

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0148.11
(156 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
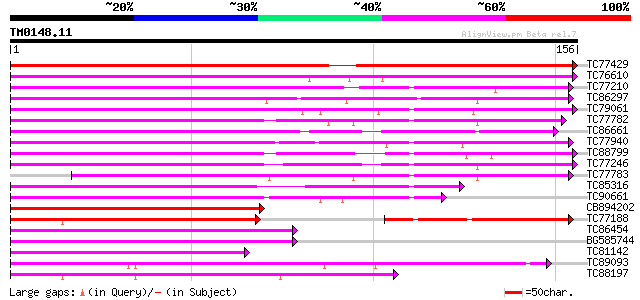
Score E
Sequences producing significant alignments: (bits) Value
TC77429 similar to GP|15215664|gb|AAK91377.1 AT5g25110/T11H3_120... 201 1e-52
TC76610 similar to PIR|T50802|T50802 serine/threonine protein ki... 121 1e-28
TC77210 similar to GP|21954717|gb|AAM83095.1 SOS2-like protein k... 112 6e-26
TC86297 similar to GP|15912283|gb|AAL08275.1 At1g30270/F12P21_6 ... 109 4e-25
TC79061 similar to GP|23428550|gb|AAL23677.1 Serine/threonine Ki... 108 1e-24
TC77782 similar to GP|13448037|gb|AAK26845.1 SOS2-like protein k... 101 1e-22
TC86661 similar to GP|9798599|dbj|BAB11737.1 serine/threonine pr... 96 8e-21
TC77940 similar to PIR|C84667|C84667 probable protein kinase [im... 92 1e-19
TC88799 similar to PIR|C71408|C71408 probable protein kinase - A... 80 3e-16
TC77246 similar to PIR|C71408|C71408 probable protein kinase - A... 80 3e-16
TC77783 similar to GP|19172407|gb|AAL85889.1 putative serine thr... 78 1e-15
TC85316 weakly similar to PIR|E84707|E84707 probable protein kin... 77 3e-15
TC90661 weakly similar to GP|7682805|gb|AAF67384.1| contains sim... 72 7e-14
CB894202 weakly similar to GP|23428550|gb Serine/threonine Kinas... 68 2e-12
TC77188 similar to PIR|T04862|T04862 probable serine/threonine-s... 67 2e-12
TC86454 homologue to GP|4567091|gb|AAD23582.1| SNF-1-like serine... 65 1e-11
BG585744 homologue to GP|4567091|gb| SNF-1-like serine/threonine... 62 7e-11
TC81142 similar to GP|8978255|dbj|BAA98146.1 serine/threonine pr... 60 3e-10
TC89093 similar to GP|15011837|gb|AAK26164.2 calcium-dependent c... 53 6e-08
TC88197 homologue to GP|6739629|gb|AAF27340.1| abscisic acid-act... 52 1e-07
>TC77429 similar to GP|15215664|gb|AAK91377.1 AT5g25110/T11H3_120
{Arabidopsis thaliana}, partial (79%)
Length = 1869
Score = 201 bits (511), Expect = 1e-52
Identities = 104/156 (66%), Positives = 118/156 (74%)
Frame = +3
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LYALLAG LPFQHENL++MYNKV + E+QFPPWFSPESK+LISKILVADP RITISSIM
Sbjct: 840 LYALLAGFLPFQHENLISMYNKVFKEEYQFPPWFSPESKRLISKILVADPERRITISSIM 1019
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
V WFQKG S D+ S +N + + +F NAFEFISSMSSGFDLS
Sbjct: 1020NVPWFQKGLCLSTTTTANDDFELESKVN-------LINSTPQFFNAFEFISSMSSGFDLS 1178
Query: 121 GFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
G FEEK++ GSVFTSKCSVSEI +KIE AAK LRF+
Sbjct: 1179GLFEEKKKRGSVFTSKCSVSEIVTKIESAAKNLRFK 1286
>TC76610 similar to PIR|T50802|T50802 serine/threonine protein kinase-like
protein - Arabidopsis thaliana, partial (73%)
Length = 1799
Score = 121 bits (304), Expect = 1e-28
Identities = 71/169 (42%), Positives = 93/169 (55%), Gaps = 13/169 (7%)
Frame = +2
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ALL G LPFQ EN+M +Y+K +A++ P W SP +KKLIS +LV DP R +I IM
Sbjct: 896 LFALLCGYLPFQGENVMRIYSKSFKADYVLPEWISPGAKKLISNLLVVDPEKRFSIPDIM 1075
Query: 61 RVSWFQKGFSASIPIPDPDES-NFNSDLNSSSE-----QSTKVVAAA-------KFINAF 107
+ WFQ GF I + + + N D + E S +VV NAF
Sbjct: 1076 KDPWFQIGFMRPIAFSMKESAVDDNIDFSGDDEGNGDGNSVEVVGVTGTKHSRRPSYNAF 1255
Query: 108 EFISSMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
E ISS+S GFDL FE ++R S+F SK S S + K+E AK L R
Sbjct: 1256 EIISSLSHGFDLRNLFETRKRSPSMFISKLSASAVVGKLENVAKKLNLR 1402
>TC77210 similar to GP|21954717|gb|AAM83095.1 SOS2-like protein kinase
{Glycine max}, partial (88%)
Length = 1831
Score = 112 bits (280), Expect = 6e-26
Identities = 66/158 (41%), Positives = 94/158 (58%), Gaps = 3/158 (1%)
Frame = +1
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAG LPFQ +NL+ MY K+ R +F+ PPWFS E+++LI+K+L +PN RI+IS IM
Sbjct: 925 LYVLLAGFLPFQDDNLVAMYKKIYRGDFKCPPWFSLEARRLITKLLDPNPNTRISISKIM 1104
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
SWF+K SI + ++ + E ++ +NAF I S+S GFDLS
Sbjct: 1105 DSSWFKKPVPKSIAMRKKEKEEEEDKVFEFME----CEKSSTTMNAFHII-SLSEGFDLS 1269
Query: 121 GFFEEKRRGGSV---FTSKCSVSEIASKIEGAAKGLRF 155
FEEK+R F + + S + SK+E AK ++F
Sbjct: 1270 PLFEEKKREEREEMRFATGGTPSRVISKLEQVAKAVKF 1383
>TC86297 similar to GP|15912283|gb|AAL08275.1 At1g30270/F12P21_6 {Arabidopsis
thaliana}, partial (90%)
Length = 2043
Score = 109 bits (273), Expect = 4e-25
Identities = 67/167 (40%), Positives = 98/167 (58%), Gaps = 12/167 (7%)
Frame = +2
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPF+ +NL+ +Y K+ +A+F PPWFS +KKLI +IL P RI I+ ++
Sbjct: 1022 LFVLMAGYLPFEEDNLVALYKKIFKADFTCPPWFSSSAKKLIKRILDPSPITRIKIAEVI 1201
Query: 61 RVSWFQKGF------SASIPIPDPDESNFNSDLNSSS---EQSTKVVAAAKFINAFEFIS 111
WF+KG+ A + + D + S FN ++S + E+ + A +NAFE IS
Sbjct: 1202 ENEWFKKGYKPPVFEQADVSLDDVN-SIFNGYMDSDNIVVERQEEGPVAPVTMNAFELIS 1378
Query: 112 SMSSGFDLSGFFEEKR---RGGSVFTSKCSVSEIASKIEGAAKGLRF 155
S G +LS FE++ + + FTSKCS +EI SKIE A L F
Sbjct: 1379 K-SQGLNLSSLFEKQMGLVKRETRFTSKCSANEIISKIEKTAGPLGF 1516
>TC79061 similar to GP|23428550|gb|AAL23677.1 Serine/threonine Kinase
{Persea americana}, partial (72%)
Length = 1367
Score = 108 bits (269), Expect = 1e-24
Identities = 70/168 (41%), Positives = 94/168 (55%), Gaps = 12/168 (7%)
Frame = +2
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAG LPF +NLM MY K+ + EF+FP WF+PE ++L+SKIL RI+++ IM
Sbjct: 392 LYVLLAGYLPFHDQNLMEMYRKIGKGEFKFPKWFAPEVRRLLSKILDPSLKTRISMAKIM 571
Query: 61 RVSWFQKGFSASIPIPDPD--------ESNFN-SDLNSSSEQSTKVVAA--AKFINAFEF 109
SWF+KG + I + E F S+ S TK + A +NAF+
Sbjct: 572 ENSWFKKGLEKPVVIETENNELTSLHAEGVFEVSENGGDSNTETKQLQAKPCNNLNAFDI 751
Query: 110 ISSMSSGFDLSGFFEEK-RRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
I S SSGFDLSG FE+ ++ FTS S I SK+E K L+ +
Sbjct: 752 I-SFSSGFDLSGLFEDTIQKKEMRFTSNKPASTIISKLEEICKCLQLK 892
>TC77782 similar to GP|13448037|gb|AAK26845.1 SOS2-like protein kinase PKS6
{Arabidopsis thaliana}, partial (77%)
Length = 1236
Score = 101 bits (251), Expect = 1e-22
Identities = 66/162 (40%), Positives = 91/162 (55%), Gaps = 9/162 (5%)
Frame = +2
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPF NL+ +Y K+ RA+F+ P WFSPE+KKL++ IL +P RI I I+
Sbjct: 758 LFVLMAGYLPFDEPNLIALYRKIGRADFKCPSWFSPEAKKLLTSILNPNPLTRIKIPEIL 937
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSD-----LNSSSEQ-STKVVAAAKFINAFEFISSMS 114
WF+KG+ P +E + N D N S E T+ +NAFE I S S
Sbjct: 938 EDEWFRKGYK---PACFTEEEDVNVDDVAAAFNDSRENLVTETKEKPVSMNAFELI-SRS 1105
Query: 115 SGFDLSGFFEEKR---RGGSVFTSKCSVSEIASKIEGAAKGL 153
F+L FE ++ + + FTS+ +EI SKIE AAK L
Sbjct: 1106ESFNLDNLFEREKGVVKRETHFTSQRPANEIMSKIEEAAKPL 1231
>TC86661 similar to GP|9798599|dbj|BAB11737.1 serine/threonine protein kinase
{Arabidopsis thaliana}, partial (58%)
Length = 1840
Score = 95.5 bits (236), Expect = 8e-21
Identities = 55/151 (36%), Positives = 81/151 (53%)
Frame = +2
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPF NLM MY K+ EF+ P WFSP+ K+ +S+++ +P RIT+ I+
Sbjct: 878 LFVLVAGYLPFNDPNLMVMYRKIYSGEFKCPRWFSPDLKRFLSRLMDTNPETRITVDEIL 1057
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
R WF+KG+ + E F +N+ E+ + NAF+ IS S G +LS
Sbjct: 1058 RDPWFRKGYKEVKFYEEGFE--FEKKVNNGEEEGKPL-----DFNAFDIISFFSRGLNLS 1216
Query: 121 GFFEEKRRGGSVFTSKCSVSEIASKIEGAAK 151
GFF + G F + ++ K E AAK
Sbjct: 1217 GFFIDGGE-GERFMLRELPEKVVEKAEAAAK 1306
>TC77940 similar to PIR|C84667|C84667 probable protein kinase [imported] -
Arabidopsis thaliana, partial (64%)
Length = 1258
Score = 91.7 bits (226), Expect = 1e-19
Identities = 61/163 (37%), Positives = 91/163 (55%), Gaps = 8/163 (4%)
Frame = +2
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPF NLM +Y K+ A+F PPW S ++KLI++IL +P RIT++ I+
Sbjct: 182 LFVLVAGYLPFDDPNLMELYKKISSADFTCPPWLSFSARKLITRILDPNPMTRITMAEIL 361
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAK------FINAFEFISSMS 114
WF+K + + + E+N + D+ + + S + K +NAFE I SMS
Sbjct: 362 DDEWFKKDYKPPV-FEESGETNLD-DVEAVFKDSEEHHVTEKKEEQPTSMNAFELI-SMS 532
Query: 115 SGFDLSGFF--EEKRRGGSVFTSKCSVSEIASKIEGAAKGLRF 155
G +L F E+ + + FTS+ EI +KIE AAK L F
Sbjct: 533 RGLNLENLFDVEQGFKRETRFTSQSPADEIINKIEEAAKPLGF 661
>TC88799 similar to PIR|C71408|C71408 probable protein kinase - Arabidopsis
thaliana, partial (57%)
Length = 1648
Score = 80.5 bits (197), Expect = 3e-16
Identities = 52/166 (31%), Positives = 81/166 (48%), Gaps = 10/166 (6%)
Frame = +1
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAG LPF N+ M K+ R +++FP W S ++ +I ++L +P R+++ +
Sbjct: 706 LYVLLAGYLPFDDLNIPAMCKKISRRDYRFPDWVSKRARIVIYRLLDPNPETRMSLEGLF 885
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
WF+K P +P++S F D + +NAF I SMSSG DLS
Sbjct: 886 GNEWFKKSLK---PEAEPEKSIFGCDYGKEGKNLG--------LNAFHII-SMSSGLDLS 1029
Query: 121 GFFE--EKRRGGS--------VFTSKCSVSEIASKIEGAAKGLRFR 156
G FE +GG+ FTS ++ + K++ L F+
Sbjct: 1030GLFETTSSSQGGNNNNTWREKRFTSSANIEVVGEKVKEVGGVLGFK 1167
>TC77246 similar to PIR|C71408|C71408 probable protein kinase - Arabidopsis
thaliana, partial (65%)
Length = 1715
Score = 80.1 bits (196), Expect = 3e-16
Identities = 51/157 (32%), Positives = 80/157 (50%), Gaps = 1/157 (0%)
Frame = +3
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAG LPF N+ M ++ R ++QFP W S ++ LI ++L +P RI + ++
Sbjct: 660 LYVLLAGHLPFDDSNIPAMCKRITRRDYQFPEWISKPARYLIYQLLDPNPKTRIRLENVF 839
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
+WF+K +P+E F+ + + ++ +NAF+ I S+SSG +LS
Sbjct: 840 GNNWFKKSLR-----EEPEEKLFDDKYSFDGYKKLEL-----GMNAFDII-SLSSGLNLS 986
Query: 121 GFFEEKR-RGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
G FE R FTS V + K++ L FR
Sbjct: 987 GLFETTTLRSEKRFTSSEEVGVVEEKVKEIGVRLGFR 1097
>TC77783 similar to GP|19172407|gb|AAL85889.1 putative serine threonine
kinase {Sandersonia aurantiaca}, partial (85%)
Length = 990
Score = 78.2 bits (191), Expect = 1e-15
Identities = 53/143 (37%), Positives = 78/143 (54%), Gaps = 5/143 (3%)
Frame = +1
Query: 18 TMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIMRVSWFQKGFS-ASIPIP 76
T +++ +F+FP WFSP +KKL+ IL +P RI I I++ WF+KG+ A
Sbjct: 34 TCIEELVEPDFKFPSWFSPGAKKLLRSILNPNPITRIKIPEILQHEWFRKGYKPAFFTEE 213
Query: 77 DPDESNFNSDLNSSSEQ-STKVVAAAKFINAFEFISSMSSGFDLSGFFEEKR---RGGSV 132
D + + + N S E T+ +NAFE I S S F+L FE+++ + +
Sbjct: 214 DVNVDDVAAAFNDSKENLVTETKEKPVSMNAFELI-SRSDSFNLDNLFEKEKGVVKRETH 390
Query: 133 FTSKCSVSEIASKIEGAAKGLRF 155
FTS+ +EI SKIE AAK L F
Sbjct: 391 FTSQRPANEIMSKIEEAAKPLGF 459
>TC85316 weakly similar to PIR|E84707|E84707 probable protein kinase
[imported] - Arabidopsis thaliana, partial (66%)
Length = 1528
Score = 77.0 bits (188), Expect = 3e-15
Identities = 42/125 (33%), Positives = 63/125 (49%)
Frame = +1
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ + AG LPF N+ +Y K+ R +F+FP W S + K L+S++L +P RI++ I+
Sbjct: 838 LFTVTAGYLPFNDYNVTVLYRKIYRGQFRFPKWMSCDLKNLLSRMLDTNPKTRISVDEIL 1017
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
WF G + D Q + K +NAF+ I S S+G D+S
Sbjct: 1018EDPWFSSG-------------GYKLDRVLVKSQMEESRTGFKSLNAFDLI-SFSTGLDMS 1155
Query: 121 GFFEE 125
G FEE
Sbjct: 1156GMFEE 1170
>TC90661 weakly similar to GP|7682805|gb|AAF67384.1| contains similarity to
Pfam family PF00069 (Eukaryotic protein kinase domain)
score=310.0 E=2., partial (36%)
Length = 677
Score = 72.4 bits (176), Expect = 7e-14
Identities = 45/128 (35%), Positives = 68/128 (52%), Gaps = 8/128 (6%)
Frame = +2
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY L+AG LPF+ +L T++ ++ EF P WFS +K I KIL DP R+ I I
Sbjct: 296 LYVLMAGYLPFEEADLPTLFRRISAGEFVCPVWFSAGAKTFIHKILDPDPKTRVKIVEIR 475
Query: 61 RVSWFQKGFSASIPIPDPDESNFN------SDLNSS--SEQSTKVVAAAKFINAFEFISS 112
+ WF+K +S I + + ++ N + D+ SE+S +NAFE I +
Sbjct: 476 KDPWFRKNYS-PIKLREDEQVNLDDIKAVFDDIEDQYVSERSEIAEGGPLIMNAFEMI-T 649
Query: 113 MSSGFDLS 120
+S G +LS
Sbjct: 650 LSQGLNLS 673
>CB894202 weakly similar to GP|23428550|gb Serine/threonine Kinase {Persea
americana}, partial (24%)
Length = 827
Score = 67.8 bits (164), Expect = 2e-12
Identities = 31/70 (44%), Positives = 45/70 (64%)
Frame = +2
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY L+ PF NLM MY K + E++ P WFS E ++L+S+IL +P+ RI+ + IM
Sbjct: 149 LYVLMTRYYPFYDRNLMEMYRKSNKGEYKCPDWFSVEIRRLLSQILNPNPDSRISTAKIM 328
Query: 61 RVSWFQKGFS 70
WF+KGF+
Sbjct: 329 ESRWFRKGFT 358
>TC77188 similar to PIR|T04862|T04862 probable serine/threonine-specific
protein kinase (EC 2.7.1.-) F28A21.110 - Arabidopsis
thaliana, partial (79%)
Length = 1881
Score = 67.4 bits (163), Expect = 2e-12
Identities = 32/70 (45%), Positives = 44/70 (62%), Gaps = 1/70 (1%)
Frame = +1
Query: 1 LYALLAGCLPFQH-ENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSI 59
L+ L+AG LPF N+M MY K+ + EF+ P WFSPE L++++L P RI+I I
Sbjct: 760 LFVLMAGYLPFHDPNNVMAMYKKIYKGEFRCPRWFSPELVSLLTRLLDIKPQTRISIPEI 939
Query: 60 MRVSWFQKGF 69
M WF+ GF
Sbjct: 940 MENRWFKIGF 969
Score = 42.7 bits (99), Expect = 6e-05
Identities = 26/52 (50%), Positives = 33/52 (63%)
Frame = +2
Query: 104 INAFEFISSMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRF 155
+NAF+ IS S GFDLSG FEEK + F S SV +I SK+E + +RF
Sbjct: 1175 LNAFDIIS-FSQGFDLSGLFEEK-GDEARFVSSASVPKIISKLEEVGQMVRF 1324
>TC86454 homologue to GP|4567091|gb|AAD23582.1| SNF-1-like serine/threonine
protein kinase {Glycine max}, partial (90%)
Length = 2194
Score = 65.1 bits (157), Expect = 1e-11
Identities = 31/79 (39%), Positives = 44/79 (55%)
Frame = +3
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LYALL G LPF EN+ ++ K+ + P SP ++ LI ++LV DP R+TI I
Sbjct: 678 LYALLCGTLPFDDENIPNLFKKIKGGIYTLPSHLSPGARDLIPRLLVVDPMKRMTIPEIR 857
Query: 61 RVSWFQKGFSASIPIPDPD 79
+ WFQ + +P PD
Sbjct: 858 QHPWFQLHLPRYLAVPPPD 914
>BG585744 homologue to GP|4567091|gb| SNF-1-like serine/threonine protein
kinase {Glycine max}, partial (36%)
Length = 597
Score = 62.4 bits (150), Expect = 7e-11
Identities = 30/79 (37%), Positives = 43/79 (53%)
Frame = +2
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LYALL G LPF EN+ ++ K+ + P SP ++ LI ++LV DP R+TI I
Sbjct: 131 LYALLCGTLPFDDENIPNLFKKIKGGIYTLPSHLSPGARDLIPRLLVVDPMKRMTIPEIR 310
Query: 61 RVSWFQKGFSASIPIPDPD 79
+ WF + +P PD
Sbjct: 311 QHPWFLLHLPRYLAVPPPD 367
>TC81142 similar to GP|8978255|dbj|BAA98146.1 serine/threonine protein
kinase SOS2 {Arabidopsis thaliana}, partial (54%)
Length = 784
Score = 60.5 bits (145), Expect = 3e-10
Identities = 28/66 (42%), Positives = 39/66 (58%)
Frame = +2
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY L+AG LPF+ +L T++ ++ EF P WFS +K I KIL DP R+ I I
Sbjct: 587 LYVLMAGYLPFEEADLPTLFRRISAGEFVCPVWFSAGAKTFIHKILDPDPKTRVKIVEIR 766
Query: 61 RVSWFQ 66
+ WF+
Sbjct: 767 KDPWFR 784
>TC89093 similar to GP|15011837|gb|AAK26164.2 calcium-dependent
calmodulin-independent protein kinase 5 {Cucumis
sativus}, partial (42%)
Length = 690
Score = 52.8 bits (125), Expect = 6e-08
Identities = 46/159 (28%), Positives = 75/159 (46%), Gaps = 10/159 (6%)
Frame = +1
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFP--PW--FSPESKKLISKILVADPNLRITI 56
LY LL+G PF EN +++ +LR F PW S +K LI K+L ADP RI+
Sbjct: 193 LYILLSGVPPFWAENEQGIFDAILRGHLDFASDPWPKISSIAKDLIKKMLRADPKERISA 372
Query: 57 SSIMRVSWFQKGFSASIPIPDPDESNFNS--DLNSSSEQSTKVVA----AAKFINAFEFI 110
++ SW ++ ++ P+ + +N + + KV+A + I E
Sbjct: 373 VEVLDHSWMKEDGASDKPLDIAVLTRMKQFRAMNKLKKVALKVIAENLSEEEIIGLKEMF 552
Query: 111 SSMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGA 149
SM + + FEE + G +K S SE+ +I+G+
Sbjct: 553 KSMDTDNSGTITFEELKAGLPKLGTKISESEV-RQIDGS 666
>TC88197 homologue to GP|6739629|gb|AAF27340.1| abscisic acid-activated
protein kinase {Vicia faba}, partial (96%)
Length = 1158
Score = 51.6 bits (122), Expect = 1e-07
Identities = 34/121 (28%), Positives = 55/121 (45%), Gaps = 14/121 (11%)
Frame = +1
Query: 1 LYALLAGCLPFQH----ENLMTMYNKVLRAEFQFPPW--FSPESKKLISKILVADPNLRI 54
LY +L G PF+ ++ +VL ++ P + SPE +++IS+I V DP RI
Sbjct: 685 LYVMLVGSYPFEDPDNPKDFRKTIQRVLSVQYSVPDFVQISPECREVISRIFVFDPAKRI 864
Query: 55 TISSIMRVSWFQKGFSASI--------PIPDPDESNFNSDLNSSSEQSTKVVAAAKFINA 106
TI I++ WF K A + +PD+ + D V AA +++
Sbjct: 865 TIPEILKNEWFLKNLPADLVNEKLTDNQFEEPDQPMQSMDTIMQIISEATVPAAGSYLDQ 1044
Query: 107 F 107
F
Sbjct: 1045F 1047
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.133 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,479,359
Number of Sequences: 36976
Number of extensions: 53747
Number of successful extensions: 383
Number of sequences better than 10.0: 93
Number of HSP's better than 10.0 without gapping: 370
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 371
length of query: 156
length of database: 9,014,727
effective HSP length: 88
effective length of query: 68
effective length of database: 5,760,839
effective search space: 391737052
effective search space used: 391737052
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0148.11


