

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0153b.2
(402 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
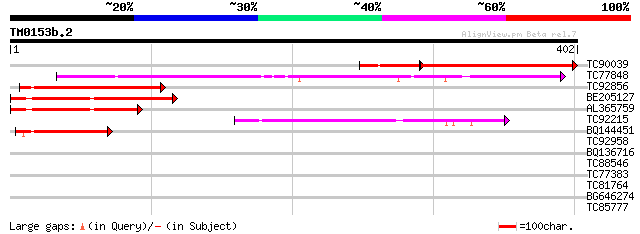
Score E
Sequences producing significant alignments: (bits) Value
TC90039 similar to GP|17381270|gb|AAL36053.1 AT3g24470/MXP5_4 {A... 216 3e-65
TC77848 similar to PIR|F86296|F86296 hypothetical protein AAF185... 195 3e-50
TC92856 similar to GP|16604677|gb|AAL24131.1 unknown protein {Ar... 147 7e-36
BE205127 similar to GP|16604677|gb| unknown protein {Arabidopsis... 136 2e-32
AL365759 similar to GP|17381270|gb| AT3g24470/MXP5_4 {Arabidopsi... 106 1e-23
TC92215 similar to GP|6862921|gb|AAF30310.1| hypothetical protei... 99 4e-21
BQ144451 weakly similar to GP|16604677|gb| unknown protein {Arab... 79 3e-15
TC92958 30 1.3
BQ136716 similar to PIR|S51590|S515 mitochondrial processing pep... 29 2.8
TC88546 similar to GP|18150126|dbj|BAB83633. spermine synthase {... 29 3.7
TC77383 similar to SP|O48922|C982_SOYBN Cytochrome P450 98A2 (EC... 28 4.8
TC81764 similar to GP|17104805|gb|AAL34291.1 putative glucan end... 28 6.3
BG646274 similar to GP|14517434|gb| AT4g01940/T7B11_20 {Arabidop... 28 8.2
TC85777 similar to PIR|F86442|F86442 unknown protein [imported] ... 28 8.2
>TC90039 similar to GP|17381270|gb|AAL36053.1 AT3g24470/MXP5_4 {Arabidopsis
thaliana}, partial (24%)
Length = 809
Score = 216 bits (549), Expect(2) = 3e-65
Identities = 97/111 (87%), Positives = 104/111 (93%)
Frame = +3
Query: 292 LVVGILALVIATFSTGIDSKCFQYRKGDKPAEEDDVPYGYGFFHFVFATGAMYFAMLLVG 351
L GILA+V+ TFSTGIDSKCFQ RKGDKPAEEDDVPYGYGFFHFVFATGAMYFAMLL+G
Sbjct: 132 LCYGILAIVMQTFSTGIDSKCFQLRKGDKPAEEDDVPYGYGFFHFVFATGAMYFAMLLIG 311
Query: 352 WNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAPIIWKNIHTEST 402
WN+HHSM+KW++DVGWTSAWVRIVNEWLAVCVYLWMLIAPIIWK TEST
Sbjct: 312 WNTHHSMRKWSLDVGWTSAWVRIVNEWLAVCVYLWMLIAPIIWKARQTEST 464
Score = 50.8 bits (120), Expect(2) = 3e-65
Identities = 28/46 (60%), Positives = 31/46 (66%)
Frame = +1
Query: 249 SPGLMGLYVVFLCWCAIRSEPEGYECIRKSDSPNKTDWQNIISLVV 294
SPGLMGLYV L SEPEG +CIR S + K DWQNIIS V+
Sbjct: 7 SPGLMGLYVSSLV-VGN*SEPEGDQCIRTSGTVTKPDWQNIISFVM 141
>TC77848 similar to PIR|F86296|F86296 hypothetical protein AAF18512.1
[imported] - Arabidopsis thaliana, complete
Length = 1592
Score = 195 bits (495), Expect = 3e-50
Identities = 121/389 (31%), Positives = 190/389 (48%), Gaps = 28/389 (7%)
Frame = +2
Query: 34 KMARYVYALIFLVCNLLAWASRD-ELPGRSVLTKLKGLKTCKDPKDCLGTNGVLRVSMGC 92
+ AR Y +F + ++AW R+ P + + K ++ T+ VLRVS G
Sbjct: 224 RSARIAYCGLFALSLVVAWMLREVAAPLMESIPWINHFKQTPS-REWFETDAVLRVSFGN 400
Query: 93 FLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPSELIDLYGEVAHF 152
FLFF ++ + RD H G W +KI+ W + IF F LP+E+I Y ++ F
Sbjct: 401 FLFFTILAAMMVGVKTQKDPRDGLHHGGWMMKIICWCLLVIFMFFLPNEIISFYETISKF 580
Query: 153 GAGVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLFATASYFICMVGV-----ILM 207
G+G+FL +Q++ ++ F+ ND + Y E+ ++ LF + +C V +L
Sbjct: 581 GSGMFLLVQVVLLLDFVHRWNDTWVG--YDEQFW-YIALFVVS--LVCYVATFVFSGVLF 745
Query: 208 YIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKVNAGILSPGLMGLYVVFLCWCAIRS 267
+ + C NI FI+ TL+L + V+LHP VN +L ++ Y ++LC+ A+ S
Sbjct: 746 HFFTPSGQDCGTNIFFISMTLMLAFVFAIVALHPAVNGSVLPASVISFYCMYLCYSALAS 925
Query: 268 EPEGYEC--IRKSDSPNKTDWQNIISLVVGILALVIATFSTG------------------ 307
EP YEC + K T + LV +L++V + G
Sbjct: 926 EPRDYECNGLHKHSKAVSTG-SLTLGLVTTVLSVVYSAVRAGSSATVLSPPSSPRAGKPL 1102
Query: 308 --IDSKCFQYRKGDKPAEEDDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDV 365
+D+K + + KP V Y Y FFH +F+ +MY AMLL GW++ +DV
Sbjct: 1103LPLDAKDEESNEKAKP-----VTYSYAFFHLIFSLASMYSAMLLTGWSTSVGESGKLVDV 1267
Query: 366 GWTSAWVRIVNEWLAVCVYLWMLIAPIIW 394
GW S WVRIV W +YLW L+API++
Sbjct: 1268GWPSVWVRIVTCWATALLYLWSLVAPIMF 1354
>TC92856 similar to GP|16604677|gb|AAL24131.1 unknown protein {Arabidopsis
thaliana}, partial (10%)
Length = 821
Score = 147 bits (371), Expect = 7e-36
Identities = 74/103 (71%), Positives = 80/103 (76%)
Frame = +1
Query: 8 SSNQQGCGVLLKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLTKL 67
+SNQQGCG LK SSW QF+NASNP MARYVY LIFL N+LAWA+RDEL S LT+L
Sbjct: 517 NSNQQGCG--LKDSSWCCQFKNASNPLMARYVYGLIFLAANMLAWATRDELSSISALTEL 690
Query: 68 KGLKTCKDPKDCLGTNGVLRVSMGCFLFFMMMFWSTARASKLN 110
KG K CK KDCLG +GVLRVSMGCFLFFMMM ST R S LN
Sbjct: 691 KGFKACKVGKDCLGAHGVLRVSMGCFLFFMMMXGSTTRTSNLN 819
>BE205127 similar to GP|16604677|gb| unknown protein {Arabidopsis thaliana},
partial (24%)
Length = 559
Score = 136 bits (342), Expect = 2e-32
Identities = 70/119 (58%), Positives = 86/119 (71%)
Frame = +3
Query: 1 MVTVEDNSSNQQGCGVLLKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPG 60
M T E N+ N++ C ++ SSW SQFRNASNP MARY YA IFL+ NLLAWA+RD G
Sbjct: 213 METGESNNGNER-C-IISNDSSWFSQFRNASNPWMARYAYAFIFLLANLLAWAARDY--G 380
Query: 61 RSVLTKLKGLKTCKDPKDCLGTNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSG 119
RS LT+++ LK C KDCLG VLRVS+GCF F+++MF ST SKL + R+TWHSG
Sbjct: 381 RSALTEMERLKGCNGGKDCLGAERVLRVSLGCFTFYIIMFLSTTGTSKLKQERNTWHSG 557
>AL365759 similar to GP|17381270|gb| AT3g24470/MXP5_4 {Arabidopsis thaliana},
partial (17%)
Length = 497
Score = 106 bits (265), Expect = 1e-23
Identities = 56/94 (59%), Positives = 68/94 (71%)
Frame = -1
Query: 1 MVTVEDNSSNQQGCGVLLKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPG 60
M T E N+ N++ C ++ SSW SQFRNASNP MARY YA IFL+ N LAWA+RD G
Sbjct: 284 METGESNNGNER-C-IISNDSSWFSQFRNASNPWMARYAYAFIFLLANSLAWAARDY--G 117
Query: 61 RSVLTKLKGLKTCKDPKDCLGTNGVLRVSMGCFL 94
RS LT+++ LK C KDCLG GVLRVS+GCF+
Sbjct: 116 RSALTEMERLKGCNGGKDCLGAEGVLRVSLGCFV 15
>TC92215 similar to GP|6862921|gb|AAF30310.1| hypothetical protein
{Arabidopsis thaliana}, partial (63%)
Length = 666
Score = 98.6 bits (244), Expect = 4e-21
Identities = 64/219 (29%), Positives = 108/219 (49%), Gaps = 24/219 (10%)
Frame = +1
Query: 160 IQLISIISFITWLNDFFASEKYAERCQIHVMLFATASYFICMVGVILMYIWYAPQP-SCL 218
IQ+I ++ ND + EK ++ I +++ + Y +++IW+ P C
Sbjct: 4 IQVIILLDCTHNWNDSWV-EKDEQKWYIALLVVSIGCYIAAFTLSGILFIWFNPGGYDCG 180
Query: 219 LNISFITFTLVLLQIMTSVSLHPKVNAGILSPGLMGLYVVFLCWCAIRSEPEGYECIRKS 278
LN+ F++ +++L + V+LHPKVN +L ++ LY ++C+ + SEP GYEC
Sbjct: 181 LNVFFLSMSMILAFVFGVVALHPKVNGSLLPASVISLYCAYVCYTGLSSEPRGYEC---- 348
Query: 279 DSPNKTDWQNIISLVVGILALVIATFSTGI---DSKCF-----------------QYRKG 318
+ NK+ + +LV+G+L V++ + + S F + +G
Sbjct: 349 NGLNKSRAVSTGTLVLGMLTTVLSVLYSALRAGSSTTFLSPPSSPKAGESKPLLEEVEEG 528
Query: 319 DKPAEEDD---VPYGYGFFHFVFATGAMYFAMLLVGWNS 354
EE + V Y Y FFH +FA +MY AMLL GW S
Sbjct: 529 KSKKEEKEARPVSYSYSFFHLIFALASMYSAMLLSGWTS 645
>BQ144451 weakly similar to GP|16604677|gb| unknown protein {Arabidopsis
thaliana}, partial (9%)
Length = 758
Score = 79.0 bits (193), Expect = 3e-15
Identities = 43/71 (60%), Positives = 47/71 (65%), Gaps = 2/71 (2%)
Frame = +1
Query: 5 EDNS--SNQQGCGVLLKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRS 62
E NS SNQQGCG LK SSW QF+NASNP MA YVY LIFL N LAWA+R+EL
Sbjct: 298 ESNSINSNQQGCG--LKDSSWCCQFKNASNPLMATYVYGLIFLTANTLAWATREELSSLV 471
Query: 63 VLTKLKGLKTC 73
L +L K C
Sbjct: 472 ALAELYCFKAC 504
>TC92958
Length = 652
Score = 30.4 bits (67), Expect = 1.3
Identities = 11/31 (35%), Positives = 16/31 (51%)
Frame = +2
Query: 326 DVPYGYGFFHFVFATGAMYFAMLLVGWNSHH 356
D+ Y +G FH F G+ + L + W S H
Sbjct: 302 DLLYPFGSFHLCFLIGSFHLCFLTISWGS*H 394
>BQ136716 similar to PIR|S51590|S515 mitochondrial processing peptidase (EC
3.4.24.64) alpha-II chain precursor - potato, partial
(3%)
Length = 958
Score = 29.3 bits (64), Expect = 2.8
Identities = 21/69 (30%), Positives = 34/69 (48%), Gaps = 2/69 (2%)
Frame = -2
Query: 246 GILSPGLMGLYVVFL-CWCAIRSEPEGYECIRKSDSP-NKTDWQNIISLVVGILALVIAT 303
G+L +M Y++ + C CAI E EC+ KS + W+ +S+ + A+V+
Sbjct: 915 GVLDQWIMSEYILCISC*CAIIGSCETEECVEKSHVR*FLSFWE--VSVYALLYAVVV*D 742
Query: 304 FSTGIDSKC 312
GID C
Sbjct: 741 IYVGIDGLC 715
>TC88546 similar to GP|18150126|dbj|BAB83633. spermine synthase {Arabidopsis
thaliana}, partial (66%)
Length = 1823
Score = 28.9 bits (63), Expect = 3.7
Identities = 20/83 (24%), Positives = 32/83 (38%)
Frame = +1
Query: 310 SKCFQYRKGDKPAEEDDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTS 369
+KCFQ R FFH+++ A++F W +KK + V +
Sbjct: 1618 NKCFQSRY---------------FFHYMYFLSALHF---FTPWLVLSPLKKRLVSVASMN 1743
Query: 370 AWVRIVNEWLAVCVYLWMLIAPI 392
N+W C +W+ PI
Sbjct: 1744 ------NQWFLSCASIWICSLPI 1794
>TC77383 similar to SP|O48922|C982_SOYBN Cytochrome P450 98A2 (EC 1.14.-.-).
[Soybean] {Glycine max}, complete
Length = 1782
Score = 28.5 bits (62), Expect = 4.8
Identities = 21/79 (26%), Positives = 35/79 (43%)
Frame = +2
Query: 177 ASEKYAERCQIHVMLFATASYFICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTS 236
AS + C ++F + V +++W P CL N ++ T +L++MT+
Sbjct: 1550 ASVGVVQTCDS*YLIFFSYCNVAVFQNVESIFLWDLFLPLCLCNYNWNTM*GTMLRLMTN 1729
Query: 237 VSLHPKVNAGILSPGLMGL 255
V L N +L P GL
Sbjct: 1730 VRL--VTNLTLLHPSDFGL 1780
>TC81764 similar to GP|17104805|gb|AAL34291.1 putative glucan endo-1
3-beta-glucosidase precursor {Arabidopsis thaliana},
partial (71%)
Length = 1560
Score = 28.1 bits (61), Expect = 6.3
Identities = 24/54 (44%), Positives = 28/54 (51%), Gaps = 4/54 (7%)
Frame = +3
Query: 206 LMYIWYAPQPSCLLNISF---ITFT-LVLLQIMTSVSLHPKVNAGILSPGLMGL 255
L+ + + PQ LL SF I FT L LLQ +TSVS N ILS L L
Sbjct: 186 LLAVRFFPQSQMLLPFSFLP*ILFTKLWLLQTLTSVSRFQLPNLWILSLSLFHL 347
>BG646274 similar to GP|14517434|gb| AT4g01940/T7B11_20 {Arabidopsis
thaliana}, partial (35%)
Length = 634
Score = 27.7 bits (60), Expect = 8.2
Identities = 12/16 (75%), Positives = 14/16 (87%)
Frame = -1
Query: 66 KLKGLKTCKDPKDCLG 81
KLKGL CK+P+DCLG
Sbjct: 625 KLKGL-ICKEPEDCLG 581
>TC85777 similar to PIR|F86442|F86442 unknown protein [imported] -
Arabidopsis thaliana, partial (89%)
Length = 2381
Score = 27.7 bits (60), Expect = 8.2
Identities = 12/16 (75%), Positives = 14/16 (87%)
Frame = -1
Query: 66 KLKGLKTCKDPKDCLG 81
KLKGL CK+P+DCLG
Sbjct: 359 KLKGL-ICKEPEDCLG 315
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.328 0.140 0.466
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,140,747
Number of Sequences: 36976
Number of extensions: 295068
Number of successful extensions: 2075
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 2041
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2067
length of query: 402
length of database: 9,014,727
effective HSP length: 98
effective length of query: 304
effective length of database: 5,391,079
effective search space: 1638888016
effective search space used: 1638888016
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0153b.2


