

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
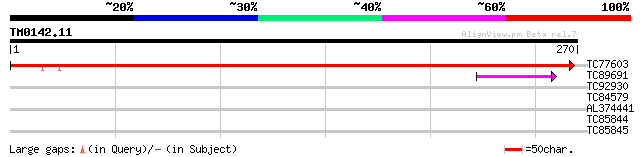
Query= TM0142.11
(270 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC77603 similar to GP|11994356|dbj|BAB02315. gene_id:MSJ11.24~un... 439 e-124
TC89691 similar to GP|8918359|dbj|BAA97583.1 RuBisCO activase la... 41 6e-04
TC92930 similar to GP|13430332|gb|AAK25798.1 rubisco activase {Z... 40 0.001
TC84579 28 4.8
AL374441 27 6.3
TC85844 homologue to SP|P12886|ADH1_PEA Alcohol dehydrogenase 1 ... 27 8.3
TC85845 homologue to SP|P12886|ADH1_PEA Alcohol dehydrogenase 1 ... 27 8.3
>TC77603 similar to GP|11994356|dbj|BAB02315. gene_id:MSJ11.24~unknown
protein {Arabidopsis thaliana}, partial (79%)
Length = 1141
Score = 439 bits (1129), Expect = e-124
Identities = 212/272 (77%), Positives = 238/272 (86%), Gaps = 3/272 (1%)
Frame = +2
Query: 1 MYDSSDLLFFSVPP-SNPITSRH--TFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPA 57
++ SS FSV S+ IT+ FLNVSLPRCY +KER+VKV +NAA AVAT+P
Sbjct: 143 IFASSTQPCFSVTTRSHSITNTFGSNFLNVSLPRCYPMKERHVKVRHVVNAAAAVATSPT 322
Query: 58 QEIEEYKIPSWANFELGKASVYWKTMNGLPPTSGEKLKLFYNPTSTQLAPNEEFGIAFNG 117
+EI+EYK+PSWA FELGKA+VYWKT NG+ PTSGEKLKLFYNP + QLAPNEEFGIAFNG
Sbjct: 323 EEIQEYKLPSWAMFELGKAAVYWKTTNGVAPTSGEKLKLFYNPAAAQLAPNEEFGIAFNG 502
Query: 118 GFNQPIMCGGEPRAMLRKDRGKADAPIYSIQICVPKHALNLIFSFTNGVDWDGPYRLQFQ 177
GFNQPIMCGGEPRAMLRKDRGKAD+PIYSIQICVPKHALNLIFSFTNGVDWDGPYRLQFQ
Sbjct: 503 GFNQPIMCGGEPRAMLRKDRGKADSPIYSIQICVPKHALNLIFSFTNGVDWDGPYRLQFQ 682
Query: 178 VPKVLQNKPIEFFNEGLAEELGKEGACEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNL 237
VPK LQNKPIEFFNEGLAEEL KEGACEQAIFPD+ VI KCAM+GNL+ EGGDRC+LNL
Sbjct: 683 VPKPLQNKPIEFFNEGLAEELSKEGACEQAIFPDTTAVIEKCAMIGNLSKEGGDRCELNL 862
Query: 238 VEGCTDPSSHLYNPLANVDDGTCPLDLDSDSE 269
V GC DPSS LY+P+ANVDDG+CP+++DSDS+
Sbjct: 863 VPGCIDPSSPLYDPMANVDDGSCPIEVDSDSD 958
>TC89691 similar to GP|8918359|dbj|BAA97583.1 RuBisCO activase large isoform
precursor {Oryza sativa (japonica cultivar-group)},
partial (59%)
Length = 1079
Score = 40.8 bits (94), Expect = 6e-04
Identities = 17/38 (44%), Positives = 23/38 (59%)
Frame = +1
Query: 223 GNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
GN +G + L + EGC DPS+ Y+P A DDG+C
Sbjct: 727 GNFYGQGAQQVPLPVQEGCADPSAENYDPTARSDDGSC 840
>TC92930 similar to GP|13430332|gb|AAK25798.1 rubisco activase {Zantedeschia
aethiopica}, partial (42%)
Length = 765
Score = 40.0 bits (92), Expect = 0.001
Identities = 26/91 (28%), Positives = 44/91 (47%), Gaps = 6/91 (6%)
Frame = +2
Query: 176 FQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNKVITKCAMLGNLTVEG 229
F PK+ K +E+ N + E+ + E A D+N+ K G+ +
Sbjct: 284 FDQPKMSLEKLLEYGNMLVQEQENVKRVQLADKYLEGAALGDANQDAIKS---GSFYGKA 454
Query: 230 GDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ ++ + EGCTDP++ ++P A DDGTC
Sbjct: 455 AQQVNIPIPEGCTDPNAKNFDPTARSDDGTC 547
>TC84579
Length = 698
Score = 27.7 bits (60), Expect = 4.8
Identities = 15/48 (31%), Positives = 22/48 (45%)
Frame = +1
Query: 169 DGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGACEQAIFPDSNKVI 216
+G + + LQNK EF A LGK GA + PD++ +
Sbjct: 283 EGSMQKVYAANSPLQNKQEEFQPSESAAALGKSGAANSSEMPDASDAL 426
>AL374441
Length = 357
Score = 27.3 bits (59), Expect = 6.3
Identities = 11/18 (61%), Positives = 14/18 (77%)
Frame = +3
Query: 251 PLANVDDGTCPLDLDSDS 268
PLAN+D TCP+ LD +S
Sbjct: 48 PLANLDIWTCPIKLDCNS 101
>TC85844 homologue to SP|P12886|ADH1_PEA Alcohol dehydrogenase 1 (EC
1.1.1.1). [Garden pea] {Pisum sativum}, complete
Length = 1502
Score = 26.9 bits (58), Expect = 8.3
Identities = 21/94 (22%), Positives = 38/94 (40%), Gaps = 7/94 (7%)
Frame = +2
Query: 97 FYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAMLRKDRGKADAPIYSIQICVPK--- 153
F NP + NGG ++ + C G +AM+ D ++ + VP
Sbjct: 818 FVNPKDHDKPVQQVIAEMTNGGVDRAVECTGSIQAMISAFECVHDGWGVAVLVGVPNKDD 997
Query: 154 ----HALNLIFSFTNGVDWDGPYRLQFQVPKVLQ 183
H +NL+ T + G Y+ + +P V++
Sbjct: 998 AFKTHPMNLLNERTLKGTFYGNYKPRTDLPNVVE 1099
>TC85845 homologue to SP|P12886|ADH1_PEA Alcohol dehydrogenase 1 (EC
1.1.1.1). [Garden pea] {Pisum sativum}, partial (45%)
Length = 734
Score = 26.9 bits (58), Expect = 8.3
Identities = 21/94 (22%), Positives = 38/94 (40%), Gaps = 7/94 (7%)
Frame = +1
Query: 97 FYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAMLRKDRGKADAPIYSIQICVPK--- 153
F NP + NGG ++ + C G +AM+ D ++ + VP
Sbjct: 112 FVNPKDHDKPVQQVIAEMTNGGVDRAVECTGSIQAMISAFECVHDGWGVAVLVGVPNKDD 291
Query: 154 ----HALNLIFSFTNGVDWDGPYRLQFQVPKVLQ 183
H +NL+ T + G Y+ + +P V++
Sbjct: 292 AFQTHPMNLLNERTLKGTFYGNYKPRTDLPNVVE 393
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,643,235
Number of Sequences: 36976
Number of extensions: 115336
Number of successful extensions: 499
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 497
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 499
length of query: 270
length of database: 9,014,727
effective HSP length: 95
effective length of query: 175
effective length of database: 5,502,007
effective search space: 962851225
effective search space used: 962851225
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0142.11


