

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0148.13
(480 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
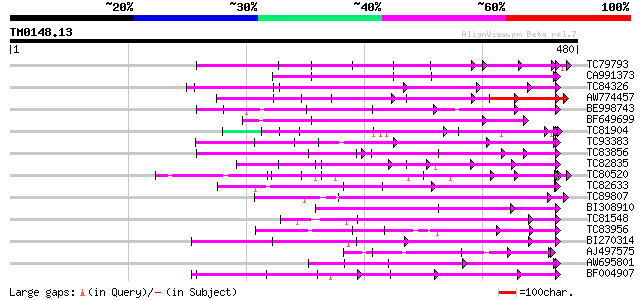
Score E
Sequences producing significant alignments: (bits) Value
TC79793 weakly similar to GP|6630464|gb|AAF19552.1| F23N19.4 {Ar... 132 2e-31
CA991373 weakly similar to PIR|D86260|D86 protein T12C24.22 [imp... 121 7e-28
TC84326 weakly similar to PIR|F85225|F85225 hypothetical protein... 113 2e-25
AW774457 weakly similar to GP|10176973|dbj gb|AAF19552.1~gene_id... 112 3e-25
BE998743 similar to GP|22128591|gb| fertility restorer-like prot... 108 6e-24
BF649699 similar to GP|9757911|dbj gb|AAF19552.1~gene_id:MMN10.1... 103 1e-22
TC81904 weakly similar to GP|8778410|gb|AAF79418.1| F16A14.3 {Ar... 98 6e-21
TC93383 similar to PIR|T47786|T47786 hypothetical protein F17J16... 95 5e-20
TC83856 weakly similar to PIR|B96656|B96656 unknown protein 419... 94 2e-19
TC82835 weakly similar to GP|21741712|emb|CAD41335. oj991113_30.... 93 3e-19
TC80520 weakly similar to PIR|T02562|T02562 probable salt-induci... 91 7e-19
TC82633 similar to GP|15450347|gb|AAK96467.1 At1g20300/F14O10_8 ... 87 2e-17
TC89807 weakly similar to PIR|B96600|B96600 protein F14J16.14 [i... 86 4e-17
BI308910 similar to GP|6682243|gb| hypothetical protein {Arabido... 83 3e-16
TC81548 weakly similar to GP|1931651|gb|AAB65486.1| membrane-ass... 82 3e-16
TC83956 weakly similar to GP|15982931|gb|AAL09812.1 AT3g53700/F4... 82 6e-16
BI270314 similar to GP|9759030|dbj contains similarity to salt-i... 80 1e-15
AJ497575 similar to PIR|T05642|T05 hypothetical protein F20D10.2... 79 4e-15
AW695801 weakly similar to PIR|T01622|T01 probable salt-inducibl... 79 4e-15
BF004907 weakly similar to GP|8953393|emb putative protein {Arab... 78 7e-15
>TC79793 weakly similar to GP|6630464|gb|AAF19552.1| F23N19.4 {Arabidopsis
thaliana}, partial (7%)
Length = 1580
Score = 132 bits (333), Expect = 2e-31
Identities = 68/239 (28%), Positives = 128/239 (53%)
Frame = +1
Query: 228 SYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEM 287
+YN+++ ++ + + ++ M G + DI +Y+I++ ++ K D+ LFKEM
Sbjct: 562 TYNSLMDGYCLVKEVNKAKSIFNTMAQGGVNPDIQSYSIMINGFCKIKKFDEAMNLFKEM 741
Query: 288 GRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNL 347
R PD TY+ L+ L K + A+ L+ QM + G+ P + Y +++D L + +
Sbjct: 742 HRKNIIPDVVTYSSLIDGLSKSGRISYALQLVDQMHDRGVPPNICTYNSILDALCKTHQV 921
Query: 348 HACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSM 407
+ G PD+ Y+++I G +G+LE A+++F ++ K NV TY M
Sbjct: 922 DKAIALLTKFKDKGFQPDISTYSILIKGLCQSGKLEDARKVFEGLLVKGHNLNVDTYTIM 1101
Query: 408 IRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
I+G C+ G F+EA ++L +ME GC P++ Y ++ L + A +++R+M+ +G
Sbjct: 1102IQGFCVEGLFNEALALLSKMEDNGCIPDAKTYEIIILSLFKKDENDMAEKLLREMIARG 1278
Score = 85.5 bits (210), Expect = 4e-17
Identities = 44/149 (29%), Positives = 84/149 (55%)
Frame = +2
Query: 256 GFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAA 315
G+ D +T+ + G++ Q + ++ GF D +Y L+H L K + AA
Sbjct: 14 GYVPDTITFTTLSKGLCLKGQIQQAFLFHDKVVALGFHFDQISYGTLIHGLCKVGETRAA 193
Query: 316 VNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITG 375
++LL+++ ++P V+ Y T+ID + + ++ F EM+ G PDVV Y+ +I+G
Sbjct: 194 LDLLQRVDGNLVQPNVVMYNTIIDSMCKVKLVNEAFDLFSEMVSKGISPDVVTYSALISG 373
Query: 376 YVVAGELEKAQEMFHEMISKEQVPNVYTY 404
+ + G+L+ A ++F++MI + P+VYT+
Sbjct: 374 FCILGKLKDAIDLFNKMILENIKPDVYTF 460
Score = 71.6 bits (174), Expect = 6e-13
Identities = 41/138 (29%), Positives = 68/138 (48%)
Frame = +2
Query: 326 GIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKA 385
G P + +TTL GL G + F D+++ G D ++Y +I G GE A
Sbjct: 14 GYVPDTITFTTLSKGLCLKGQIQQAFLFHDKVVALGFHFDQISYGTLIHGLCKVGETRAA 193
Query: 386 QEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSC 445
++ + PNV YN++I +C +EA + EM +KG SP+ V Y +L+S
Sbjct: 194 LDLLQRVDGNLVQPNVVMYNTIIDSMCKVKLVNEAFDLFSEMVSKGISPDVVTYSALISG 373
Query: 446 LQNAGKVANAREVMRQMM 463
GK+ +A ++ +M+
Sbjct: 374 FCILGKLKDAIDLFNKMI 427
Score = 71.2 bits (173), Expect = 8e-13
Identities = 53/237 (22%), Positives = 102/237 (42%)
Frame = +1
Query: 159 YHLIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNILIRTCGEAGLAKDLVETFIKSK 218
Y+ +M+ Y +E + M + G ++++I+I + + + F +
Sbjct: 565 YNSLMDGYCLVKEVNKAKSIFNTMAQGGVNPDIQSYSIMINGFCKIKKFDEAMNLFKEMH 744
Query: 219 TFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLD 278
N P +Y++++ L + + QM G +I TYN ++ A + ++D
Sbjct: 745 RKNIIPDVVTYSSLIDGLSKSGRISYALQLVDQMHDRGVPPNICTYNSILDALCKTHQVD 924
Query: 279 QFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLI 338
+ L + GF PD TY++L+ L + K A + + + G V YT +I
Sbjct: 925 KAIALLTKFKDKGFQPDISTYSILIKGLCQSGKLEDARKVFEGLLVKGHNLNVDTYTIMI 1104
Query: 339 DGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISK 395
G G + +M NGC+PD Y ++I E + A+++ EMI++
Sbjct: 1105QGFCVEGLFNEALALLSKMEDNGCIPDAKTYEIIILSLFKKDENDMAEKLLREMIAR 1275
Score = 63.9 bits (154), Expect = 1e-10
Identities = 40/145 (27%), Positives = 66/145 (44%)
Frame = +2
Query: 291 GFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHAC 350
G+ PD T+ L L + + A ++ G + Y TLI GL + G A
Sbjct: 14 GYVPDTITFTTLSKGLCLKGQIQQAFLFHDKVVALGFHFDQISYGTLIHGLCKVGETRAA 193
Query: 351 KHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRG 410
+ N P+VV Y +I + +A ++F EM+SK P+V TY+++I G
Sbjct: 194 LDLLQRVDGNLVQPNVVMYNTIIDSMCKVKLVNEAFDLFSEMVSKGISPDVVTYSALISG 373
Query: 411 LCMAGKFDEACSMLKEMETKGCSPN 435
C+ GK +A + +M + P+
Sbjct: 374 FCILGKLKDAIDLFNKMILENIKPD 448
Score = 52.0 bits (123), Expect = 5e-07
Identities = 29/125 (23%), Positives = 64/125 (51%), Gaps = 6/125 (4%)
Frame = +2
Query: 357 MIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGK 416
++K G +PD + +T + G + G++++A ++++ + +Y ++I GLC G+
Sbjct: 2 ILKMGYVPDTITFTTLSKGLCLKGQIQQAFLFHDKVVALGFHFDQISYGTLIHGLCKVGE 181
Query: 417 FDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG------KYAH 470
A +L+ ++ PN V+Y +++ + V A ++ +M+ KG Y+
Sbjct: 182 TRAALDLLQRVDGNLVQPNVVMYNTIIDSMCKVKLVNEAFDLFSEMVSKGISPDVVTYSA 361
Query: 471 LLSRF 475
L+S F
Sbjct: 362 LISGF 376
Score = 37.4 bits (85), Expect = 0.013
Identities = 25/104 (24%), Positives = 51/104 (49%)
Frame = +2
Query: 228 SYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEM 287
SY ++H L + + + + Q++ + +++ YN ++ + ++ +++ LF EM
Sbjct: 140 SYGTLIHGLCKVGETRAALDLLQRVDGNLVQPNVVMYNTIIDSMCKVKLVNEAFDLFSEM 319
Query: 288 GRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTV 331
G SPD TY+ L+ K A++L +M I+P V
Sbjct: 320 VSKGISPDVVTYSALISGFCILGKLKDAIDLFNKMILENIKPDV 451
Score = 36.6 bits (83), Expect = 0.022
Identities = 26/119 (21%), Positives = 52/119 (42%)
Frame = +2
Query: 181 EMIEKGFPATARTFNILIRTCGEAGLAKDLVETFIKSKTFNYRPFKHSYNAILHSLLVLN 240
+++ GF ++ LI + G + ++ + +P YN I+ S+ +
Sbjct: 104 KVVALGFHFDQISYGTLIHGLCKVGETRAALDLLQRVDGNLVQPNVVMYNTIIDSMCKVK 283
Query: 241 QYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTY 299
++ +M+ G S D++TY+ ++ LGKL LF +M PD +T+
Sbjct: 284 LVNEAFDLFSEMVSKGISPDVVTYSALISGFCILGKLKDAIDLFNKMILENIKPDVYTF 460
>CA991373 weakly similar to PIR|D86260|D86 protein T12C24.22 [imported] -
Arabidopsis thaliana, partial (6%)
Length = 756
Score = 121 bits (303), Expect = 7e-28
Identities = 69/244 (28%), Positives = 125/244 (50%)
Frame = +2
Query: 223 RPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQI 282
+P +Y +++ ++ + + + M G + DI +YNI++ ++ K+D+
Sbjct: 17 KPIVVTYCSLMDGYCLVKEVNKAKSILYTMSQRGVNPDIQSYNILIDGFCKIKKVDEAMN 196
Query: 283 LFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLS 342
LFKEM PD TYN L+ L K K A+ L+ +M + G+ P ++ Y++++D L
Sbjct: 197 LFKEMHHKHIIPDVVTYNSLIDGLCKLGKISYALKLVDEMHDRGVPPDIITYSSILDALC 376
Query: 343 RAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVY 402
+ + ++ G P++ YT++I G G LE A +F +++ K V
Sbjct: 377 KNHQVDKAIALLTKLKDQGIRPNMYTYTILIDGLCKGGRLEDAHNIFEDLLVKGYNITVN 556
Query: 403 TYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQM 462
TY MI G C G FDEA ++L +M+ C P++V Y ++ L + + A E +R+M
Sbjct: 557 TYTVMIHGFCNKGLFDEALTLLSKMKDNSCIPDAVTYEIIIRSLFDKDENDKA-EKLREM 733
Query: 463 MEKG 466
+ +G
Sbjct: 734 ITRG 745
Score = 97.8 bits (242), Expect = 8e-21
Identities = 55/209 (26%), Positives = 105/209 (49%)
Frame = +2
Query: 256 GFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAA 315
G ++TY +M + ++++ + + M + G +PD +YN+L+ K K A
Sbjct: 11 GIKPIVVTYCSLMDGYCLVKEVNKAKSILYTMSQRGVNPDIQSYNILIDGFCKIKKVDEA 190
Query: 316 VNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITG 375
+NL K+M I P V+ Y +LIDGL + G + DEM G PD++ Y+ ++
Sbjct: 191 MNLFKEMHHKHIIPDVVTYNSLIDGLCKLGKISYALKLVDEMHDRGVPPDIITYSSILDA 370
Query: 376 YVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPN 435
+++KA + ++ + PN+YTY +I GLC G+ ++A ++ +++ KG +
Sbjct: 371 LCKNHQVDKAIALLTKLKDQGIRPNMYTYTILIDGLCKGGRLEDAHNIFEDLLVKGYNIT 550
Query: 436 SVVYVSLLSCLQNAGKVANAREVMRQMME 464
Y ++ N G A ++ +M +
Sbjct: 551 VNTYTVMIHGFCNKGLFDEALTLLSKMKD 637
Score = 91.7 bits (226), Expect = 6e-19
Identities = 45/141 (31%), Positives = 78/141 (54%)
Frame = +2
Query: 326 GIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKA 385
GI+P V+ Y +L+DG ++ K M + G PD+ +Y ++I G+ ++++A
Sbjct: 11 GIKPIVVTYCSLMDGYCLVKEVNKAKSILYTMSQRGVNPDIQSYNILIDGFCKIKKVDEA 190
Query: 386 QEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSC 445
+F EM K +P+V TYNS+I GLC GK A ++ EM +G P+ + Y S+L
Sbjct: 191 MNLFKEMHHKHIIPDVVTYNSLIDGLCKLGKISYALKLVDEMHDRGVPPDIITYSSILDA 370
Query: 446 LQNAGKVANAREVMRQMMEKG 466
L +V A ++ ++ ++G
Sbjct: 371 LCKNHQVDKAIALLTKLKDQG 433
Score = 73.2 bits (178), Expect = 2e-13
Identities = 37/109 (33%), Positives = 62/109 (55%)
Frame = +2
Query: 358 IKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKF 417
+K G P VV Y ++ GY + E+ KA+ + + M + P++ +YN +I G C K
Sbjct: 2 MKQGIKPIVVTYCSLMDGYCLVKEVNKAKSILYTMSQRGVNPDIQSYNILIDGFCKIKKV 181
Query: 418 DEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
DEA ++ KEM K P+ V Y SL+ L GK++ A +++ +M ++G
Sbjct: 182 DEAMNLFKEMHHKHIIPDVVTYNSLIDGLCKLGKISYALKLVDEMHDRG 328
>TC84326 weakly similar to PIR|F85225|F85225 hypothetical protein AT4g19900
[imported] - Arabidopsis thaliana, partial (18%)
Length = 756
Score = 113 bits (282), Expect = 2e-25
Identities = 65/219 (29%), Positives = 107/219 (48%)
Frame = +1
Query: 224 PFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQIL 283
P ++Y ++ ++ + M +GFS ++ TYN ++ + G++ + +
Sbjct: 91 PNTNTYTTLIDGHCKAGNFERAYDLMNLMSSEGFSPNLCTYNAIVNGLCKRGRVQEAYKM 270
Query: 284 FKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSR 343
++ +NG PD TYN+L+ K++ A+ L +M + GI+P + YTTLI R
Sbjct: 271 LEDGFQNGLKPDKFTYNILMSEHCKQENIRQALALFNKMLKIGIQPDIHSYTTLIAVFCR 450
Query: 344 AGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYT 403
+ + FF+E ++ G +P YT MI GY G L A + FH + P T
Sbjct: 451 ENRMKESEMFFEEAVRIGIIPTNKTYTSMICGYCREGNLTLAMKFFHRLSDHGCAPESIT 630
Query: 404 YNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSL 442
Y ++I GLC K DEA S+ M KG P V ++L
Sbjct: 631 YGAIISGLCKQSKRDEARSLYDSMIEKGLVPCEVTKITL 747
Score = 104 bits (260), Expect = 7e-23
Identities = 61/238 (25%), Positives = 108/238 (44%)
Frame = +1
Query: 229 YNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMG 288
Y A++ ++ E + +M G + TY ++ + G ++ L M
Sbjct: 1 YTAMIRGYCREDKLNRAEMLLSRMKEQGLVPNTNTYTTLIDGHCKAGNFERAYDLMNLMS 180
Query: 289 RNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLH 348
GFSP+ TYN +++ L K + A +L+ + G++P Y L+ + N+
Sbjct: 181 SEGFSPNLCTYNAIVNGLCKRGRVQEAYKMLEDGFQNGLKPDKFTYNILMSEHCKQENIR 360
Query: 349 ACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMI 408
F++M+K G PD+ +YT +I + +++++ F E + +P TY SMI
Sbjct: 361 QALALFNKMLKIGIQPDIHSYTTLIAVFCRENRMKESEMFFEEAVRIGIIPTNKTYTSMI 540
Query: 409 RGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
G C G A + GC+P S+ Y +++S L K AR + M+EKG
Sbjct: 541 CGYCREGNLTLAMKFFHRLSDHGCAPESITYGAIISGLCKQSKRDEARSLYDSMIEKG 714
Score = 74.3 bits (181), Expect = 9e-14
Identities = 52/243 (21%), Positives = 98/243 (39%)
Frame = +1
Query: 157 NAYHLIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNILIRTCGEAGLAKDLVETFIK 216
N Y +++ + + ++ + L+ M +GF T+N ++ + G ++ +
Sbjct: 100 NTYTTLIDGHCKAGNFERAYDLMNLMSSEGFSPNLCTYNAIVNGLCKRGRVQEAYKMLED 279
Query: 217 SKTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGK 276
+P K +Y NI+M +
Sbjct: 280 GFQNGLKPDKFTY-----------------------------------NILMSEHCKQEN 354
Query: 277 LDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTT 336
+ Q LF +M + G PD H+Y L+ +E++ + ++ GI PT YT+
Sbjct: 355 IRQALALFNKMLKIGIQPDIHSYTTLIAVFCRENRMKESEMFFEEAVRIGIIPTNKTYTS 534
Query: 337 LIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKE 396
+I G R GNL FF + +GC P+ + Y +I+G + ++A+ ++ MI K
Sbjct: 535 MICGYCREGNLTLAMKFFHRLSDHGCAPESITYGAIISGLCKQSKRDEARSLYDSMIEKG 714
Query: 397 QVP 399
VP
Sbjct: 715 LVP 723
Score = 48.9 bits (115), Expect = 4e-06
Identities = 40/188 (21%), Positives = 76/188 (40%)
Frame = +1
Query: 150 EAYQHTANAYHLIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNILIRTCGEAGLAKD 209
E + Y+ I+N + + ++++ + + G T+NIL+ + +
Sbjct: 184 EGFSPNLCTYNAIVNGLCKRGRVQEAYKMLEDGFQNGLKPDKFTYNILMSEHCKQENIRQ 363
Query: 210 LVETFIKSKTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMY 269
+ F K +P HSY ++ N+ K E +++ + G TY ++
Sbjct: 364 ALALFNKMLKIGIQPDIHSYTTLIAVFCRENRMKESEMFFEEAVRIGIIPTNKTYTSMIC 543
Query: 270 AKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEP 329
R G L F + +G +P+ TY ++ L K+ K A +L M E G+ P
Sbjct: 544 GYCREGNLTLAMKFFHRLSDHGCAPESITYGAIISGLCKQSKRDEARSLYDSMIEKGLVP 723
Query: 330 TVLHYTTL 337
+ TL
Sbjct: 724 CEVTKITL 747
>AW774457 weakly similar to GP|10176973|dbj
gb|AAF19552.1~gene_id:MTG13.9~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (13%)
Length = 650
Score = 112 bits (280), Expect = 3e-25
Identities = 59/194 (30%), Positives = 104/194 (53%)
Frame = -1
Query: 273 RLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVL 332
++ +D+ LF +M G +PD TYN L+ L K + A L+ +M + G +
Sbjct: 632 KIKMVDEALSLFNDMQFKGIAPDKVTYNSLIDGLCKSGRISYAWELVDEMHDNGQPANIF 453
Query: 333 HYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEM 392
Y LID L + ++ ++ G PD+ + ++I G G L+ AQ++F ++
Sbjct: 452 TYNCLIDALCKNHHVDQAIALVKKIKDQGIQPDMYTFNILIYGLCKVGRLKNAQDVFQDL 273
Query: 393 ISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKV 452
+SK N +TYN M+ GLC G FDEA ++L +M+ G P++V Y +L+ L + +
Sbjct: 272 LSKGYSVNAWTYNIMVNGLCKEGLFDEAEALLSKMDDNGIIPDAVTYETLIQALFHKDEN 93
Query: 453 ANAREVMRQMMEKG 466
A +++R+M+ +G
Sbjct: 92 EKAEKLLREMIARG 51
Score = 86.7 bits (213), Expect = 2e-17
Identities = 48/184 (26%), Positives = 92/184 (49%)
Frame = -1
Query: 248 VYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLG 307
++ M G + D +TYN ++ + G++ L EM NG + TYN L+ L
Sbjct: 602 LFNDMQFKGIAPDKVTYNSLIDGLCKSGRISYAWELVDEMHDNGQPANIFTYNCLIDALC 423
Query: 308 KEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVV 367
K A+ L+K++++ GI+P + + LI GL + G L + F +++ G +
Sbjct: 422 KNHHVDQAIALVKKIKDQGIQPDMYTFNILIYGLCKVGRLKNAQDVFQDLLSKGYSVNAW 243
Query: 368 AYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEM 427
Y +M+ G G ++A+ + +M +P+ TY ++I+ L + ++A +L+EM
Sbjct: 242 TYNIMVNGLCKEGLFDEAEALLSKMDDNGIIPDAVTYETLIQALFHKDENEKAEKLLREM 63
Query: 428 ETKG 431
+G
Sbjct: 62 IARG 51
Score = 76.3 bits (186), Expect = 2e-14
Identities = 45/172 (26%), Positives = 84/172 (48%)
Frame = -1
Query: 224 PFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQIL 283
P K +YN+++ L + + +M +G ++I TYN ++ A + +DQ L
Sbjct: 569 PDKVTYNSLIDGLCKSGRISYAWELVDEMHDNGQPANIFTYNCLIDALCKNHHVDQAIAL 390
Query: 284 FKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSR 343
K++ G PD +T+N+L++ L K + A ++ + + G Y +++GL +
Sbjct: 389 VKKIKDQGIQPDMYTFNILIYGLCKVGRLKNAQDVFQDLLSKGYSVNAWTYNIMVNGLCK 210
Query: 344 AGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISK 395
G + +M NG +PD V Y +I E EKA+++ EMI++
Sbjct: 209 EGLFDEAEALLSKMDDNGIIPDAVTYETLIQALFHKDENEKAEKLLREMIAR 54
Score = 74.7 bits (182), Expect = 7e-14
Identities = 45/152 (29%), Positives = 75/152 (48%)
Frame = -1
Query: 176 WRLVGEMIEKGFPATARTFNILIRTCGEAGLAKDLVETFIKSKTFNYRPFKHSYNAILHS 235
W LV EM + G PA T+N LI + + K K +P +++N +++
Sbjct: 503 WELVDEMHDNGQPANIFTYNCLIDALCKNHHVDQAIALVKKIKDQGIQPDMYTFNILIYG 324
Query: 236 LLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPD 295
L + + K + V+Q +L G+S + TYNI++ + G D+ + L +M NG PD
Sbjct: 323 LCKVGRLKNAQDVFQDLLSKGYSVNAWTYNIMVNGLCKEGLFDEAEALLSKMDDNGIIPD 144
Query: 296 FHTYNLLLHFLGKEDKPLAAVNLLKQMRETGI 327
TY L+ L +D+ A LL++M G+
Sbjct: 143 AVTYETLIQALFHKDENEKAEKLLREMIARGL 48
Score = 73.6 bits (179), Expect = 2e-13
Identities = 37/130 (28%), Positives = 68/130 (51%)
Frame = -1
Query: 337 LIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKE 396
+I+G + + F++M G PD V Y +I G +G + A E+ EM
Sbjct: 650 MINGFCKIKMVDEALSLFNDMQFKGIAPDKVTYNSLIDGLCKSGRISYAWELVDEMHDNG 471
Query: 397 QVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAR 456
Q N++TYN +I LC D+A +++K+++ +G P+ + L+ L G++ NA+
Sbjct: 470 QPANIFTYNCLIDALCKNHHVDQAIALVKKIKDQGIQPDMYTFNILIYGLCKVGRLKNAQ 291
Query: 457 EVMRQMMEKG 466
+V + ++ KG
Sbjct: 290 DVFQDLLSKG 261
Score = 49.3 bits (116), Expect = 3e-06
Identities = 23/67 (34%), Positives = 41/67 (60%)
Frame = -1
Query: 407 MIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
MI G C DEA S+ +M+ KG +P+ V Y SL+ L +G+++ A E++ +M + G
Sbjct: 650 MINGFCKIKMVDEALSLFNDMQFKGIAPDKVTYNSLIDGLCKSGRISYAWELVDEMHDNG 471
Query: 467 KYAHLLS 473
+ A++ +
Sbjct: 470 QPANIFT 450
>BE998743 similar to GP|22128591|gb| fertility restorer-like protein {Petunia
x hybrida}, partial (7%)
Length = 829
Score = 108 bits (269), Expect = 6e-24
Identities = 58/215 (26%), Positives = 113/215 (51%)
Frame = +2
Query: 252 MLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDK 311
M+ D++TYN +M + ++++ + + + R G +PD +YN++++ +
Sbjct: 149 MIKGSIKPDVVTYNSLMDGYCLVNEVNKAKHVLSTIARMGVAPDAQSYNIMVNGFCRISH 328
Query: 312 PLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTV 371
A L+ +M G P + Y++LID L + +L ++ G P++ Y +
Sbjct: 329 ---AWKLVDEMHVNGQPPDIFTYSSLIDALCKNNHLDKAIALVKKIKDQGIQPNMYTYNI 499
Query: 372 MITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKG 431
+I G G L+ AQ++F ++++K N+ TYN +I GLC G FD+A ++L +ME
Sbjct: 500 LIDGLCKGGRLKNAQDVFQDLLTKGYSLNIRTYNILINGLCKEGLFDKAEALLSKMEDND 679
Query: 432 CSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
+PN V Y +++ L A +++R+M+ +G
Sbjct: 680 INPNVVTYETIIRSLFYKDYNEKAEKLLREMVARG 784
Score = 92.8 bits (229), Expect = 3e-19
Identities = 47/159 (29%), Positives = 84/159 (52%)
Frame = +2
Query: 308 KEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVV 367
KE A N L M + I+P V+ Y +L+DG ++ KH + + G PD
Sbjct: 107 KEGNVKGAKNALAMMIKGSIKPDVVTYNSLMDGYCLVNEVNKAKHVLSTIARMGVAPDAQ 286
Query: 368 AYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEM 427
+Y +M+ G+ + A ++ EM Q P+++TY+S+I LC D+A +++K++
Sbjct: 287 SYNIMVNGFC---RISHAWKLVDEMHVNGQPPDIFTYSSLIDALCKNNHLDKAIALVKKI 457
Query: 428 ETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
+ +G PN Y L+ L G++ NA++V + ++ KG
Sbjct: 458 KDQGIQPNMYTYNILIDGLCKGGRLKNAQDVFQDLLTKG 574
Score = 72.4 bits (176), Expect = 4e-13
Identities = 60/254 (23%), Positives = 108/254 (41%), Gaps = 4/254 (1%)
Frame = +2
Query: 182 MIEKGFPATARTFNILIR---TCGEAGLAKDLVETFIKSKTFNYRPFKHSYNAILHSLL- 237
MI+ T+N L+ E AK ++ T + P SYN +++
Sbjct: 149 MIKGSIKPDVVTYNSLMDGYCLVNEVNKAKHVLSTIAR---MGVAPDAQSYNIMVNGFCR 319
Query: 238 VLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFH 297
+ + +KL++ +M ++G DI TY+ ++ A + LD+ L K++ G P+ +
Sbjct: 320 ISHAWKLVD----EMHVNGQPPDIFTYSSLIDALCKNNHLDKAIALVKKIKDQGIQPNMY 487
Query: 298 TYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEM 357
TYN+ LIDGL + G L + F ++
Sbjct: 488 TYNI-----------------------------------LIDGLCKGGRLKNAQDVFQDL 562
Query: 358 IKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKF 417
+ G ++ Y ++I G G +KA+ + +M + PNV TY ++IR L
Sbjct: 563 LTKGYSLNIRTYNILINGLCKEGLFDKAEALLSKMEDNDINPNVVTYETIIRSLFYKDYN 742
Query: 418 DEACSMLKEMETKG 431
++A +L+EM +G
Sbjct: 743 EKAEKLLREMVARG 784
Score = 60.8 bits (146), Expect = 1e-09
Identities = 47/205 (22%), Positives = 92/205 (43%)
Frame = +2
Query: 159 YHLIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNILIRTCGEAGLAKDLVETFIKSK 218
Y+ +M+ Y E ++ + G A+++NI++ A LV+ +
Sbjct: 185 YNSLMDGYCLVNEVNKAKHVLSTIARMGVAPDAQSYNIMVNGFCRISHAWKLVD---EMH 355
Query: 219 TFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLD 278
P +Y++++ +L N + +++ G ++ TYNI++ + G+L
Sbjct: 356 VNGQPPDIFTYSSLIDALCKNNHLDKAIALVKKIKDQGIQPNMYTYNILIDGLCKGGRLK 535
Query: 279 QFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLI 338
Q +F+++ G+S + TYN+L++ L KE A LL +M + I P V+ Y T+I
Sbjct: 536 NAQDVFQDLLTKGYSLNIRTYNILINGLCKEGLFDKAEALLSKMEDNDINPNVVTYETII 715
Query: 339 DGLSRAGNLHACKHFFDEMIKNGCM 363
L + EM+ G +
Sbjct: 716 RSLFYKDYNEKAEKLLREMVARGLL 790
Score = 33.9 bits (76), Expect = 0.14
Identities = 34/166 (20%), Positives = 68/166 (40%), Gaps = 6/166 (3%)
Frame = +2
Query: 103 VLDELQV--RPSGLLVREVLVGILKSINHENRTRCAKLAYKFFVWCSQQEAYQHTANAYH 160
++DE+ V +P + L+ L NH ++ K + Q Y+
Sbjct: 338 LVDEMHVNGQPPDIFTYSSLIDALCKNNHLDKAIALVKKIK-------DQGIQPNMYTYN 496
Query: 161 LIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNILIRTCGEAGLAKDLVETFIKSKTF 220
++++ + K + +++ KG+ RT+NILI + GL K +
Sbjct: 497 ILIDGLCKGGRLKNAQDVFQDLLTKGYSLNIRTYNILINGLCKEGLFDKAEALLSKMEDN 676
Query: 221 NYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDG----FSSDIL 262
+ P +Y I+ SL + + E + ++M+ G F+ D+L
Sbjct: 677 DINPNVVTYETIIRSLFYKDYNEKAEKLLREMVARGLL*EFTLDLL 814
>BF649699 similar to GP|9757911|dbj gb|AAF19552.1~gene_id:MMN10.14~similar to
unknown protein {Arabidopsis thaliana}, partial (11%)
Length = 617
Score = 103 bits (258), Expect = 1e-22
Identities = 57/184 (30%), Positives = 95/184 (50%)
Frame = +1
Query: 256 GFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAA 315
GF ++ YN ++ A + LD+ ++L+K M + TY++L+ K A
Sbjct: 64 GFLPNLFVYNALINALCKGEDLDKAELLYKNMHSMNLPLNDVTYSILIDSFCKRGMLDVA 243
Query: 316 VNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITG 375
+ +M E GI T+ Y +LI+G + G+L A + + +MI G P +T +I+G
Sbjct: 244 ESYFGRMIEDGIRETIYPYNSLINGHCKFGDLSAAEFLYTKMINEGLEPTATTFTTLISG 423
Query: 376 YVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPN 435
Y ++EKA +++ EM KE P+VYT+ ++I GLC + EA + EM + P
Sbjct: 424 YCKDLQVEKAFKLYREMNEKEXAPSVYTFTALIYGLCSTNEMAEASKLFDEMVERKIKPT 603
Query: 436 SVVY 439
V Y
Sbjct: 604 XVTY 615
Score = 79.0 bits (193), Expect = 4e-15
Identities = 57/207 (27%), Positives = 94/207 (44%)
Frame = +1
Query: 198 IRTCGEAGLAKDLVETFIKSKTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGF 257
+R G A DLV +K F + P YNA++++L E +Y+ M
Sbjct: 4 LRKKGNIDSAYDLV---VKLGRFGFLPNLFVYNALINALCKGEDLDKAELLYKNMHSMNL 174
Query: 258 SSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVN 317
+ +TY+I++ + + G LD + F M +G + YN L++ K AA
Sbjct: 175 PLNDVTYSILIDSFCKRGMLDVAESYFGRMIEDGIRETIYPYNSLINGHCKFGDLSAAEF 354
Query: 318 LLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYV 377
L +M G+EPT +TTLI G + + + EM + P V +T +I G
Sbjct: 355 LYTKMINEGLEPTATTFTTLISGYCKDLQVEKAFKLYREMNEKEXAPSVYTFTALIYGLC 534
Query: 378 VAGELEKAQEMFHEMISKEQVPNVYTY 404
E+ +A ++F EM+ ++ P TY
Sbjct: 535 STNEMAEASKLFDEMVERKIKPTXVTY 615
Score = 40.8 bits (94), Expect = 0.001
Identities = 22/87 (25%), Positives = 40/87 (45%)
Frame = +1
Query: 380 GELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVY 439
G ++ A ++ ++ +PN++ YN++I LC D+A + K M + N V Y
Sbjct: 16 GNIDSAYDLVVKLGRFGFLPNLFVYNALINALCKGEDLDKAELLYKNMHSMNLPLNDVTY 195
Query: 440 VSLLSCLQNAGKVANAREVMRQMMEKG 466
L+ G + A +M+E G
Sbjct: 196 SILIDSFCKRGMLDVAESYFGRMIEDG 276
>TC81904 weakly similar to GP|8778410|gb|AAF79418.1| F16A14.3 {Arabidopsis
thaliana}, partial (7%)
Length = 978
Score = 98.2 bits (243), Expect = 6e-21
Identities = 65/257 (25%), Positives = 122/257 (47%), Gaps = 6/257 (2%)
Frame = +1
Query: 214 FIKSKTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYR 273
F++ + P + Y A++H + VY+ M+ G ++ + ++ +++
Sbjct: 31 FLEMEKQGLVPDVYVYCALVHGYCNSRNFDKALAVYKSMISRGIKTNCVIFSCILHCLDE 210
Query: 274 LGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLH 333
+G+ + +F+E +G D YN+L L K K AV +L +++ ++ + H
Sbjct: 211 MGRALEVVDMFEEFKESGLFIDRKAYNILFDALCKLGKVDDAVGMLDELKSMQLDVDMKH 390
Query: 334 YTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMI 393
YTTLI+G G + F EM + G PDVVAY V+ G+ +A ++ + M
Sbjct: 391 YTTLINGYFLQGKPIEAQSLFKEMEERGFKPDVVAYNVLAAGFFRNRTDFEAMDLLNYME 570
Query: 394 SKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVA 453
S+ PN T+ +I GLC AGK +EA ++ + + +Y +L++ A +
Sbjct: 571 SQGVEPNSTTHKIIIEGLCSAGKVEEAEEFFNWLKGESVEISVEIYTALVNGYCEAALIE 750
Query: 454 NARE------VMRQMME 464
+ E ++R M+E
Sbjct: 751 KSHELKEAFILLRTMLE 801
Score = 90.5 bits (223), Expect = 1e-18
Identities = 57/212 (26%), Positives = 98/212 (45%)
Frame = +1
Query: 246 EWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHF 305
E V+ +M G D+ Y +++ D+ ++K M G + ++ +LH
Sbjct: 22 ESVFLEMEKQGLVPDVYVYCALVHGYCNSRNFDKALAVYKSMISRGIKTNCVIFSCILHC 201
Query: 306 LGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPD 365
L + + L V++ ++ +E+G+ Y L D L + G + DE+ D
Sbjct: 202 LDEMGRALEVVDMFEEFKESGLFIDRKAYNILFDALCKLGKVDDAVGMLDELKSMQLDVD 381
Query: 366 VVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLK 425
+ YT +I GY + G+ +AQ +F EM + P+V YN + G EA +L
Sbjct: 382 MKHYTTLINGYFLQGKPIEAQSLFKEMEERGFKPDVVAYNVLAAGFFRNRTDFEAMDLLN 561
Query: 426 EMETKGCSPNSVVYVSLLSCLQNAGKVANARE 457
ME++G PNS + ++ L +AGKV A E
Sbjct: 562 YMESQGVEPNSTTHKIIIEGLCSAGKVEEAEE 657
Score = 85.9 bits (211), Expect = 3e-17
Identities = 59/244 (24%), Positives = 106/244 (43%), Gaps = 6/244 (2%)
Frame = +1
Query: 229 YNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMG 288
++ ILH L + + + ++++ G D YNI+ A +LGK+D + E+
Sbjct: 181 FSCILHCLDEMGRALEVVDMFEEFKESGLFIDRKAYNILFDALCKLGKVDDAVGMLDELK 360
Query: 289 RNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLH 348
D Y L++ + KP+ A +L K+M E G +P V+ Y L G R
Sbjct: 361 SMQLDVDMKHYTTLINGYFLQGKPIEAQSLFKEMEERGFKPDVVAYNVLAAGFFRNRTDF 540
Query: 349 ACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMI 408
+ M G P+ + ++I G AG++E+A+E F+ + + +V Y +++
Sbjct: 541 EAMDLLNYMESQGVEPNSTTHKIIIEGLCSAGKVEEAEEFFNWLKGESVEISVEIYTALV 720
Query: 409 RGLCMAG------KFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQM 462
G C A + EA +L+ M P+ V+Y + + L G + A +
Sbjct: 721 NGYCEAALIEKSHELKEAFILLRTMLEMNMKPSKVMYSKIFTALCCNGNMEGAHTLFNLF 900
Query: 463 MEKG 466
G
Sbjct: 901 XHTG 912
Score = 60.8 bits (146), Expect = 1e-09
Identities = 26/88 (29%), Positives = 50/88 (56%)
Frame = +1
Query: 381 ELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYV 440
+L++A+ +F EM + VP+VY Y +++ G C + FD+A ++ K M ++G N V++
Sbjct: 7 KLDEAESVFLEMEKQGLVPDVYVYCALVHGYCNSRNFDKALAVYKSMISRGIKTNCVIFS 186
Query: 441 SLLSCLQNAGKVANAREVMRQMMEKGKY 468
+L CL G+ ++ + E G +
Sbjct: 187 CILHCLDEMGRALEVVDMFEEFKESGLF 270
Score = 57.8 bits (138), Expect = 9e-09
Identities = 38/158 (24%), Positives = 68/158 (42%)
Frame = +1
Query: 309 EDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVA 368
E K A ++ +M + G+ P V Y L+ G + N + MI G + V
Sbjct: 1 ETKLDEAESVFLEMEKQGLVPDVYVYCALVHGYCNSRNFDKALAVYKSMISRGIKTNCVI 180
Query: 369 YTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEME 428
++ ++ G + +MF E + YN + LC GK D+A ML E++
Sbjct: 181 FSCILHCLDEMGRALEVVDMFEEFKESGLFIDRKAYNILFDALCKLGKVDDAVGMLDELK 360
Query: 429 TKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
+ + Y +L++ GK A+ + ++M E+G
Sbjct: 361 SMQLDVDMKHYTTLINGYFLQGKPIEAQSLFKEMEERG 474
Score = 43.1 bits (100), Expect = 2e-04
Identities = 45/232 (19%), Positives = 83/232 (35%), Gaps = 41/232 (17%)
Frame = +1
Query: 181 EMIEKGFPATARTFNILIRTCGEAGLAKDLVETFIKSKTFNYRPFKHSYNAILHSLLVLN 240
E E G + +NIL + G D V + K+ Y +++ +
Sbjct: 247 EFKESGLFIDRKAYNILFDALCKLGKVDDAVGMLDELKSMQLDVDMKHYTTLINGYFLQG 426
Query: 241 QYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYN 300
+ + ++++M GF D++ YN++ +R + L M G P+ T+
Sbjct: 427 KPIEAQSLFKEMEERGFKPDVVAYNVLAAGFFRNRTDFEAMDLLNYMESQGVEPNSTTHK 606
Query: 301 LLLHFL---GKEDKP-------------------LAAVN-------------------LL 319
+++ L GK ++ A VN LL
Sbjct: 607 IIIEGLCSAGKVEEAEEFFNWLKGESVEISVEIYTALVNGYCEAALIEKSHELKEAFILL 786
Query: 320 KQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTV 371
+ M E ++P+ + Y+ + L GN+ F+ G PD V YT+
Sbjct: 787 RTMLEMNMKPSKVMYSKIFTALCCNGNMEGAHTLFNLFXHTGFTPDAVTYTI 942
>TC93383 similar to PIR|T47786|T47786 hypothetical protein F17J16.90 -
Arabidopsis thaliana, partial (43%)
Length = 1038
Score = 95.1 bits (235), Expect = 5e-20
Identities = 54/205 (26%), Positives = 101/205 (48%)
Frame = +1
Query: 262 LTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQ 321
+TYN +M + ++ ++ +M R PD +Y LL++ GK + A+ + ++
Sbjct: 7 VTYNSLMSFETNYKEVSN---IYDQMQRADLRPDVVSYALLINAYGKARREEEALAVFEE 177
Query: 322 MRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGE 381
M + G+ PT Y L+D S +G + + F M ++ MPD+ +YT M++ YV A +
Sbjct: 178 MLDAGVRPTRKAYNILLDAFSISGMVEQARIVFKSMRRDKYMPDLCSYTTMLSAYVNAPD 357
Query: 382 LEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVS 441
+E A++ F +I PNV TY ++I+G A ++ +EM +G N + +
Sbjct: 358 MEGAEKFFKRLIQDGFEPNVVTYGTLIKGYAKANDIEKVMEKYEEMLGRGIKANQTILTT 537
Query: 442 LLSCLQNAGKVANAREVMRQMMEKG 466
++ G +A ++M G
Sbjct: 538 IMDAHGKNGDFDSAVNWFKEMALNG 612
Score = 94.4 bits (233), Expect = 9e-20
Identities = 59/223 (26%), Positives = 101/223 (44%)
Frame = +1
Query: 242 YKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNL 301
YK + +Y QM D+++Y +++ A + + ++ +F+EM G P YN+
Sbjct: 43 YKEVSNIYDQMQRADLRPDVVSYALLINAYGKARREEEALAVFEEMLDAGVRPTRKAYNI 222
Query: 302 LLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNG 361
LL A + K MR P + YTT++ A ++ + FF +I++G
Sbjct: 223 LLDAFSISGMVEQARIVFKSMRRDKYMPDLCSYTTMLSAYVNAPDMEGAEKFFKRLIQDG 402
Query: 362 CMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEAC 421
P+VV Y +I GY A ++EK E + EM+ + N +++ G FD A
Sbjct: 403 FEPNVVTYGTLIKGYAKANDIEKVMEKYEEMLGRGIKANQTILTTIMDAHGKNGDFDSAV 582
Query: 422 SMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMME 464
+ KEM G P+ LLS + + A E++ +E
Sbjct: 583 NWFKEMALNGLLPDQKAKNILLSLAKTEEDIKEANELVLHSIE 711
Score = 70.5 bits (171), Expect = 1e-12
Identities = 47/201 (23%), Positives = 93/201 (45%)
Frame = +1
Query: 208 KDLVETFIKSKTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIV 267
K++ + + + + RP SY ++++ + + V+++ML G YNI+
Sbjct: 46 KEVSNIYDQMQRADLRPDVVSYALLINAYGKARREEEALAVFEEMLDAGVRPTRKAYNIL 225
Query: 268 MYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGI 327
+ A G ++Q +I+FK M R+ + PD +Y +L A K++ + G
Sbjct: 226 LDAFSISGMVEQARIVFKSMRRDKYMPDLCSYTTMLSAYVNAPDMEGAEKFFKRLIQDGF 405
Query: 328 EPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQE 387
EP V+ Y TLI G ++A ++ ++EM+ G + T ++ + G+ + A
Sbjct: 406 EPNVVTYGTLIKGYAKANDIEKVMEKYEEMLGRGIKANQTILTTIMDAHGKNGDFDSAVN 585
Query: 388 MFHEMISKEQVPNVYTYNSMI 408
F EM +P+ N ++
Sbjct: 586 WFKEMALNGLLPDQKAKNILL 648
Score = 57.8 bits (138), Expect = 9e-09
Identities = 36/172 (20%), Positives = 77/172 (43%)
Frame = +1
Query: 158 AYHLIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNILIRTCGEAGLAKDLVETFIKS 217
+Y L++N Y + + + EM++ G T + +NIL+ +G+ + F
Sbjct: 106 SYALLINAYGKARREEEALAVFEEMLDAGVRPTRKAYNILLDAFSISGMVEQARIVFKSM 285
Query: 218 KTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKL 277
+ Y P SY +L + + + E +++++ DGF +++TY ++ + +
Sbjct: 286 RRDKYMPDLCSYTTMLSAYVNAPDMEGAEKFFKRLIQDGFEPNVVTYGTLIKGYAKANDI 465
Query: 278 DQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEP 329
++ ++EM G + ++ GK +AVN K+M G+ P
Sbjct: 466 EKVMEKYEEMLGRGIKANQTILTTIMDAHGKNGDFDSAVNWFKEMALNGLLP 621
>TC83856 weakly similar to PIR|B96656|B96656 unknown protein 41955-40111
[imported] - Arabidopsis thaliana, partial (12%)
Length = 662
Score = 93.6 bits (231), Expect = 2e-19
Identities = 45/146 (30%), Positives = 79/146 (53%)
Frame = +2
Query: 294 PDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHF 353
PD +T+N+L+ K K A+ L+ +M + G P ++ Y++++D L + +
Sbjct: 74 PDVYTFNILVDVFCKSGKISYALKLVDEMHDRGQPPNIVTYSSILDALCKTHRVDKAVAL 253
Query: 354 FDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCM 413
++ G P++ YT++I G +G+LE A+ +F +++ K V TY M G C
Sbjct: 254 LTKLKDQGIRPNMHTYTILIDGLCTSGKLEDARNIFEDLLVKGYDITVVTYIVMFYGFCK 433
Query: 414 AGKFDEACSMLKEMETKGCSPNSVVY 439
G FDEA ++L +ME GC P++ Y
Sbjct: 434 KGLFDEASALLSKMEENGCIPDAKTY 511
Score = 83.6 bits (205), Expect = 2e-16
Identities = 51/167 (30%), Positives = 83/167 (49%)
Frame = +2
Query: 254 LDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPL 313
L+ D+ T+NI++ + GK+ L EM G P+ TY+ +L L K +
Sbjct: 59 LENIKPDVYTFNILVDVFCKSGKISYALKLVDEMHDRGQPPNIVTYSSILDALCKTHRVD 238
Query: 314 AAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMI 373
AV LL ++++ GI P + YT LIDGL +G L ++ F++++ G VV Y VM
Sbjct: 239 KAVALLTKLKDQGIRPNMHTYTILIDGLCTSGKLEDARNIFEDLLVKGYDITVVTYIVMF 418
Query: 374 TGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEA 420
G+ G ++A + +M +P+ TY + L G+ D A
Sbjct: 419 YGFCKKGLFDEASALLSKMEENGCIPDAKTYELIKLSLFKKGENDMA 559
Score = 69.3 bits (168), Expect = 3e-12
Identities = 30/103 (29%), Positives = 60/103 (58%)
Frame = +2
Query: 364 PDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSM 423
PDV + +++ + +G++ A ++ EM + Q PN+ TY+S++ LC + D+A ++
Sbjct: 74 PDVYTFNILVDVFCKSGKISYALKLVDEMHDRGQPPNIVTYSSILDALCKTHRVDKAVAL 253
Query: 424 LKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
L +++ +G PN Y L+ L +GK+ +AR + ++ KG
Sbjct: 254 LTKLKDQGIRPNMHTYTILIDGLCTSGKLEDARNIFEDLLVKG 382
Score = 65.5 bits (158), Expect = 4e-11
Identities = 40/160 (25%), Positives = 76/160 (47%)
Frame = +2
Query: 307 GKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDV 366
G + L + L+K E I+P V + L+D ++G + DEM G P++
Sbjct: 11 GTRAERLQLICLIK*YLEN-IKPDVYTFNILVDVFCKSGKISYALKLVDEMHDRGQPPNI 187
Query: 367 VAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKE 426
V Y+ ++ ++KA + ++ + PN++TY +I GLC +GK ++A ++ ++
Sbjct: 188 VTYSSILDALCKTHRVDKAVALLTKLKDQGIRPNMHTYTILIDGLCTSGKLEDARNIFED 367
Query: 427 METKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
+ KG V Y+ + G A ++ +M E G
Sbjct: 368 LLVKGYDITVVTYIVMFYGFCKKGLFDEASALLSKMEENG 487
Score = 61.6 bits (148), Expect = 6e-10
Identities = 31/144 (21%), Positives = 68/144 (46%)
Frame = +2
Query: 159 YHLIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNILIRTCGEAGLAKDLVETFIKSK 218
++++++++ + + +LV EM ++G P T++ ++ + V K K
Sbjct: 89 FNILVDVFCKSGKISYALKLVDEMHDRGQPPNIVTYSSILDALCKTHRVDKAVALLTKLK 268
Query: 219 TFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLD 278
RP H+Y ++ L + + +++ +L+ G+ ++TY ++ Y + G D
Sbjct: 269 DQGIRPNMHTYTILIDGLCTSGKLEDARNIFEDLLVKGYDITVVTYIVMFYGFCKKGLFD 448
Query: 279 QFQILFKEMGRNGFSPDFHTYNLL 302
+ L +M NG PD TY L+
Sbjct: 449 EASALLSKMEENGCIPDAKTYELI 520
>TC82835 weakly similar to GP|21741712|emb|CAD41335. oj991113_30.17 {Oryza
sativa}, partial (5%)
Length = 915
Score = 92.8 bits (229), Expect = 3e-19
Identities = 51/166 (30%), Positives = 90/166 (53%), Gaps = 1/166 (0%)
Frame = +3
Query: 265 NIVMYAKYRLGKLDQFQILFKEMG-RNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMR 323
N +++A + G+ D +F+ + +N D TYN++L LG++ + +++ ++
Sbjct: 18 NKIIFAFAKSGQKDSALAIFEHLREQNNSCLDLITYNIVLDILGRKGRVDEMLDMFASLK 197
Query: 324 ETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELE 383
ETG P + Y TLI+GL + G C +F EM +NG PD++ YT +I AG +E
Sbjct: 198 ETGFVPDTISYNTLINGLRKVGRSDMCFEYFKEMKENGNEPDLLTYTALIDISGRAGNIE 377
Query: 384 KAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMET 429
++ + F EM K +P++ Y S+I L + A +L+EM +
Sbjct: 378 ESLKFFMEMKLKGILPSIQIYRSLIHNLNKTENIELATELLEEMNS 515
Score = 86.3 bits (212), Expect = 2e-17
Identities = 49/135 (36%), Positives = 70/135 (51%)
Frame = +3
Query: 260 DILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLL 319
D++TYNIV+ R G++D+ +F + GF PD +YN L++ L K +
Sbjct: 111 DLITYNIVLDILGRKGRVDEMLDMFASLKETGFVPDTISYNTLINGLRKVGRSDMCFEYF 290
Query: 320 KQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVA 379
K+M+E G EP +L YT LID RAGN+ FF EM G +P + Y +I
Sbjct: 291 KEMKENGNEPDLLTYTALIDISGRAGNIEESLKFFMEMKLKGILPSIQIYRSLIHNLNKT 470
Query: 380 GELEKAQEMFHEMIS 394
+E A E+ EM S
Sbjct: 471 ENIELATELLEEMNS 515
Score = 61.6 bits (148), Expect = 6e-10
Identities = 34/133 (25%), Positives = 63/133 (46%)
Frame = +3
Query: 193 TFNILIRTCGEAGLAKDLVETFIKSKTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQM 252
T+NI++ G G ++++ F K + P SYN +++ L + + + +++M
Sbjct: 120 TYNIVLDILGRKGRVDEMLDMFASLKETGFVPDTISYNTLINGLRKVGRSDMCFEYFKEM 299
Query: 253 LLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKP 312
+G D+LTY ++ R G +++ F EM G P Y L+H L K +
Sbjct: 300 KENGNEPDLLTYTALIDISGRAGNIEESLKFFMEMKLKGILPSIQIYRSLIHNLNKTENI 479
Query: 313 LAAVNLLKQMRET 325
A LL++M +
Sbjct: 480 ELATELLEEMNSS 518
Score = 58.2 bits (139), Expect = 7e-09
Identities = 34/130 (26%), Positives = 61/130 (46%)
Frame = +3
Query: 228 SYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEM 287
+YN +L L + + ++ + GF D ++YN ++ ++G+ D FKEM
Sbjct: 120 TYNIVLDILGRKGRVDEMLDMFASLKETGFVPDTISYNTLINGLRKVGRSDMCFEYFKEM 299
Query: 288 GRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNL 347
NG PD TY L+ G+ ++ +M+ GI P++ Y +LI L++ N+
Sbjct: 300 KENGNEPDLLTYTALIDISGRAGNIEESLKFFMEMKLKGILPSIQIYRSLIHNLNKTENI 479
Query: 348 HACKHFFDEM 357
+EM
Sbjct: 480 ELATELLEEM 509
Score = 55.1 bits (131), Expect = 6e-08
Identities = 31/132 (23%), Positives = 64/132 (48%), Gaps = 2/132 (1%)
Frame = +3
Query: 337 LIDGLSRAGNLHACKHFFDEMIK--NGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMIS 394
+I +++G + F+ + + N C+ D++ Y +++ G +++ +MF +
Sbjct: 24 IIFAFAKSGQKDSALAIFEHLREQNNSCL-DLITYNIVLDILGRKGRVDEMLDMFASLKE 200
Query: 395 KEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVAN 454
VP+ +YN++I GL G+ D KEM+ G P+ + Y +L+ AG +
Sbjct: 201 TGFVPDTISYNTLINGLRKVGRSDMCFEYFKEMKENGNEPDLLTYTALIDISGRAGNIEE 380
Query: 455 AREVMRQMMEKG 466
+ + +M KG
Sbjct: 381 SLKFFMEMKLKG 416
>TC80520 weakly similar to PIR|T02562|T02562 probable salt-inducible protein
[imported] - Arabidopsis thaliana, partial (6%)
Length = 1911
Score = 91.3 bits (225), Expect = 7e-19
Identities = 69/290 (23%), Positives = 130/290 (44%), Gaps = 2/290 (0%)
Frame = +1
Query: 124 LKSINHENRTRCAKLAYKFFVWCSQQEAYQHTANAYHLIMNIYAECEEYKALWRLVGEMI 183
L + H+N T L+ +F W + ++ ++ + + + + KA L+
Sbjct: 181 LSYLKHQNNTF---LSLRFLHWLTSHCGFKPDQSSCNALFDALVDAGAVKAAKSLLEY-- 345
Query: 184 EKGFPATARTFNILIRTCGEAGLAKDLVETFIKSKTFNYRPFKHSYNAILHSLLVLNQYK 243
F + +R GE G+ +++ + F+ K + P S+N L + L + +
Sbjct: 346 -PDFVPKNDSLEGYVRLLGENGMVEEVFDVFVSLKKVGFLPSASSFNVCLLACLKVGRTD 522
Query: 244 LIEWVYQQMLLDGF--SSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNL 301
L+ +Y+ M+ G + D+ T ++ A K+ L +++ G D +N
Sbjct: 523 LVWKLYELMIESGVGVNIDVETVGCLIKAFCAENKVFNGYELLRQVLEKGLCVDNTVFNA 702
Query: 302 LLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNG 361
L++ K+ + +L M P++ Y +I+GL + F+++ G
Sbjct: 703 LINGFCKQKQYDRVSEILHIMIAMKCNPSIYTYQEIINGLLKRRKNDEAFRVFNDLKDRG 882
Query: 362 CMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGL 411
PD V YT +I G+ G L +A++++ EMI K VPN YTYN MI G+
Sbjct: 883 YFPDRVMYTTVIKGFCDMGLLAEARKLWFEMIQKGLVPNEYTYNVMIYGI 1032
Score = 83.2 bits (204), Expect = 2e-16
Identities = 51/184 (27%), Positives = 88/184 (47%)
Frame = +3
Query: 248 VYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLG 307
+Y M G++ ++++Y+ ++ Y GK D+ LF EM R G + D +YN L+ L
Sbjct: 1065 LYDDMCGRGYAENVVSYSTMISGLYLHGKTDEALSLFHEMSRKGIARDLISYNSLIKGLC 1244
Query: 308 KEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVV 367
+E + A NLL ++ G+EP+V +T LI L + G+ +M P
Sbjct: 1245 QEGELAKATNLLNKLLIQGLEPSVSSFTPLIKCLCKVGDTEGAMRLLKDMHDRHLEPIAS 1424
Query: 368 AYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEM 427
+ MI G G+ + E +M+S + PN+ T+ +I L + D+ +L M
Sbjct: 1425 THDYMIIGLSKKGDFAQGMEWLLKMLSWKLKPNMQTFEHLIDCLKRENRLDDILIVLDLM 1604
Query: 428 ETKG 431
+G
Sbjct: 1605 FREG 1616
Score = 75.5 bits (184), Expect = 4e-14
Identities = 34/115 (29%), Positives = 66/115 (56%)
Frame = +3
Query: 351 KHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRG 410
+ +D+M G +VV+Y+ MI+G + G+ ++A +FHEM K ++ +YNS+I+G
Sbjct: 1059 RKLYDDMCGRGYAENVVSYSTMISGLYLHGKTDEALSLFHEMSRKGIARDLISYNSLIKG 1238
Query: 411 LCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEK 465
LC G+ +A ++L ++ +G P+ + L+ CL G A +++ M ++
Sbjct: 1239 LCQEGELAKATNLLNKLLIQGLEPSVSSFTPLIKCLCKVGDTEGAMRLLKDMHDR 1403
Score = 75.1 bits (183), Expect = 6e-14
Identities = 52/217 (23%), Positives = 100/217 (45%), Gaps = 6/217 (2%)
Frame = +3
Query: 265 NIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRE 324
N + Y K R + + L+ +M G++ + +Y+ ++ L K A++L +M
Sbjct: 1017 NDIWYCKVR--DFAEARKLYDDMCGRGYAENVVSYSTMISGLYLHGKTDEALSLFHEMSR 1190
Query: 325 TGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEK 384
GI ++ Y +LI GL + G L + ++++ G P V ++T +I G+ E
Sbjct: 1191 KGIARDLISYNSLIKGLCQEGELAKATNLLNKLLIQGLEPSVSSFTPLIKCLCKVGDTEG 1370
Query: 385 AQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLS 444
A + +M + P T++ MI GL G F + L +M + PN + L+
Sbjct: 1371 AMRLLKDMHDRHLEPIASTHDYMIIGLSKKGDFAQGMEWLLKMLSWKLKPNMQTFEHLID 1550
Query: 445 CLQNAGKVANAREVMRQM------MEKGKYAHLLSRF 475
CL+ ++ + V+ M +EK + L+++F
Sbjct: 1551 CLKRENRLDDILIVLDLMFREGYRLEKSRICSLVTKF 1661
Score = 72.8 bits (177), Expect = 3e-13
Identities = 55/251 (21%), Positives = 114/251 (44%), Gaps = 6/251 (2%)
Frame = +1
Query: 222 YRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRL----GKL 277
++P + S NA+ +L+ K + + + D + N + RL G +
Sbjct: 256 FKPDQSSCNALFDALVDAGAVKAAKSLLEY-------PDFVPKNDSLEGYVRLLGENGMV 414
Query: 278 DQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTT- 336
++ +F + + GF P ++N+ L K + L + M E+G+ + T
Sbjct: 415 EEVFDVFVSLKKVGFLPSASSFNVCLLACLKVGRTDLVWKLYELMIESGVGVNIDVETVG 594
Query: 337 -LIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISK 395
LI + ++++ G D + +I G+ + ++ E+ H MI+
Sbjct: 595 CLIKAFCAENKVFNGYELLRQVLEKGLCVDNTVFNALINGFCKQKQYDRVSEILHIMIAM 774
Query: 396 EQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANA 455
+ P++YTY +I GL K DEA + +++ +G P+ V+Y +++ + G +A A
Sbjct: 775 KCNPSIYTYQEIINGLLKRRKNDEAFRVFNDLKDRGYFPDRVMYTTVIKGFCDMGLLAEA 954
Query: 456 REVMRQMMEKG 466
R++ +M++KG
Sbjct: 955 RKLWFEMIQKG 987
Score = 47.4 bits (111), Expect = 1e-05
Identities = 42/252 (16%), Positives = 96/252 (37%), Gaps = 2/252 (0%)
Frame = +1
Query: 219 TFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLD 278
+FN+ K + + H + + W+ GF D + N + A G +
Sbjct: 151 SFNFSNPKFFLSYLKHQNNTFLSLRFLHWLTSHC---GFKPDQSSCNALFDALVDAGAVK 321
Query: 279 QFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLI 338
+ L + F P + + LG+ ++ +++ G P+ + +
Sbjct: 322 AAKSLLEYPD---FVPKNDSLEGYVRLLGENGMVEEVFDVFVSLKKVGFLPSASSFNVCL 492
Query: 339 DGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTV--MITGYVVAGELEKAQEMFHEMISKE 396
+ G ++ MI++G ++ TV +I + ++ E+ +++ K
Sbjct: 493 LACLKVGRTDLVWKLYELMIESGVGVNIDVETVGCLIKAFCAENKVFNGYELLRQVLEKG 672
Query: 397 QVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAR 456
+ +N++I G C ++D +L M C+P+ Y +++ L K A
Sbjct: 673 LCVDNTVFNALINGFCKQKQYDRVSEILHIMIAMKCNPSIYTYQEIINGLLKRRKNDEAF 852
Query: 457 EVMRQMMEKGKY 468
V + ++G +
Sbjct: 853 RVFNDLKDRGYF 888
>TC82633 similar to GP|15450347|gb|AAK96467.1 At1g20300/F14O10_8
{Arabidopsis thaliana}, partial (34%)
Length = 830
Score = 86.7 bits (213), Expect = 2e-17
Identities = 47/185 (25%), Positives = 97/185 (52%), Gaps = 1/185 (0%)
Frame = +1
Query: 283 LFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLS 342
++ +M R G + D +YN L+ K++ AV +L M + G+ P + ++ ++
Sbjct: 16 VYNQMKRLGCAADTISYNFLIESHCKDENLDEAVKVLDTMVKKGVAPNASTFNSIFGCIA 195
Query: 343 RAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVY 402
+++ + +M + CMP+ + Y +++ + + ++ ++ EM E PNV
Sbjct: 196 ELHDVNGAHRMYAKMKELKCMPNTLTYNILMRMFADSKSIDMVLKLKKEMDESEVEPNVN 375
Query: 403 TYNSMIRGLCMAGKFDEACSMLKEM-ETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQ 461
TY +I C G ++ A +++KEM E K PN +Y ++L L+NAG++ E++ +
Sbjct: 376 TYRILILMFCEKGHWNNAYNLMKEMVEEKCLKPNLSIYETVLELLRNAGQLKKHEELVEK 555
Query: 462 MMEKG 466
M+ +G
Sbjct: 556 MVARG 570
Score = 59.3 bits (142), Expect = 3e-09
Identities = 36/150 (24%), Positives = 63/150 (42%)
Frame = +1
Query: 316 VNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITG 375
+ + QM+ G + Y LI+ + NL D M+K G P+ + +
Sbjct: 10 LQVYNQMKRLGCAADTISYNFLIESHCKDENLDEAVKVLDTMVKKGVAPNASTFNSIFGC 189
Query: 376 YVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPN 435
++ A M+ +M + +PN TYN ++R + D + KEM+ PN
Sbjct: 190 IAELHDVNGAHRMYAKMKELKCMPNTLTYNILMRMFADSKSIDMVLKLKKEMDESEVEPN 369
Query: 436 SVVYVSLLSCLQNAGKVANAREVMRQMMEK 465
Y L+ G NA +M++M+E+
Sbjct: 370 VNTYRILILMFCEKGHWNNAYNLMKEMVEE 459
Score = 54.3 bits (129), Expect = 1e-07
Identities = 47/189 (24%), Positives = 87/189 (45%), Gaps = 4/189 (2%)
Frame = +1
Query: 177 RLVGEMIEKGFPATARTFNILIRT-CGEAGL--AKDLVETFIKSKTFNYRPFKHSYNAIL 233
++ +M G A ++N LI + C + L A +++T +K P ++N+I
Sbjct: 13 QVYNQMKRLGCAADTISYNFLIESHCKDENLDEAVKVLDTMVKK---GVAPNASTFNSIF 183
Query: 234 HSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFS 293
+ L+ +Y +M + LTYNI+M +D L KEM +
Sbjct: 184 GCIAELHDVNGAHRMYAKMKELKCMPNTLTYNILMRMFADSKSIDMVLKLKKEMDESEVE 363
Query: 294 PDFHTYNLLLHFLGKEDKPLAAVNLLKQM-RETGIEPTVLHYTTLIDGLSRAGNLHACKH 352
P+ +TY +L+ ++ A NL+K+M E ++P + Y T+++ L AG L +
Sbjct: 364 PNVNTYRILILMFCEKGHWNNAYNLMKEMVEEKCLKPNLSIYETVLELLRNAGQLKKHEE 543
Query: 353 FFDEMIKNG 361
++M+ G
Sbjct: 544 LVEKMVARG 570
Score = 40.4 bits (93), Expect = 0.002
Identities = 21/81 (25%), Positives = 37/81 (44%)
Frame = +1
Query: 384 KAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLL 443
K +++++M + +YN +I C DEA +L M KG +PN+ + S+
Sbjct: 4 KVLQVYNQMKRLGCAADTISYNFLIESHCKDENLDEAVKVLDTMVKKGVAPNASTFNSIF 183
Query: 444 SCLQNAGKVANAREVMRQMME 464
C+ V A + +M E
Sbjct: 184 GCIAELHDVNGAHRMYAKMKE 246
>TC89807 weakly similar to PIR|B96600|B96600 protein F14J16.14 [imported] -
Arabidopsis thaliana, partial (53%)
Length = 1241
Score = 85.5 bits (210), Expect = 4e-17
Identities = 63/260 (24%), Positives = 118/260 (45%), Gaps = 8/260 (3%)
Frame = +1
Query: 208 KDLVETFIKSKTFN-YRPFKHSYNAILHSLLVLNQYKLIEWV------YQQMLLDGFSSD 260
K LVE F K+ + +R Y + L +++ + + Y + +GFS+
Sbjct: 100 KTLVEKFKKASDIDRFRKKNGIYEDTVRRLAGAKRFRWVRDIIEHQKSYADISNEGFSAR 279
Query: 261 ILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLK 320
++T +Y K + + Q LF EM + + N LL + L K
Sbjct: 280 LIT----LYGKSNMHR--HAQKLFDEMPQRNCERSVLSLNALLAAYLHSKQYDVVERLFK 441
Query: 321 QMR-ETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVA 379
++ + ++P ++ Y T I L G+ + +EM K+G D++ + ++ G
Sbjct: 442 KLPVQLSVKPDLVSYNTYIKALLEKGSFDSAVSVLEEMEKDGVESDLITFNTLLDGLYSK 621
Query: 380 GELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVY 439
G E ++++ ++ K VPN+ TYN+ + GL +A + EA +EME KG P+ +
Sbjct: 622 GRFEDGEKLWEKLGEKNVVPNIRTYNARLLGLAVAKRAGEAVEFYEEMEKKGVKPDLFSF 801
Query: 440 VSLLSCLQNAGKVANAREVM 459
+L+ N G + A+ V+
Sbjct: 802 NALIKGFANEGNLDEAKNVV 861
Score = 57.0 bits (136), Expect = 2e-08
Identities = 44/196 (22%), Positives = 82/196 (41%), Gaps = 1/196 (0%)
Frame = +1
Query: 279 QFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLI 338
+ Q + ++ GFS L+ GK + A L +M + E +VL L+
Sbjct: 229 EHQKSYADISNEGFSAR------LITLYGKSNMHRHAQKLFDEMPQRNCERSVLSLNALL 390
Query: 339 DGLSRAGNLHACKHFFDEM-IKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQ 397
+ + F ++ ++ PD+V+Y I + G + A + EM
Sbjct: 391 AAYLHSKQYDVVERLFKKLPVQLSVKPDLVSYNTYIKALLEKGSFDSAVSVLEEMEKDGV 570
Query: 398 VPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANARE 457
++ T+N+++ GL G+F++ + +++ K PN Y + L L A + A E
Sbjct: 571 ESDLITFNTLLDGLYSKGRFEDGEKLWEKLGEKNVVPNIRTYNARLLGLAVAKRAGEAVE 750
Query: 458 VMRQMMEKGKYAHLLS 473
+M +KG L S
Sbjct: 751 FYEEMEKKGVKPDLFS 798
Score = 33.1 bits (74), Expect = 0.24
Identities = 28/122 (22%), Positives = 51/122 (40%)
Frame = +1
Query: 158 AYHLIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNILIRTCGEAGLAKDLVETFIKS 217
+Y+ + E + + ++ EM + G + TFN L+ G +D + + K
Sbjct: 481 SYNTYIKALLEKGSFDSAVSVLEEMEKDGVESDLITFNTLLDGLYSKGRFEDGEKLWEKL 660
Query: 218 KTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKL 277
N P +YNA L L V + Y++M G D+ ++N ++ G L
Sbjct: 661 GEKNVVPNIRTYNARLLGLAVAKRAGEAVEFYEEMEKKGVKPDLFSFNALIKGFANEGNL 840
Query: 278 DQ 279
D+
Sbjct: 841 DE 846
>BI308910 similar to GP|6682243|gb| hypothetical protein {Arabidopsis
thaliana}, partial (23%)
Length = 591
Score = 82.8 bits (203), Expect = 3e-16
Identities = 52/171 (30%), Positives = 92/171 (53%), Gaps = 2/171 (1%)
Frame = +1
Query: 260 DILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLL 319
D ++Y V+ A + G +D+ + EM R G + TYN+LL K+ + A+ LL
Sbjct: 79 DHVSYTTVVSALVKAGFMDRAHQVLAEMTRIGVPANRITYNILLKGYCKQLQMDKAMELL 258
Query: 320 KQMRET-GIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVV 378
++M E GIEP V+ Y LIDG + FF+EM G P V+YT ++ + +
Sbjct: 259 QEMAEDIGIEPDVVSYNILIDGCILVDDSAGALSFFNEMRAKGIAPTKVSYTTLMKAFAL 438
Query: 379 AGELEKAQEMFHEMISKEQV-PNVYTYNSMIRGLCMAGKFDEACSMLKEME 428
+G+ + AQ +F EM++ +V ++ +N ++ G G +E ++++M+
Sbjct: 439 SGQPKLAQRVFDEMVTDARVRVDLIAWNMLVEGYNRLGLVEEGKKVIQKMK 591
Score = 53.1 bits (126), Expect = 2e-07
Identities = 33/104 (31%), Positives = 52/104 (49%), Gaps = 1/104 (0%)
Frame = +1
Query: 364 PDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSM 423
PD V+YT +++ V AG +++A ++ EM N TYN +++G C + D+A +
Sbjct: 76 PDHVSYTTVVSALVKAGFMDRAHQVLAEMTRIGVPANRITYNILLKGYCKQLQMDKAMEL 255
Query: 424 LKEM-ETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
L+EM E G P+ V Y L+ A A +M KG
Sbjct: 256 LQEMAEDIGIEPDVVSYNILIDGCILVDDSAGALSFFNEMRAKG 387
>TC81548 weakly similar to GP|1931651|gb|AAB65486.1| membrane-associated
salt-inducible protein isolog; 88078-84012 {Arabidopsis
thaliana}, partial (23%)
Length = 882
Score = 82.4 bits (202), Expect = 3e-16
Identities = 58/221 (26%), Positives = 107/221 (48%), Gaps = 7/221 (3%)
Frame = +1
Query: 230 NAILHSLLVLNQY----KLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLDQFQIL-- 283
NA LH N++ +L W+ + LD Y+ Y K+ KLD +L
Sbjct: 241 NATLHHFANSNKFNHVSQLFLWMQENKKLD-------VYSYSHYIKFMANKLDASTMLKL 399
Query: 284 FKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSR 343
+ ++ + + N +L L K+ K + L +QM+ G+ P ++ Y+TLI G +
Sbjct: 400 YNDIQVESAKDNVYVCNSVLSCLIKKGKFDTTMKLFRQMKHDGLVPDLVTYSTLIAGCVK 579
Query: 344 AGNLHA-CKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVY 402
+ + E+ N D V Y ++ G+ E+A+ F++M S+ + PNVY
Sbjct: 580 VKDGYPKALELIQELQDNKLRMDDVIYGAILAVCASNGKWEEAEYYFNQMKSEGRSPNVY 759
Query: 403 TYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLL 443
Y+S++ G F +A +++++ME++G +PN + +LL
Sbjct: 760 HYSSLLNAYSACGDFTKADALIQDMESEGLAPNKGILTTLL 882
Score = 59.7 bits (143), Expect = 2e-09
Identities = 40/173 (23%), Positives = 78/173 (44%), Gaps = 1/173 (0%)
Frame = +1
Query: 295 DFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFF 354
D ++Y+ + F+ + + L ++ + V +++ L + G F
Sbjct: 328 DVYSYSHYIKFMANKLDASTMLKLYNDIQVESAKDNVYVCNSVLSCLIKKGKFDTTMKLF 507
Query: 355 DEMIKNGCMPDVVAYTVMITGYV-VAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCM 413
+M +G +PD+V Y+ +I G V V KA E+ E+ + + Y +++
Sbjct: 508 RQMKHDGLVPDLVTYSTLIAGCVKVKDGYPKALELIQELQDNKLRMDDVIYGAILAVCAS 687
Query: 414 AGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
GK++EA +M+++G SPN Y SLL+ G A +++ M +G
Sbjct: 688 NGKWEEAEYYFNQMKSEGRSPNVYHYSSLLNAYSACGDFTKADALIQDMESEG 846
Score = 35.4 bits (80), Expect = 0.049
Identities = 26/77 (33%), Positives = 40/77 (51%), Gaps = 3/77 (3%)
Frame = +1
Query: 400 NVYTYNSMIRGLCMAGKFDEACSMLK---EMETKGCSPNSVVYVSLLSCLQNAGKVANAR 456
+VY+Y+ I+ MA K D A +MLK +++ + N V S+LSCL GK
Sbjct: 328 DVYSYSHYIK--FMANKLD-ASTMLKLYNDIQVESAKDNVYVCNSVLSCLIKKGKFDTTM 498
Query: 457 EVMRQMMEKGKYAHLLS 473
++ RQM G L++
Sbjct: 499 KLFRQMKHDGLVPDLVT 549
>TC83956 weakly similar to GP|15982931|gb|AAL09812.1 AT3g53700/F4P12_400
{Arabidopsis thaliana}, partial (18%)
Length = 762
Score = 81.6 bits (200), Expect = 6e-16
Identities = 53/209 (25%), Positives = 101/209 (47%), Gaps = 1/209 (0%)
Frame = +1
Query: 209 DLVETFIKS-KTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIV 267
D + T +K K+ P ++ ++ S ++ + + + + L GF D YNI
Sbjct: 139 DSITTLLKQLKSSGSIPNATTFATLIQSFTNFHEIENLLKILENEL--GFKPDTNFYNIA 312
Query: 268 MYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGI 327
+ A KL ++L +M G D T+N+L+ L K + A+ +L++M G+
Sbjct: 313 LNALVEDNKLKLVEMLHSKMVNEGIVLDVSTFNVLIKALCKAHQLRPAILMLEEMANHGL 492
Query: 328 EPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQE 387
+P + +TTL+ G G+L+ +M+ GC+ V+ V++ G+ G +E+A
Sbjct: 493 KPDEITFTTLMQGFIEEGDLNGALKMKKQMLGYGCLLTNVSVKVLVNGFCKEGRVEEALR 672
Query: 388 MFHEMISKEQVPNVYTYNSMIRGLCMAGK 416
E+ + P+ T+NS++ G+C K
Sbjct: 673 FVLEVSEEGFSPDQGTFNSLVNGVCRIWK 759
Score = 73.6 bits (179), Expect = 2e-13
Identities = 42/154 (27%), Positives = 76/154 (49%)
Frame = +1
Query: 291 GFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHAC 350
GF PD + YN+ L+ L +++K L +M GI V + LI L +A L
Sbjct: 277 GFKPDTNFYNIALNALVEDNKLKLVEMLHSKMVNEGIVLDVSTFNVLIKALCKAHQLRPA 456
Query: 351 KHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRG 410
+EM +G PD + +T ++ G++ G+L A +M +M+ + + ++ G
Sbjct: 457 ILMLEEMANHGLKPDEITFTTLMQGFIEEGDLNGALKMKKQMLGYGCLLTNVSVKVLVNG 636
Query: 411 LCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLS 444
C G+ +EA + E+ +G SP+ + SL++
Sbjct: 637 FCKEGRVEEALRFVLEVSEEGFSPDQGTFNSLVN 738
Score = 65.9 bits (159), Expect = 3e-11
Identities = 40/151 (26%), Positives = 77/151 (50%), Gaps = 2/151 (1%)
Frame = +1
Query: 318 LLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKN--GCMPDVVAYTVMITG 375
LLKQ++ +G P + TLI + N H ++ ++++N G PD Y + +
Sbjct: 154 LLKQLKSSGSIPNATTFATLIQSFT---NFHEIENLL-KILENELGFKPDTNFYNIALNA 321
Query: 376 YVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPN 435
V +L+ + + +M+++ V +V T+N +I+ LC A + A ML+EM G P+
Sbjct: 322 LVEDNKLKLVEMLHSKMVNEGIVLDVSTFNVLIKALCKAHQLRPAILMLEEMANHGLKPD 501
Query: 436 SVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
+ + +L+ G + A ++ +QM+ G
Sbjct: 502 EITFTTLMQGFIEEGDLNGALKMKKQMLGYG 594
>BI270314 similar to GP|9759030|dbj contains similarity to salt-inducible
protein~gene_id:K6A12.14 {Arabidopsis thaliana}, partial
(30%)
Length = 669
Score = 80.5 bits (197), Expect = 1e-15
Identities = 52/222 (23%), Positives = 98/222 (43%), Gaps = 1/222 (0%)
Frame = +3
Query: 223 RPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKL-DQFQ 281
+P ++N ++++ Q K++E + +M G + +Y ++ A R K+ D
Sbjct: 3 KPTAVTFNILMYAYSRRMQPKIVESLLAEMKDFGLKPNANSYTCLISAYGRQKKMSDMAA 182
Query: 282 ILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGL 341
F +M + G P H+Y ++H A + + M GI+P++ YTTL+D
Sbjct: 183 DAFLKMKKVGIKPTSHSYTAMIHAYSVSGWHEKAYAVFENMIREGIKPSIETYTTLLDAF 362
Query: 342 SRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNV 401
R G+ + M+ V + +++ G+ G +A+++ E P V
Sbjct: 363 RRVGDTETLMKIWKLMMSEKVKGTQVTFNILVDGFAKQGLFMEARDVISEFGKIGLQPTV 542
Query: 402 YTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLL 443
TYN +I G +LKEME P+S+ Y +++
Sbjct: 543 MTYNMLINAYARGGLDSNIPQLLKEMEALXLRPDSITYSTVI 668
Score = 68.9 bits (167), Expect = 4e-12
Identities = 50/220 (22%), Positives = 90/220 (40%), Gaps = 36/220 (16%)
Frame = +3
Query: 155 TANAYHLIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNILIRTCGEAGLAKDLV-ET 213
TA ++++M Y+ + K + L+ EM + G A ++ LI G D+ +
Sbjct: 9 TAVTFNILMYAYSRRMQPKIVESLLAEMKDFGLKPNANSYTCLISAYGRQKKMSDMAADA 188
Query: 214 FIKSKTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYR 273
F+K K +P HSY A++H+ V ++ V++ M+ +G I TY ++ A R
Sbjct: 189 FLKMKKVGIKPTSHSYTAMIHAYSVSGWHEKAYAVFENMIREGIKPSIETYTTLLDAFRR 368
Query: 274 LGKLDQFQILFK-----------------------------------EMGRNGFSPDFHT 298
+G + ++K E G+ G P T
Sbjct: 369 VGDTETLMKIWKLMMSEKVKGTQVTFNILVDGFAKQGLFMEARDVISEFGKIGLQPTVMT 548
Query: 299 YNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLI 338
YN+L++ + LLK+M + P + Y+T+I
Sbjct: 549 YNMLINAYARGGLDSNIPQLLKEMEALXLRPDSITYSTVI 668
Score = 68.2 bits (165), Expect = 7e-12
Identities = 46/174 (26%), Positives = 79/174 (44%), Gaps = 1/174 (0%)
Frame = +3
Query: 294 PDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLH-ACKH 352
P T+N+L++ + +P +LL +M++ G++P YT LI R +
Sbjct: 6 PTAVTFNILMYAYSRRMQPKIVESLLAEMKDFGLKPNANSYTCLISAYGRQKKMSDMAAD 185
Query: 353 FFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLC 412
F +M K G P +YT MI Y V+G EKA +F MI + P++ TY +++
Sbjct: 186 AFLKMKKVGIKPTSHSYTAMIHAYSVSGWHEKAYAVFENMIREGIKPSIETYTTLLDAFR 365
Query: 413 MAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMMEKG 466
G + + K M ++ V + L+ G AR+V+ + + G
Sbjct: 366 RVGDTETLMKIWKLMMSEKVKGTQVTFNILVDGFAKQGLFMEARDVISEFGKIG 527
Score = 29.6 bits (65), Expect = 2.7
Identities = 25/123 (20%), Positives = 55/123 (44%), Gaps = 3/123 (2%)
Frame = +3
Query: 149 QEAYQHTANAYHLIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNILIRTCGEAGL-- 206
+E + + Y +++ + + + L ++ M+ + T TFNIL+ + GL
Sbjct: 309 REGIKPSIETYTTLLDAFRRVGDTETLMKIWKLMMSEKVKGTQVTFNILVDGFAKQGLFM 488
Query: 207 -AKDLVETFIKSKTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYN 265
A+D++ F K +P +YN ++++ I + ++M D +TY+
Sbjct: 489 EARDVISEFGK---IGLQPTVMTYNMLINAYARGGLDSNIPQLLKEMEALXLRPDSITYS 659
Query: 266 IVM 268
V+
Sbjct: 660 TVI 668
>AJ497575 similar to PIR|T05642|T05 hypothetical protein F20D10.270 -
Arabidopsis thaliana, partial (46%)
Length = 622
Score = 79.0 bits (193), Expect = 4e-15
Identities = 45/153 (29%), Positives = 74/153 (47%)
Frame = +2
Query: 308 KEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVV 367
+E P A + +M+ETG+ P + ++DGL + GN F M + G +PD+V
Sbjct: 170 QESMPQDADAIFNKMKETGLIPNAV---AMLDGLCKDGNFQEALKLFGLMREKGTIPDIV 340
Query: 368 AYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEM 427
YT ++ GY A + + A +F +M S PN ++Y +I+GL K +A EM
Sbjct: 341 IYTAVVDGYTKAHKADDAIRIFRKMQSNSISPNAFSYTVLIQGLYKCSKLHDAVEFCVEM 520
Query: 428 ETKGCSPNSVVYVSLLSCLQNAGKVANAREVMR 460
G S N +V L+ + A+ V++
Sbjct: 521 LEAGHSLNVTTFVGLVDGFCKENGIEEAKGVIK 619
Score = 64.3 bits (155), Expect = 1e-10
Identities = 41/143 (28%), Positives = 65/143 (44%)
Frame = +2
Query: 283 LFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLS 342
+F +M G P+ +L L K+ A+ L MRE G P ++ YT ++DG +
Sbjct: 200 IFNKMKETGLIPNAVA---MLDGLCKDGNFQEALKLFGLMREKGTIPDIVIYTAVVDGYT 370
Query: 343 RAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVY 402
+A F +M N P+ +YTV+I G +L A E EM+ NV
Sbjct: 371 KAHKADDAIRIFRKMQSNSISPNAFSYTVLIQGLYKCSKLHDAVEFCVEMLEAGHSLNVT 550
Query: 403 TYNSMIRGLCMAGKFDEACSMLK 425
T+ ++ G C +EA ++K
Sbjct: 551 TFVGLVDGFCKENGIEEAKGVIK 619
Score = 43.5 bits (101), Expect = 2e-04
Identities = 24/80 (30%), Positives = 40/80 (50%)
Frame = +2
Query: 383 EKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSL 442
+ A +F++M +PN +M+ GLC G F EA + M KG P+ V+Y ++
Sbjct: 185 QDADAIFNKMKETGLIPNAV---AMLDGLCKDGNFQEALKLFGLMREKGTIPDIVIYTAV 355
Query: 443 LSCLQNAGKVANAREVMRQM 462
+ A K +A + R+M
Sbjct: 356 VDGYTKAHKADDAIRIFRKM 415
>AW695801 weakly similar to PIR|T01622|T01 probable salt-inducible protein
At2g18940 [imported] - Arabidopsis thaliana, partial
(18%)
Length = 655
Score = 79.0 bits (193), Expect = 4e-15
Identities = 46/181 (25%), Positives = 86/181 (47%)
Frame = +3
Query: 284 FKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLIDGLSR 343
F ++ NG+ D N +L + K A +L + +G++P ++ Y +LID +R
Sbjct: 27 FHQLQNNGYKLDMVVINSMLSMFVRNQKLEKAHEMLDVIHVSGLQPNLVTYNSLIDLYAR 206
Query: 344 AGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFHEMISKEQVPNVYT 403
G+ + ++ +G PDVV+Y +I G+ G +++A + EM + P T
Sbjct: 207 VGDCWKAEEMLKDIQNSGISPDVVSYNTVIKGFCKKGLVQEAIRILSEMTANGVQPCPIT 386
Query: 404 YNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSLLSCLQNAGKVANAREVMRQMM 463
+N+ + G F EA +++ M GC PN + Y ++ A K A + + ++
Sbjct: 387 FNTFMSCYAGNGLFAEADEVIRYMIEHGCMPNELTYKIVIDGYIKAKKHKEAMDFVSKIK 566
Query: 464 E 464
E
Sbjct: 567 E 569
Score = 77.8 bits (190), Expect = 9e-15
Identities = 42/146 (28%), Positives = 72/146 (48%)
Frame = +3
Query: 321 QMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAG 380
Q++ G + ++ +++ R L D + +G P++V Y +I Y G
Sbjct: 33 QLQNNGYKLDMVVINSMLSMFVRNQKLEKAHEMLDVIHVSGLQPNLVTYNSLIDLYARVG 212
Query: 381 ELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYV 440
+ KA+EM ++ + P+V +YN++I+G C G EA +L EM G P + +
Sbjct: 213 DCWKAEEMLKDIQNSGISPDVVSYNTVIKGFCKKGLVQEAIRILSEMTANGVQPCPITFN 392
Query: 441 SLLSCLQNAGKVANAREVMRQMMEKG 466
+ +SC G A A EV+R M+E G
Sbjct: 393 TFMSCYAGNGLFAEADEVIRYMIEHG 470
Score = 66.6 bits (161), Expect = 2e-11
Identities = 33/134 (24%), Positives = 70/134 (51%)
Frame = +3
Query: 254 LDGFSSDILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPL 313
+ G +++TYN ++ R+G + + + K++ +G SPD +YN ++ K+
Sbjct: 147 VSGLQPNLVTYNSLIDLYARVGDCWKAEEMLKDIQNSGISPDVVSYNTVIKGFCKKGLVQ 326
Query: 314 AAVNLLKQMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMI 373
A+ +L +M G++P + + T + + G MI++GCMP+ + Y ++I
Sbjct: 327 EAIRILSEMTANGVQPCPITFNTFMSCYAGNGLFAEADEVIRYMIEHGCMPNELTYKIVI 506
Query: 374 TGYVVAGELEKAQE 387
GY+ A + ++A +
Sbjct: 507 DGYIKAKKHKEAMD 548
Score = 38.9 bits (89), Expect = 0.004
Identities = 21/86 (24%), Positives = 38/86 (43%)
Frame = +3
Query: 381 ELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYV 440
+L+ + FH++ + ++ NSM+ K ++A ML + G PN V Y
Sbjct: 3 QLKGMERAFHQLQNNGYKLDMVVINSMLSMFVRNQKLEKAHEMLDVIHVSGLQPNLVTYN 182
Query: 441 SLLSCLQNAGKVANAREVMRQMMEKG 466
SL+ G A E+++ + G
Sbjct: 183 SLIDLYARVGDCWKAEEMLKDIQNSG 260
>BF004907 weakly similar to GP|8953393|emb putative protein {Arabidopsis
thaliana}, partial (34%)
Length = 665
Score = 78.2 bits (191), Expect = 7e-15
Identities = 53/206 (25%), Positives = 92/206 (43%)
Frame = +3
Query: 159 YHLIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNILIRTCGEAGLAKDLVETFIKSK 218
Y ++ Y + +VGEM ++G A +N +I EAG K+ + +
Sbjct: 36 YGTLVEGYCRMRRVEKALEMVGEMTKEGIKPNAIVYNPIIDALAEAGRFKEALGMMERFH 215
Query: 219 TFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRLGKLD 278
P +YN+++ + + ++M+ GF TYN R GK+D
Sbjct: 216 VLQIGPTLSTYNSLVKGFCKAGDIEGASKILKKMISRGFLPIPTTYNYFFRYFSRCGKVD 395
Query: 279 QFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPTVLHYTTLI 338
+ L+ +M +G +PD TY+L+L L +E+K AV + +MR G + + T L
Sbjct: 396 EGMNLYTKMIESGHNPDRLTYHLVLKMLCEEEKLELAVQVSMEMRHKGYDMDLATSTMLT 575
Query: 339 DGLSRAGNLHACKHFFDEMIKNGCMP 364
L + L F++MI+ G +P
Sbjct: 576 HLLCKMHKLEEAFAEFEDMIRRGIIP 653
Score = 76.6 bits (187), Expect = 2e-14
Identities = 40/144 (27%), Positives = 65/144 (44%)
Frame = +3
Query: 323 RETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGEL 382
+ + P+V+ Y TL++G R + EM K G P+ + Y +I AG
Sbjct: 3 KNENVRPSVVTYGTLVEGYCRMRRVEKALEMVGEMTKEGIKPNAIVYNPIIDALAEAGRF 182
Query: 383 EKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYVSL 442
++A M + P + TYNS+++G C AG + A +LK+M ++G P Y
Sbjct: 183 KEALGMMERFHVLQIGPTLSTYNSLVKGFCKAGDIEGASKILKKMISRGFLPIPTTYNYF 362
Query: 443 LSCLQNAGKVANAREVMRQMMEKG 466
GKV + +M+E G
Sbjct: 363 FRYFSRCGKVDEGMNLYTKMIESG 434
Score = 75.9 bits (185), Expect = 3e-14
Identities = 44/206 (21%), Positives = 92/206 (44%)
Frame = +3
Query: 261 ILTYNIVMYAKYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLK 320
++TY ++ R+ ++++ + EM + G P+ YN ++ L + + A+ +++
Sbjct: 27 VVTYGTLVEGYCRMRRVEKALEMVGEMTKEGIKPNAIVYNPIIDALAEAGRFKEALGMME 206
Query: 321 QMRETGIEPTVLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAG 380
+ I PT+ Y +L+ G +AG++ +MI G +P Y + G
Sbjct: 207 RFHVLQIGPTLSTYNSLVKGFCKAGDIEGASKILKKMISRGFLPIPTTYNYFFRYFSRCG 386
Query: 381 ELEKAQEMFHEMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSPNSVVYV 440
++++ ++ +MI P+ TY+ +++ LC K + A + EM KG +
Sbjct: 387 KVDEGMNLYTKMIESGHNPDRLTYHLVLKMLCEEEKLELAVQVSMEMRHKGYDMDLATST 566
Query: 441 SLLSCLQNAGKVANAREVMRQMMEKG 466
L L K+ A M+ +G
Sbjct: 567 MLTHLLCKMHKLEEAFAEFEDMIRRG 644
Score = 61.2 bits (147), Expect = 8e-10
Identities = 52/224 (23%), Positives = 93/224 (41%), Gaps = 7/224 (3%)
Frame = +3
Query: 218 KTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYA------- 270
K N RP +Y ++ + + + + +M +G + + YN ++ A
Sbjct: 3 KNENVRPSVVTYGTLVEGYCRMRRVEKALEMVGEMTKEGIKPNAIVYNPIIDALAEAGRF 182
Query: 271 KYRLGKLDQFQILFKEMGRNGFSPDFHTYNLLLHFLGKEDKPLAAVNLLKQMRETGIEPT 330
K LG +++F +L P TYN L+ K A +LK+M G P
Sbjct: 183 KEALGMMERFHVL-------QIGPTLSTYNSLVKGFCKAGDIEGASKILKKMISRGFLPI 341
Query: 331 VLHYTTLIDGLSRAGNLHACKHFFDEMIKNGCMPDVVAYTVMITGYVVAGELEKAQEMFH 390
Y SR G + + + +MI++G PD + Y +++ +LE A ++
Sbjct: 342 PTTYNYFFRYFSRCGKVDEGMNLYTKMIESGHNPDRLTYHLVLKMLCEEEKLELAVQVSM 521
Query: 391 EMISKEQVPNVYTYNSMIRGLCMAGKFDEACSMLKEMETKGCSP 434
EM K ++ T + LC K +EA + ++M +G P
Sbjct: 522 EMRHKGYDMDLATSTMLTHLLCKMHKLEEAFAEFEDMIRRGIIP 653
Score = 49.7 bits (117), Expect = 2e-06
Identities = 29/144 (20%), Positives = 66/144 (45%)
Frame = +3
Query: 155 TANAYHLIMNIYAECEEYKALWRLVGEMIEKGFPATARTFNILIRTCGEAGLAKDLVETF 214
T + Y+ ++ + + + + +++ +MI +GF T+N R G + + +
Sbjct: 234 TLSTYNSLVKGFCKAGDIEGASKILKKMISRGFLPIPTTYNYFFRYFSRCGKVDEGMNLY 413
Query: 215 IKSKTFNYRPFKHSYNAILHSLLVLNQYKLIEWVYQQMLLDGFSSDILTYNIVMYAKYRL 274
K + P + +Y+ +L L + +L V +M G+ D+ T ++ + ++
Sbjct: 414 TKMIESGHNPDRLTYHLVLKMLCEEEKLELAVQVSMEMRHKGYDMDLATSTMLTHLLCKM 593
Query: 275 GKLDQFQILFKEMGRNGFSPDFHT 298
KL++ F++M R G P + T
Sbjct: 594 HKLEEAFAEFEDMIRRGIIPQYLT 665
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.139 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,923,913
Number of Sequences: 36976
Number of extensions: 207902
Number of successful extensions: 1514
Number of sequences better than 10.0: 150
Number of HSP's better than 10.0 without gapping: 1148
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1407
length of query: 480
length of database: 9,014,727
effective HSP length: 100
effective length of query: 380
effective length of database: 5,317,127
effective search space: 2020508260
effective search space used: 2020508260
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0148.13


