct: 341 EVRLALFIMHPEKAPGPDGMTTLLF 415
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 86.7 bits (213), Expect = 4e-17
Identities = 50/131 (38%), Positives = 68/131 (51%)
Frame = +3
Query: 193 QEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIARTAPVISHLLFAD 252
QE RGL+QGDPL+P+LF+L E S L+ AV + G + R +SHL +AD
Sbjct: 3 QEEISVQRGLKQGDPLAPFLFLLVAEGISGLMKNAVNRNLFQGFDVKRGGTRVSHLQYAD 182
Query: 253 DSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDCFHELRQLLTVKA 312
D++ TV ++KA+L +E ASG +N KS L NVP D + L +
Sbjct: 183 DTLCIGMPTVDNLWTLKALLQGFEMASGLKVNFHKSSL-IGINVPRDFMEAACRFLNCRE 359
Query: 313 VESYDKYLGLP 323
YLGLP
Sbjct: 360 ESIPFIYLGLP 392
>BG586862
Length = 804
Score = 85.1 bits (209), Expect = 1e-16
Identities = 66/218 (30%), Positives = 95/218 (43%), Gaps = 2/218 (0%)
Frame = -1
Query: 613 WRKLWKADALLPVRAYLHARGLVTDPTCPRCLVALETVQHALLFCPMVQPIWFASSLG-- 670
WR L A LPV+ LH RG+ CPRC +ETVQH L C + Q WF S LG
Sbjct: 612 WRLLHNA---LPVKDELHKRGIRCSLLCPRCESKIETVQHLFLNCEVTQKEWFGSQLGIN 442
Query: 671 FRLAHECRVHEFVSDFMQVADMSSLGAFLTLLYAIWMARNELCFKEKASTLEQILTHAAS 730
F + H+++++F+ D ++ A LLY+IW ARN+ F+ + ++ A+S
Sbjct: 441 FHSSGVLHFHDWITNFILKNDEETIIALTALLYSIWHARNQKVFENIDVPGDVVIQRASS 262
Query: 731 LTPLPPLEGPMVPQVHRPNTWSRPHPGTFKINFDAAMAPTGDAGFGLVARNSRGEVLAAA 790
L + V P+ A+ G G+VARN G +A+
Sbjct: 261 --SLHSFKMAQVSDSVLPSN---------------AIPSYSLWGIGVVARNCEGLAMASG 133
Query: 791 CSSHGPAASSLLAEGLSFRWALGLAMDLGFRRVVFEID 828
+ AE A+ A D GF + FE D
Sbjct: 132 TWLRHGIPCATTAEAWGIYQAMVFAGDCGFSKFEFESD 19
>TC93101
Length = 675
Score = 79.3 bits (194), Expect = 6e-15
Identities = 60/221 (27%), Positives = 91/221 (41%), Gaps = 22/221 (9%)
Frame = -1
Query: 623 LPVRAYLHARGLVTDPTCPRCLVALETVQHALLFCPMVQPIWFASSLGFRLAHECRVH-- 680
LPVR L+ RG+ P CPRC LET H + C Q +WF S L R ++
Sbjct: 663 LPVRXELNKRGVNCPPLCPRCYFNLETTNHIFMSCERTQRVWFGSQLSIRFPDNSTINFS 484
Query: 681 EFVSDFMQVADMSSLGAFLTLLYAIWMARNELCFKEKASTLEQILTHA---------ASL 731
+++ D + + + Y+IW ARN+ F+ + + + I+ A A++
Sbjct: 483 DWLFDAISNQTEEIIIKISAITYSIWHARNKAIFENQFVSEDTIIQ*AQNSILAYEQATI 304
Query: 732 TPL-PPLEGPMVPQVHRPNT----------WSRPHPGTFKINFDAAMAPTGDAGFGLVAR 780
P P + + NT W +P K N DA + G G G + R
Sbjct: 303 KPQNPNIVLSSLSATSNTNTTRRRSNVRSRWQKPLNNILKANCDANLQVQGRWGLGCIIR 124
Query: 781 NSRGEVLAAACSSHGPAASSLLAEGLSFRWALGLAMDLGFR 821
N+ GE A + AE + A+ LA D GF+
Sbjct: 123 NADGEAKVTATWCINGFDCAATAETYAILAAMYLAKDYGFK 1
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 68.6 bits (166), Expect = 1e-11
Identities = 40/99 (40%), Positives = 57/99 (57%)
Frame = +3
Query: 248 LLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDCFHELRQL 307
LLF + R A +K IL YE+ SG+ I+L KS + CSRNVPD + +
Sbjct: 279 LLFEGAPLRSLRVEEHHAQIMKNILILYEEDSGKAISLRKS*IYCSRNVPDILKTSITYI 458
Query: 308 LTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLK 346
L V+ + KYLGLP++IG+ +T F+ +K VW+K K
Sbjct: 459 LGVQFMLGTCKYLGLPSMIGRDRTTTFSSIKGGVWQKNK 575
Score = 61.6 bits (148), Expect(2) = 6e-12
Identities = 31/69 (44%), Positives = 41/69 (58%)
Frame = +2
Query: 178 CVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIK 237
CV + D+ +++N +P P RGL+QGD LSPY+FI+C E S LI A HG
Sbjct: 104 CVESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIICVEGLSFLIPHAKERGDTHGTS 283
Query: 238 IARTAPVIS 246
I R AP +S
Sbjct: 284 I*RGAPPVS 310
Score = 27.7 bits (60), Expect(2) = 6e-12
Identities = 11/29 (37%), Positives = 19/29 (64%)
Frame = +1
Query: 144 LDMSKAYDRVEWPFLASVLQQMGFPASWV 172
L +SK Y+RV+ +L ++ +MGF W+
Sbjct: 1 LYISKVYNRVD*DYLKEIMIKMGFNNRWI 87
>BQ148771
Length = 680
Score = 68.6 bits (166), Expect = 1e-11
Identities = 41/134 (30%), Positives = 59/134 (43%)
Frame = -3
Query: 325 IIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREVLVKAVAHAIPSYVMSCFLLPDGLC 384
++ KS + V +V L WK L A R L K+V A+P Y M ++P
Sbjct: 633 LVRKSTKKSRFSVYYQVHVMLANWKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACI 454
Query: 385 NQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNGRLGFRDFKMFNSALVAKNWWRIKS 444
+I+ + +F WG R H + W+ + K K LG R + N A + K W I S
Sbjct: 453 EEIQKLQRKFVWGDTEVSRRYHAVGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYS 274
Query: 445 NPESLMGKVYRAVY 458
SL +V R Y
Sbjct: 273 GSNSLCTEVMRGKY 232
>BG646730
Length = 799
Score = 66.2 bits (160), Expect = 5e-11
Identities = 35/125 (28%), Positives = 61/125 (48%), Gaps = 2/125 (1%)
Frame = +1
Query: 470 GYRPSYAWSSIMRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSL 529
G +PSYAW S+ ++ G W+I NG++V+ W D W+P + +
Sbjct: 388 GNQPSYAWRSMFNIKDVIDLGSRWSISNGQNVRIWKDDWLPNQTGFKVCSPMVDFEEATC 567
Query: 530 VSDLMLPGLLAWNRELINFICCPPTARAIMAIPLP--LVPHDDVLYWPFSSDGHYSTKSG 587
+S+L+ + R+L+ + A+ I+ IPL L P +LYW D +YS +S
Sbjct: 568 ISELIDTNTKSSKRDLVRNVFSAFEAKQILNIPLSWRLWPDRRILYW--ERDRNYSVRSA 741
Query: 588 YQFLR 592
+ ++
Sbjct: 742 HHLIK 756
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 65.9 bits (159), Expect = 7e-11
Identities = 60/230 (26%), Positives = 94/230 (40%), Gaps = 27/230 (11%)
Frame = -3
Query: 467 AKRGYRPSYAWSSIMRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFG 526
A G SYAW SI + + G IGNG++ W + + +H
Sbjct: 756 APLGSWASYAWRSIHSAQHLIKQGAKVIIGNGENTNIWEREMAWKLTCVTNHPN--KHSS 583
Query: 527 VSLVSDLML---------PGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFS 577
+ + + P N LIN I T R I++I +D W +S
Sbjct: 582 RAY*APTLYGYEGCRSDDPMRRERNANLINSIFPEGTRRKILSIHPQGPIGEDSYSWEYS 403
Query: 578 SDGHYSTKSGYQFLRHGAEAVVASSSQVPSL--PRAD--WRKLWKADAL----------- 622
GHYS KSGY + ++A+++Q ++ P D ++++WK +
Sbjct: 402 KSGHYSVKSGYYVQTN----IIAAANQRGTVDQPSLDDLYQRVWKYNTSPKVRHFLWRCI 235
Query: 623 ---LPVRAYLHARGLVTDPTCPRCLVALETVQHALLFCPMVQPIWFASSL 669
LP A + +R + D +C RC + ETV H L CP + IW S +
Sbjct: 234 SNSLPTAANMRSRHISKDGSCSRCGMESETVNHILFQCPYARLIWATSPI 85
>BG584442
Length = 775
Score = 57.0 bits (136), Expect = 3e-08
Identities = 34/96 (35%), Positives = 54/96 (55%), Gaps = 1/96 (1%)
Frame = +1
Query: 343 KKLKGWKERSLYRAGREVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYW-GGDVT 401
KK+ + + L + EV++K +I SYVMS FLL + ++IE +++ F W
Sbjct: 379 KKINF*RNKCLSKVM*EVMIKYALQSISSYVMSIFLLLNSQVDEIEKIMNTFSWVHVGEN 558
Query: 402 RRGMHCLSWKDLCKAKDNGRLGFRDFKMFNSALVAK 437
R+GMH +S + L K+ G +GF DF FN ++ K
Sbjct: 559 RKGMHWMS*EKLFVHKNYGGMGFTDFTTFNIPMLGK 666
>AJ497561 weakly similar to GP|10140689|gb putative non-LTR retroelement
reverse transcriptase {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 621
Score = 50.4 bits (119), Expect = 3e-06
Identities = 21/68 (30%), Positives = 34/68 (49%)
Frame = +2
Query: 448 SLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSIMRSGEIFATGGMWNIGNGKSVKAWSDK 507
SL+ K+ + YFP+ A G+ PS+ W S++ + + G W IG+G + S
Sbjct: 410 SLLSKILKFKYFPQWDFSYANLGHNPSFTWRSLLSTQSLLTLGHRWMIGDGSQINVSSMS 589
Query: 508 WVPGTDTL 515
W+ TL
Sbjct: 590 WIRNRPTL 613
>BF640104
Length = 344
Score = 50.4 bits (119), Expect = 3e-06
Identities = 34/101 (33%), Positives = 46/101 (44%), Gaps = 2/101 (1%)
Frame = -1
Query: 751 WSRPHPGTFKINFDAAMAPTGDAGFGLVARNSRGEVLAAACSSHGPAASSLLAEGLSFRW 810
W +PH G KIN DA + G G++ RN G V+A+ + + AE
Sbjct: 338 WIKPHQGVIKINCDANLTSEDVWGIGVITRNDNGIVMASGTWNRPGFMCPITAEAWGVYQ 159
Query: 811 ALGLAMDLGFRRVVFEIDCLLLYEAWKR--RGGCSILDSLI 849
A A+D GF+ V+FE D L R G S L S+I
Sbjct: 158 AALFALDQGFQNVLFENDNEKLISMLSREEEGHRSYLGSII 36
>BF631765
Length = 498
Score = 47.4 bits (111), Expect = 3e-05
Identities = 42/144 (29%), Positives = 62/144 (42%), Gaps = 4/144 (2%)
Frame = +1
Query: 667 SSLGFRLAHECRVHEFVSDFMQVADMSSLGAFLTLLYAIWMARNELCFK--EKASTLEQI 724
S LG + + + ++ + V + F T L+ IW RN+ F + L
Sbjct: 25 SPLGIHVPVQLDLLSWMEKCLAVPEQRVKQLFGTSLWLIWKHRNQKVFNNVDFDPPLVSA 204
Query: 725 LTHAASLTPLPPLEGPM--VPQVHRPNTWSRPHPGTFKINFDAAMAPTGDAGFGLVARNS 782
L S+ LP G + VP+ P G+ K+N A G G+G +ARN+
Sbjct: 205 LVEEFSIANLPLKRGQVCVVPR*QPPPF------GSIKVNVGAGCFGDGSTGWGFIARNA 366
Query: 783 RGEVLAAACSSHGPAASSLLAEGL 806
G L+A S G + S LLAE L
Sbjct: 367 AGLALSAETKSDGISYSPLLAECL 438
>TC82442
Length = 578
Score = 47.0 bits (110), Expect = 3e-05
Identities = 32/109 (29%), Positives = 51/109 (46%), Gaps = 4/109 (3%)
Frame = -3
Query: 771 GDAGFGLVARNSRGEVLAAACSSHGPAASSLLAEGLSFRWALGLAMDLGFRRVVFEIDC- 829
G G G + NS G+ + +A S + AE + A+ LA+ + ++FE DC
Sbjct: 330 GRWGLGCIIHNSAGQAMTSATWSRNGFNCAATAEAYAILAAMHLALTSDLKHMIFESDCE 151
Query: 830 ---LLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFDFSFIRRTGNSVA 875
L ++ S L S++ + ++S FD F FI R+GN VA
Sbjct: 150 AVIRKLNRGIEKDIDISYLGSVLEEIWKISEKFDMCSFRFIPRSGNRVA 4
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.139 0.444
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,392,419
Number of Sequences: 36976
Number of extensions: 535504
Number of successful extensions: 3213
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 2438
Number of HSP's successfully gapped in prelim test: 88
Number of HSP's that attempted gapping in prelim test: 725
Number of HSP's gapped (non-prelim): 2592
length of query: 906
length of database: 9,014,727
effective HSP length: 105
effective length of query: 801
effective length of database: 5,132,247
effective search space: 4110929847
effective search space used: 4110929847
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0154.14


