

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
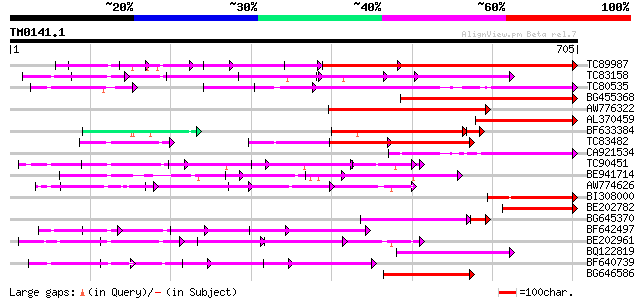
Query= TM0141.1
(705 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC89987 weakly similar to PIR|D86443|D86443 probable PPR-repeat ... 276 3e-91
TC83158 weakly similar to PIR|H86191|H86191 hypothetical protein... 212 3e-55
TC80535 weakly similar to GP|14335136|gb|AAK59848.1 At2g15690/F9... 211 6e-55
BG455368 weakly similar to GP|9759524|dbj selenium-binding prote... 191 6e-49
AW776322 weakly similar to PIR|H84859|H84 hypothetical protein A... 170 2e-42
AL370459 weakly similar to GP|20146256|d hypothetical protein {O... 157 1e-38
BF633384 similar to PIR|T46179|T46 hypothetical protein T8H10.30... 145 5e-38
TC83482 weakly similar to GP|9758453|dbj|BAB08982.1 selenium-bin... 152 4e-37
CA921534 similar to PIR|D96516|D96 F16N3.14 [imported] - Arabido... 150 2e-36
TC90451 similar to PIR|H84508|H84508 hypothetical protein At2g13... 102 2e-33
BE941714 weakly similar to GP|11994734|dbj selenium-binding prot... 133 2e-31
AW774626 similar to PIR|T47909|T47 hypothetical protein T20K12.7... 123 3e-28
BI308000 similar to GP|7363286|dbj| Similar to Arabidopsis thali... 115 6e-26
BE202782 weakly similar to GP|22093801|dbj hypothetical protein~... 112 4e-25
BG645370 weakly similar to PIR|T00405|T004 hypothetical protein ... 100 2e-24
BF642497 weakly similar to PIR|H96713|H96 hypothetical protein T... 108 5e-24
BE202961 weakly similar to GP|6016735|gb|A hypothetical protein ... 108 9e-24
BQ122819 similar to PIR|C84748|C84 hypothetical protein At2g3368... 104 1e-22
BF640739 weakly similar to GP|7363286|dbj Similar to Arabidopsis... 100 2e-21
BG646586 weakly similar to GP|10177683|dbj selenium-binding prot... 99 4e-21
>TC89987 weakly similar to PIR|D86443|D86443 probable PPR-repeat protein
[imported] - Arabidopsis thaliana, partial (62%)
Length = 1559
Score = 276 bits (706), Expect(2) = 3e-91
Identities = 136/316 (43%), Positives = 204/316 (64%)
Frame = +2
Query: 390 VFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLAL 449
VF + ++ ++ MI G + HG G+EAL+VF +M G +P++V +VGV SAC+H L
Sbjct: 581 VFKNMSEKNRYSYTVMISGLAIHGRGKEALKVFSEMIEEGLAPDDVVYVGVFSACSHAGL 760
Query: 450 VDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLL 509
V++GL + HKIEP ++HY CMV L R G+L++A +K+ +K + V WR+LL
Sbjct: 761 VEEGLQCFKSMQFEHKIEPTVQHYGCMVDLLGRFGMLKEAYELIKSMSIKPNDVIWRSLL 940
Query: 510 NSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKK 569
++C+VH N ++GK AE++ ++ + G Y +L+NM+A A WD V +IR + ERN+ +
Sbjct: 941 SACKVHHNLEIGKIAAENLFMLNQNNSGDYLVLANMYAKAQKWDDVAKIRTKLAERNLVQ 1120
Query: 570 EPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQ 629
PG S +E + + FVS+ P+ N I + + Q+ +K GY+P+ S VL DV+DE+
Sbjct: 1121TPGFSLIEAKRKVYKFVSQDKSIPQWNIIYEMIHQMEWQVKFEGYIPDTSQVLLDVDDEE 1300
Query: 630 KEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDAN 689
K+ L HS+KLAIA+GL+ P+RI +NLRMC DCHT K IS + R I VRD
Sbjct: 1301KKERLKFHSQKLAIAFGLIHTSEGXPLRITRNLRMCSDCHTYTKYISMIYEREITVRDRX 1480
Query: 690 RFHHFRDGSCTCADHW 705
RF HF++GSC+C D+W
Sbjct: 1481RFXHFKNGSCSCKDYW 1528
Score = 80.9 bits (198), Expect = 2e-15
Identities = 50/145 (34%), Positives = 76/145 (51%), Gaps = 2/145 (1%)
Frame = +1
Query: 137 SCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDEND 196
+CS G V EG+Q HG++FK GL V+++L NMY +C + A V N G DE
Sbjct: 130 ACSLLGVVDEGIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACDVFN---GMDEKS 300
Query: 197 IFLYNSLLNVLVESGRWEEAIGVLRRMGNE--CMVWDNATYVGVMGLCAQIGDLQLGLRV 254
+ +++++ W E + +L +M +E C V + +T V V+ C +G LG +
Sbjct: 301 VASWSAIIGAHACVEMWNECLMLLGKMSSEGRCRV-EESTLVNVLSACTHLGSPDLGKCI 477
Query: 255 HARLLRGGLLFDEFVGSKLIDMYGK 279
H LLR + V + LIDMY K
Sbjct: 478 HGILLRNISELNVVVKTSLIDMYVK 552
Score = 78.2 bits (191), Expect(2) = 3e-91
Identities = 47/148 (31%), Positives = 78/148 (51%), Gaps = 1/148 (0%)
Frame = +1
Query: 242 CAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVW 301
C+ +G + G++VH + + GL D V + LI+MYGKC + NA VF+G+ ++V W
Sbjct: 133 CSLLGVVDEGIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACDVFNGMDEKSVASW 312
Query: 302 TSLMSAYLQNRYFEETLDLLTCMDQEDTL-PNECTFKVLLGACAGIALLKHGDLLHARLE 360
++++ A+ + E L LL M E E T +L AC + G +H L
Sbjct: 313 SAIIGAHACVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSACTHLGSPDLGKCIHGILL 492
Query: 361 KLGFKNYVIVNNALINMYSKSGDIDSSY 388
+ + V+V +LI+MY KSG ++ +
Sbjct: 493 RNISELNVVVKTSLIDMYVKSGCLEKRF 576
Score = 74.3 bits (181), Expect = 1e-13
Identities = 46/148 (31%), Positives = 79/148 (53%), Gaps = 1/148 (0%)
Frame = +1
Query: 342 ACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTT 401
AC+ + ++ G +H + K+G + VIV N+LINMY K G+I ++ +VF+ + V +
Sbjct: 130 ACSLLGVVDEGIQVHGHVFKMGLEGDVIVQNSLINMYGKCGEIKNACDVFNGMDEKSVAS 309
Query: 402 WNSMICGYSHHGLGQEALRVFQDMKSAGE-SPNNVTFVGVLSACAHLALVDKGLDYLYNE 460
W+++I ++ + E L + M S G T V VLSAC HL D G ++
Sbjct: 310 WSAIIGAHACVEMWNECLMLLGKMSSEGRCRVEESTLVNVLSACTHLGSPDLG-KCIHGI 486
Query: 461 MKNHKIEPGLEHYTCMVVLYCRAGLLEK 488
+ + E + T ++ +Y ++G LEK
Sbjct: 487 LLRNISELNVVVKTSLIDMYVKSGCLEK 570
Score = 56.2 bits (134), Expect = 4e-08
Identities = 36/119 (30%), Positives = 59/119 (49%)
Frame = +1
Query: 57 DVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLV 116
DV+ NSLI++Y KCG ++ A +F+ M + V SW+ ++ + + E L+L +
Sbjct: 205 DVIVQNSLINMYGKCGEIKNACDVFNGMDEKSVASWSAIIGAHACVEMWNECLMLLGKMS 384
Query: 117 SSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYAR 175
S + E VLS+C+H G G HG L + VK++L +MY +
Sbjct: 385 SEGRCR---VEESTLVNVLSACTHLGSPDLGKCIHGILLRNISELNVVVKTSLIDMYVK 552
Score = 45.8 bits (107), Expect = 6e-05
Identities = 42/195 (21%), Positives = 82/195 (41%), Gaps = 39/195 (20%)
Frame = +2
Query: 74 LRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTT 133
LR +F M ++ +S+ +++G G E L +F ++ P++ V+
Sbjct: 563 LRKGLRVFKNMSEKNRYSYTVMISGLAIHGRGKEALKVFSEMIEEGLA----PDDVVYVG 730
Query: 134 VLSSCSHSGRVREGMQCH--------------------GYLFKYGLV--SYQFVKSALF- 170
V S+CSH+G V EG+QC L ++G++ +Y+ +KS
Sbjct: 731 VFSACSHAGLVEEGLQCFKSMQFEHKIEPTVQHYGCMVDLLGRFGMLKEAYELIKSMSIK 910
Query: 171 -------NMYARC-LHVNLA--------LQVLNTESGGDENDIFLYNSLLNVLVESGRWE 214
++ + C +H NL L +LN + GD Y L N+ ++ +W+
Sbjct: 911 PNDVIWRSLLSACKVHHNLEIGKIAAENLFMLNQNNSGD------YLVLANMYAKAQKWD 1072
Query: 215 EAIGVLRRMGNECMV 229
+ + ++ +V
Sbjct: 1073DVAKIRTKLAERNLV 1117
>TC83158 weakly similar to PIR|H86191|H86191 hypothetical protein [imported]
- Arabidopsis thaliana, partial (42%)
Length = 1194
Score = 212 bits (540), Expect = 3e-55
Identities = 115/343 (33%), Positives = 194/343 (56%)
Frame = +2
Query: 285 NARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACA 344
+A K+FD L +NVV WT ++ +++ +EE L+ M +P+ T ++ ACA
Sbjct: 14 DALKLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREMQLAGVVPDFVTVIAIISACA 193
Query: 345 GIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNS 404
+ L G +H + K F++ V V N+LI+MY++ G I+ + VF R++ +WNS
Sbjct: 194 NLGALGLGLWVHRLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFDGMSQRNLVSWNS 373
Query: 405 MICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNH 464
+I G++ +GL +AL F+ MK G PN V++ L+AC+H L+D+GL + ++H
Sbjct: 374 IIVGFAVNGLADKALSFFRSMKKEGLEPNGVSYTSALTACSHAGLIDEGLKIFADIKRDH 553
Query: 465 KIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRI 524
+ P +EHY C+V LY RAG L++A + +K + + V +LL +CR + +L +++
Sbjct: 554 RNSPRIEHYGCLVDLYSRAGRLKEAWDVIKKMPMMPNEVVLGSLLAACRTQGDVELAEKV 733
Query: 525 AESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHV 584
+ +++ P Y L SN++A WDG ++R+ MKER ++K S +EI + H
Sbjct: 734 MKYQVELYPGGDSNYVLFSNIYAAVGKWDGASKVRREMKERGLQKNLAFSSIEIDSGIHK 913
Query: 585 FVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVED 627
FVS H E++ I ++ L + GYVP+ S DV+D
Sbjct: 914 FVSGDKYHEENDYIYSALELLSFELHLYGYVPDFSGKESDVDD 1042
Score = 91.3 bits (225), Expect = 1e-18
Identities = 70/281 (24%), Positives = 132/281 (46%), Gaps = 6/281 (2%)
Frame = +2
Query: 196 DIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVH 255
++ + ++ V+ +EEA+ R M +V D T + ++ CA +G L LGL VH
Sbjct: 50 NVVSWTVVIGGFVKKECYEEALECFREMQLAGVVPDFVTVIAIISACANLGALGLGLWVH 229
Query: 256 ARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFE 315
+++ + V + LIDMY +C + AR+VFDG+ RN+V W S++ + N +
Sbjct: 230 RLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFDGMSQRNLVSWNSIIVGFAVNGLAD 409
Query: 316 ETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNN--A 373
+ L M +E PN ++ L AC+ L+ G + A + K +N + +
Sbjct: 410 KALSFFRSMKKEGLEPNGVSYTSALTACSHAGLIDEGLKIFADI-KRDHRNSPRIEHYGC 586
Query: 374 LINMYSKSGDIDSSYNVFSD-TMCRDVTTWNSMICGYSHHG---LGQEALRVFQDMKSAG 429
L+++YS++G + +++V M + S++ G L ++ ++ ++ G
Sbjct: 587 LVDLYSRAGRLKEAWDVIKKMPMMPNEVVLGSLLAACRTQGDVELAEKVMKYQVELYPGG 766
Query: 430 ESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGL 470
+S N V F + +A G + EMK ++ L
Sbjct: 767 DS-NYVLFSNIYAAVGKW----DGASKVRREMKERGLQKNL 874
Score = 79.0 bits (193), Expect = 6e-15
Identities = 73/317 (23%), Positives = 133/317 (41%), Gaps = 3/317 (0%)
Frame = +2
Query: 77 ARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLS 136
A +FDK+P+++V SW ++ G++ Y E L F+ + + P+ ++S
Sbjct: 17 ALKLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREMQLAGVV----PDFVTVIAIIS 184
Query: 137 SCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDEND 196
+C++ G + G+ H + K V ++L +MYARC + LA QV + G + +
Sbjct: 185 ACANLGALGLGLWVHRLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFD---GMSQRN 355
Query: 197 IFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHA 256
+ +NS++ +G ++A+ R M E + + +Y + C+ G + GL++ A
Sbjct: 356 LVSWNSIIVGFAVNGLADKALSFFRSMKKEGLEPNGVSYTSALTACSHAGLIDEGLKIFA 535
Query: 257 RLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEE 316
+ R + S I+ YG L+ Y + +E
Sbjct: 536 DIKR------DHRNSPRIEHYG------------------------CLVDLYSRAGRLKE 625
Query: 317 TLDLLTCMDQEDTLPNECTFKVLLGAC---AGIALLKHGDLLHARLEKLGFKNYVIVNNA 373
D++ M +PNE LL AC + L + L G NYV+ +
Sbjct: 626 AWDVIKKMPM---MPNEVVLGSLLAACRTQGDVELAEKVMKYQVELYPGGDSNYVLFS-- 790
Query: 374 LINMYSKSGDIDSSYNV 390
N+Y+ G D + V
Sbjct: 791 --NIYAAVGKWDGASKV 835
Score = 73.6 bits (179), Expect = 3e-13
Identities = 39/133 (29%), Positives = 73/133 (54%)
Frame = +2
Query: 17 LPPLEYIVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRL 76
+P ++ ++ A+ L G +H ++ ++ R +V LNSLI +Y +CG + L
Sbjct: 149 VPDFVTVIAIISACANLGALGLGLWVHRLVMKKEF---RDNVKVLNSLIDMYARCGCIEL 319
Query: 77 ARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLS 136
AR +FD M R++ SWN ++ G+ +G + L F+++ PN +T+ L+
Sbjct: 320 ARQVFDGMSQRNLVSWNSIIVGFAVNGLADKALSFFRSMKKEGLE----PNGVSYTSALT 487
Query: 137 SCSHSGRVREGMQ 149
+CSH+G + EG++
Sbjct: 488 ACSHAGLIDEGLK 526
Score = 62.8 bits (151), Expect = 4e-10
Identities = 32/128 (25%), Positives = 68/128 (53%)
Frame = +2
Query: 382 GDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVL 441
GD+D + +F ++V +W +I G+ +EAL F++M+ AG P+ VT + ++
Sbjct: 2 GDVDDALKLFDKLPVKNVVSWTVVIGGFVKKECYEEALECFREMQLAGVVPDFVTVIAII 181
Query: 442 SACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWD 501
SACA+L + GL +++ + + ++ ++ +Y R G +E A + +
Sbjct: 182 SACANLGALGLGL-WVHRLVMKKEFRDNVKVLNSLIDMYARCGCIELARQVFDGMSQR-N 355
Query: 502 IVAWRTLL 509
+V+W +++
Sbjct: 356 LVSWNSII 379
>TC80535 weakly similar to GP|14335136|gb|AAK59848.1 At2g15690/F9O13.24
{Arabidopsis thaliana}, partial (55%)
Length = 1727
Score = 211 bits (538), Expect = 6e-55
Identities = 130/403 (32%), Positives = 209/403 (51%), Gaps = 2/403 (0%)
Frame = +2
Query: 305 MSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGF 364
++ + Q +E L+L+ + D C F++L C ++ +H + F
Sbjct: 209 LTRFCQEGKVKEALELMEKGIKADA---NC-FEILFDLCGKSKSVEDAKKVHDYFLQSTF 376
Query: 365 KNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQD 424
++ ++N +I MY + + VF R++ +W+ MI GY++ +G E L++F+
Sbjct: 377 RSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQ 556
Query: 425 MKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAG 484
M G + T + VLSAC V+ YL + + IEPG+EHY ++ + ++G
Sbjct: 557 MNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSG 736
Query: 485 LLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSN 544
L++AE F++ + + + TL N R+H + DL + E I+ +DP
Sbjct: 737 YLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSK--------- 889
Query: 545 MHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQ 604
A AN + KK S L+ +N + K+P + K ++
Sbjct: 890 --AVANK----------IPTPPPKKYTAISMLDGKNRIIEY-----KNPT---LYKDDEK 1009
Query: 605 LLAM--IKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNL 662
L+AM +K GYVP+ VLHD++ E KE L HSE+LAIAYGL+ P P+RIIKNL
Sbjct: 1010LIAMNSMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNL 1189
Query: 663 RMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
R+C DCH A+K++S++ R +IVRD RFHHF+DG C+C D+W
Sbjct: 1190RVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 1318
Score = 55.1 bits (131), Expect = 9e-08
Identities = 37/143 (25%), Positives = 67/143 (45%), Gaps = 1/143 (0%)
Frame = +2
Query: 241 LCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVV 300
LC + ++ +VH L+ D + +K+I+MYG C + +AR+VFD + NRN+
Sbjct: 308 LCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDS 487
Query: 301 WTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDL-LHARL 359
W ++ Y + +E L L M++ T +L AC ++ + L +
Sbjct: 488 WHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMK 667
Query: 360 EKLGFKNYVIVNNALINMYSKSG 382
K G + V L+++ +SG
Sbjct: 668 SKYGIEPGVEHYMGLLDVLGQSG 736
Score = 49.7 bits (117), Expect = 4e-06
Identities = 40/141 (28%), Positives = 64/141 (45%), Gaps = 7/141 (4%)
Frame = +2
Query: 26 LLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMP 85
L L +K + K +H L Q++ RSD N +I +Y C + AR +FD MP
Sbjct: 299 LFDLCGKSKSVEDAKKVHDYFL---QSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMP 469
Query: 86 LRDVFSWNDLMTGYMHSGNYLEVLVLFKNL------VSSSTTQHACPNEYVFTTVLSSCS 139
R++ SW+ ++ GY +S E L LF+ + ++S T VLS+C
Sbjct: 470 NRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSET----------MLAVLSACG 619
Query: 140 HSGRVREG-MQCHGYLFKYGL 159
+ V + + KYG+
Sbjct: 620 SAEAVEDAYIYLESMKSKYGI 682
>BG455368 weakly similar to GP|9759524|dbj selenium-binding protein-like
{Arabidopsis thaliana}, partial (33%)
Length = 687
Score = 191 bits (486), Expect = 6e-49
Identities = 91/221 (41%), Positives = 141/221 (63%), Gaps = 1/221 (0%)
Frame = +1
Query: 486 LEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNM 545
+EKA ++T ++ + W LL + +H N D+ + + S+ +++P ++G Y LLS
Sbjct: 7 VEKALQLVQTMPMEPNGGVWGALLGASHIHGNPDVAEIASRSLFELEPDNLGNYLLLSKT 186
Query: 546 HATANMWDGVVEIRKMMKERNIKKEPGASWLEIRN-VNHVFVSEGCKHPESNEINKKVQQ 604
+A A WD V +RK+M+E+ ++K PG SW+E +N + H F + KHPE NEI K +
Sbjct: 187 YALAAKWDDVSRVRKLMREKQLRKNPGCSWVEAKNGIIHEFFAGDVKHPEINEIKKALDD 366
Query: 605 LLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRM 664
LL +K GY P +++V +D++DE K L HSEKLA+AYGL+ + + I+I+KNLR+
Sbjct: 367 LLQRLKCTGYQPKLNSVPYDIDDEGKRCLLVSHSEKLALAYGLLSTDAGSTIKIMKNLRI 546
Query: 665 CDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
C+DCH + S +T R IIVRD FHHF +G+C+C + W
Sbjct: 547 CEDCHIVMCGASXLTGRKIIVRDNMXFHHFLNGACSCNNFW 669
>AW776322 weakly similar to PIR|H84859|H84 hypothetical protein At2g42920
[imported] - Arabidopsis thaliana, partial (27%)
Length = 633
Score = 170 bits (430), Expect = 2e-42
Identities = 81/203 (39%), Positives = 131/203 (63%), Gaps = 1/203 (0%)
Frame = +1
Query: 397 RDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGE-SPNNVTFVGVLSACAHLALVDKGLD 455
R ++ WNS+I G + +G +EA F ++S+ P++V+F+GVL+AC HL ++K D
Sbjct: 16 RGLSCWNSIIIGLAMNGHEREAFEFFSKLESSKLLKPDSVSFIGVLTACKHLGAINKARD 195
Query: 456 YLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVH 515
Y M ++IEP ++HYTC+V + +AGLLE+AE +K +K D + W +LL+SCR H
Sbjct: 196 YFELMMNKYEIEPSIKHYTCIVDVLGQAGLLEEAEELIKGMPLKPDAIIWGSLLSSCRKH 375
Query: 516 RNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASW 575
RN + +R A+ + +++P D Y L+SN+HA +N ++ +E R +MKE +KEPG S
Sbjct: 376 RNVQIARRAAQRVYELNPSDASGYVLMSNVHAASNKFEEAIEQRLLMKENLTEKEPGCSS 555
Query: 576 LEIRNVNHVFVSEGCKHPESNEI 598
+E+ H F++ G HP++ EI
Sbjct: 556 IELYGEVHEFIAGGRLHPKTQEI 624
>AL370459 weakly similar to GP|20146256|d hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (11%)
Length = 477
Score = 157 bits (397), Expect = 1e-38
Identities = 72/126 (57%), Positives = 92/126 (72%)
Frame = +1
Query: 580 NVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSE 639
N H F HP++ +I +K+ +L IK +GYVPN VLHDVEDEQKE YL HSE
Sbjct: 4 NQVHKFHVGDTLHPKAQQIYEKLDELALKIKNVGYVPNTDFVLHDVEDEQKEQYLFQHSE 183
Query: 640 KLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSC 699
KLA+A+ L+ P+P PIR+ KNLR+C DCHTA+K IS V+ R I+VRDANRFHH +DG+C
Sbjct: 184 KLAVAFALISTPNPKPIRVFKNLRVCGDCHTAIKYISMVSGREIVVRDANRFHHMKDGTC 363
Query: 700 TCADHW 705
+C D+W
Sbjct: 364 SCNDYW 381
>BF633384 similar to PIR|T46179|T46 hypothetical protein T8H10.30 -
Arabidopsis thaliana, partial (22%)
Length = 595
Score = 145 bits (367), Expect(2) = 5e-38
Identities = 66/175 (37%), Positives = 111/175 (62%), Gaps = 6/175 (3%)
Frame = +1
Query: 401 TWNSMICGYSHHGLGQEALRVFQDMKSAGES-----PNNVTFVGVLSACAHLALVDKGLD 455
TWN +I Y HG G+EAL++F+ M G++ PN VT++ + ++ +H +VD+GL+
Sbjct: 4 TWNVLIMAYGMHGKGEEALKLFRRMVEEGDNNREIRPNEVTYIAIFASLSHSGMVDEGLN 183
Query: 456 YLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIV-AWRTLLNSCRV 514
Y H IEP +HY C+V L R+G +E+A N +KT V AW +LL +C++
Sbjct: 184 LFYTMKAKHGIEPTSDHYACLVDLLGRSGQIEEAYNLIKTMPSNMKKVDAWSSLLGACKI 363
Query: 515 HRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKK 569
H+N ++G+ A+++ +DP Y LLSN++++A +WD +++RK MKE+ ++K
Sbjct: 364 HQNLEIGEIAAKNLFVLDPNVASYYVLLSNIYSSAGLWDQAIDVRKKMKEKGVRK 528
Score = 44.7 bits (104), Expect = 1e-04
Identities = 49/183 (26%), Positives = 74/183 (39%), Gaps = 35/183 (19%)
Frame = +1
Query: 91 SWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHAC-PNEYVFTTVLSSCSHSGRVREGMQ 149
+WN L+ Y G E L LF+ +V PNE + + +S SHSG V EG+
Sbjct: 4 TWNVLIMAYGMHGKGEEALKLFRRMVEEGDNNREIRPNEVTYIAIFASLSHSGMVDEGLN 183
Query: 150 C-------HG-------------YLFKYGLV--SYQFVKSALFNMY----------ARCL 177
HG L + G + +Y +K+ NM A +
Sbjct: 184 LFYTMKAKHGIEPTSDHYACLVDLLGRSGQIEEAYNLIKTMPSNMKKVDAWSSLLGACKI 363
Query: 178 HVNLALQVLNTES--GGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATY 235
H NL + + ++ D N Y L N+ +G W++AI V ++M E V N
Sbjct: 364 HQNLEIGEIAAKNLFVLDPNVASYYVLLSNIYSSAGLWDQAIDVRKKM-KEKGVRKNLDV 540
Query: 236 VGV 238
VG+
Sbjct: 541 VGL 549
Score = 30.8 bits (68), Expect(2) = 5e-38
Identities = 12/23 (52%), Positives = 16/23 (69%)
Frame = +3
Query: 568 KKEPGASWLEIRNVNHVFVSEGC 590
KKEPG SW+E + H F++ GC
Sbjct: 522 KKEPGCSWIEHGDEVHKFLAGGC 590
>TC83482 weakly similar to GP|9758453|dbj|BAB08982.1 selenium-binding
protein-like {Arabidopsis thaliana}, partial (10%)
Length = 895
Score = 152 bits (384), Expect = 4e-37
Identities = 75/180 (41%), Positives = 111/180 (61%)
Frame = +2
Query: 398 DVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYL 457
D+ +WN++I GY+ HGL AL F MK G +P+ VTFV VLSAC H LV++G +
Sbjct: 2 DLVSWNAIIGGYASHGLATRALEEFDRMKVVG-TPDEVTFVNVLSACVHAGLVEEGEKHF 178
Query: 458 YNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRN 517
+ + + I+ +EHY+CMV LY RAG ++AEN +K + D+V W LL +C +H N
Sbjct: 179 TDMLTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLIKNMPFEPDVVLWGALLAACGLHSN 358
Query: 518 YDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLE 577
+LG+ AE I +++ +Y++LS + +W V E+R MKER IKK+ SW+E
Sbjct: 359 LELGEYAAERIRRLESSHPVSYSVLSKIQGEKGVWSSVNELRDTMKERGIKKQTAISWVE 538
Score = 45.8 bits (107), Expect = 6e-05
Identities = 35/118 (29%), Positives = 55/118 (45%), Gaps = 1/118 (0%)
Frame = +2
Query: 88 DVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREG 147
D+ SWN ++ GY G L F + T P+E F VLS+C H+G V EG
Sbjct: 2 DLVSWNAIIGGYASHGLATRALEEFDRMKVVGT-----PDEVTFVNVLSACVHAGLVEEG 166
Query: 148 -MQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLL 204
L KYG+ + S + ++Y R + A ++ ++ E D+ L+ +LL
Sbjct: 167 EKHFTDMLTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLI--KNMPFEPDVVLWGALL 334
Score = 43.1 bits (100), Expect = 4e-04
Identities = 44/185 (23%), Positives = 87/185 (46%), Gaps = 5/185 (2%)
Frame = +2
Query: 297 NVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDL-L 355
++V W +++ Y + L+ M T P+E TF +L AC L++ G+
Sbjct: 2 DLVSWNAIIGGYASHGLATRALEEFDRMKVVGT-PDEVTFVNVLSACVHAGLVEEGEKHF 178
Query: 356 HARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCR-DVTTWNSMI--CG-YSH 411
L K G + + + ++++Y ++G D + N+ + DV W +++ CG +S+
Sbjct: 179 TDMLTKYGIQAEMEHYSCMVDLYGRAGRFDEAENLIKNMPFEPDVVLWGALLAACGLHSN 358
Query: 412 HGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLE 471
LG+ A + + ES + V++ VLS +KG+ NE+++ E G++
Sbjct: 359 LELGEYAAERIRRL----ESSHPVSY-SVLSKIQG----EKGVWSSVNELRDTMKERGIK 511
Query: 472 HYTCM 476
T +
Sbjct: 512 KQTAI 526
>CA921534 similar to PIR|D96516|D96 F16N3.14 [imported] - Arabidopsis
thaliana, partial (18%)
Length = 790
Score = 150 bits (378), Expect = 2e-36
Identities = 89/237 (37%), Positives = 131/237 (54%), Gaps = 2/237 (0%)
Frame = -1
Query: 471 EHYTCMVVLY-CRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESIL 529
E Y +V ++ C GL+ KA FM+ ++ + W TL N R+H + + E +
Sbjct: 784 EPYLGVVNIFGCAVGLM-KAHEFMRNMPIEPGVELWETLRNFARIHGDLEREDCADELLT 608
Query: 530 QMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSE- 588
+DP A A+ + + +R KK+ + LE +N VSE
Sbjct: 607 VLDPSK-----------AAAD--------KVPLPQR--KKQSAINMLEEKNR----VSEY 503
Query: 589 GCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLM 648
C P E + K++ L ++ GYVP+ VLHD+++E+KE L HSE+LAIAYGL+
Sbjct: 502 RCNMPYKEEGDVKLRGLTGQMREAGYVPDTRYVLHDIDEEEKEKALQYHSERLAIAYGLI 323
Query: 649 KMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
P +RIIKNLR+C DCH A+K++SK+ R +IVRD RFHHF+DG C+C D+W
Sbjct: 322 STPPRTTLRIIKNLRICGDCHNAIKIMSKIVGRELIVRDNKRFHHFKDGKCSCGDYW 152
>TC90451 similar to PIR|H84508|H84508 hypothetical protein At2g13600
[imported] - Arabidopsis thaliana, partial (12%)
Length = 1362
Score = 102 bits (255), Expect = 4e-22
Identities = 69/239 (28%), Positives = 117/239 (48%), Gaps = 4/239 (1%)
Frame = +2
Query: 90 FSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQ 149
F+WN+++ Y + L ++ +++ + P+ Y VL + S S ++ G Q
Sbjct: 383 FNWNNIIRSYTRLESPQNALRIYVSMLRAGVL----PDRYTLPIVLKAVSQSFAIQLGQQ 550
Query: 150 CHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVE 209
H Y K GL S ++ +S N+Y + + A +V + E + +N+L++ L +
Sbjct: 551 VHSYGIKLGLQSNEYCESGFINLYCKAGDFDSAHKVFDE---NHEPKLGSWNALISGLSQ 721
Query: 210 SGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEF- 268
G +AI V M D T V VM C IGDL L L++H + + +E+
Sbjct: 722 GGLAMDAIVVFVDMKRHGFEPDGITMVSVMSACGSIGDLYLALQLHKYVFQAKT--NEWT 895
Query: 269 ---VGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCM 324
+ + LIDMYGKC + A +VF +++RNV WTS++ Y + + +E L CM
Sbjct: 896 VILMSNSLIDMYGKCGRMDLAYEVFATMEDRNVSSWTSMIVGYAMHGHAKEALGCFHCM 1072
Score = 95.9 bits (237), Expect(2) = 2e-33
Identities = 70/239 (29%), Positives = 117/239 (48%), Gaps = 7/239 (2%)
Frame = +2
Query: 198 FLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHAR 257
F +N+++ + A+ + M ++ D T V+ +Q +QLG +VH+
Sbjct: 383 FNWNNIIRSYTRLESPQNALRIYVSMLRAGVLPDRYTLPIVLKAVSQSFAIQLGQQVHSY 562
Query: 258 LLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEET 317
++ GL +E+ S I++Y K +A KVFD + W +L+S Q +
Sbjct: 563 GIKLGLQSNEYCESGFINLYCKAGDFDSAHKVFDENHEPKLGSWNALISGLSQGGLAMDA 742
Query: 318 LDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDL-LHARLEKLGFK------NYVIV 370
+ + M + P+ T ++ AC I GDL L +L K F+ +++
Sbjct: 743 IVVFVDMKRHGFEPDGITMVSVMSACGSI-----GDLYLALQLHKYVFQAKTNEWTVILM 907
Query: 371 NNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAG 429
+N+LI+MY K G +D +Y VF+ R+V++W SMI GY+ HG +EAL F MK G
Sbjct: 908 SNSLIDMYGKCGRMDLAYEVFATMEDRNVSSWTSMIVGYAMHGHAKEALGCFHCMKGVG 1084
Score = 90.9 bits (224), Expect = 2e-18
Identities = 55/221 (24%), Positives = 110/221 (48%), Gaps = 6/221 (2%)
Frame = +2
Query: 301 WTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLE 360
W +++ +Y + + L + M + LP+ T ++L A + ++ G +H+
Sbjct: 389 WNNIIRSYTRLESPQNALRIYVSMLRAGVLPDRYTLPIVLKAVSQSFAIQLGQQVHSYGI 568
Query: 361 KLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALR 420
KLG ++ + IN+Y K+GD DS++ VF + + +WN++I G S GL +A+
Sbjct: 569 KLGLQSNEYCESGFINLYCKAGDFDSAHKVFDENHEPKLGSWNALISGLSQGGLAMDAIV 748
Query: 421 VFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVV-- 478
VF DMK G P+ +T V V+SAC + G YL ++ + + +T +++
Sbjct: 749 VFVDMKRHGFEPDGITMVSVMSACGSI-----GDLYLALQLHKYVFQAKTNEWTVILMSN 913
Query: 479 ----LYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVH 515
+Y + G ++ A T + + ++ +W +++ +H
Sbjct: 914 SLIDMYGKCGRMDLAYEVFATMEDR-NVSSWTSMIVGYAMH 1033
Score = 65.5 bits (158), Expect(2) = 2e-33
Identities = 37/80 (46%), Positives = 48/80 (59%), Gaps = 1/80 (1%)
Frame = +1
Query: 427 SAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKN-HKIEPGLEHYTCMVVLYCRAGL 485
S G PN VTF+GVLSAC H V +G Y ++ MKN + I P L+HY CMV L RAGL
Sbjct: 1078 SRGVKPNYVTFIGVLSACVHGGTVQEGRFY-FDMMKNIYGITPQLQHYGCMVDLLGRAGL 1254
Query: 486 LEKAENFMKTTKVKWDIVAW 505
+ A ++ +K + V W
Sbjct: 1255 FDDARRMVEEMPMKPNSVVW 1314
Score = 60.1 bits (144), Expect = 3e-09
Identities = 48/215 (22%), Positives = 100/215 (46%), Gaps = 2/215 (0%)
Frame = +2
Query: 11 VVNSRYLPPLEYIVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVK 70
V+ RY P+ +LK + + + G+ +H+ + + +S+ + I+LY K
Sbjct: 473 VLPDRYTLPI-----VLKAVSQSFAIQLGQQVHSYGI---KLGLQSNEYCESGFINLYCK 628
Query: 71 CGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYV 130
G A +FD+ + SWN L++G G ++ +V+F ++ P+
Sbjct: 629 AGDFDSAHKVFDENHEPKLGSWNALISGLSQGGLAMDAIVVFVDMKRHGFE----PDGIT 796
Query: 131 FTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFV--KSALFNMYARCLHVNLALQVLNT 188
+V+S+C G + +Q H Y+F+ + + ++L +MY +C ++LA +V T
Sbjct: 797 MVSVMSACGSIGDLYLALQLHKYVFQAKTNEWTVILMSNSLIDMYGKCGRMDLAYEVFAT 976
Query: 189 ESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRM 223
++ ++ + S++ G +EA+G M
Sbjct: 977 M---EDRNVSSWTSMIVGYAMHGHAKEALGCFHCM 1072
Score = 28.5 bits (62), Expect = 9.2
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = +1
Query: 126 PNEYVFTTVLSSCSHSGRVREG 147
PN F VLS+C H G V+EG
Sbjct: 1093 PNYVTFIGVLSACVHGGTVQEG 1158
>BE941714 weakly similar to GP|11994734|dbj selenium-binding protein-like
{Arabidopsis thaliana}, partial (10%)
Length = 619
Score = 133 bits (335), Expect = 2e-31
Identities = 70/201 (34%), Positives = 114/201 (55%), Gaps = 5/201 (2%)
Frame = +2
Query: 368 VIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKS 427
V ++ +L++MY+K G+++ + +F RDV WN+MI G + HG G+ AL++F DM+
Sbjct: 14 VRLSTSLLDMYAKCGNLELAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDMEK 193
Query: 428 AGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLE 487
G P+++TF+ V +AC++ + +GL L + I P EHY C+V L RAGL E
Sbjct: 194 VGVKPDDITFIAVFTACSYSGMAYEGLMLLDKMCSVYNIVPKSEHYGCLVDLLSRAGLFE 373
Query: 488 KAENFMKTTKVKW----DIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQ-DMGTYTLL 542
+A ++ W + +AWR L++C H L + AE +LQ+D G Y LL
Sbjct: 374 EAMVMIRKITNSWNGSEETLAWRAFLSACCNHGETQLAELAAEKVLQLDNHIHSGVYVLL 553
Query: 543 SNMHATANMWDGVVEIRKMMK 563
SN++A + + ++ MMK
Sbjct: 554 SNLYAASGKHNDARRVKDMMK 616
Score = 66.2 bits (160), Expect = 4e-11
Identities = 55/233 (23%), Positives = 100/233 (42%), Gaps = 4/233 (1%)
Frame = +2
Query: 63 SLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQ 122
SL+ +Y KCG L LA+ +FD M +RDV WN +++G G+ L LF ++
Sbjct: 29 SLLDMYAKCGNLELAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDMEKVGVK- 205
Query: 123 HACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLA 182
P++ F V ++CS+SG EG+ + +++N+ + H
Sbjct: 206 ---PDDITFIAVFTACSYSGMAYEGLMLLDKM------------CSVYNIVPKSEH---- 328
Query: 183 LQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNA----TYVGV 238
Y L+++L +G +EEA+ ++R++ N W+ + +
Sbjct: 329 -----------------YGCLVDLLSRAGLFEEAMVMIRKITNS---WNGSEETLAWRAF 448
Query: 239 MGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFD 291
+ C G+ QL ++L+ V L ++Y +AR+V D
Sbjct: 449 LSACCNHGETQLAELAAEKVLQLDNHIHSGVYVLLSNLYAASGKHNDARRVKD 607
Score = 51.2 bits (121), Expect = 1e-06
Identities = 38/164 (23%), Positives = 78/164 (47%), Gaps = 14/164 (8%)
Frame = +2
Query: 269 VGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQED 328
+ + L+DMY KC ++ A+++FD + R+VV W +++S + + L L M++
Sbjct: 20 LSTSLLDMYAKCGNLELAKRLFDSMNMRDVVCWNAMISGMAMHGDGKGALKLFYDMEKVG 199
Query: 329 TLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNN----ALINMYSKSG-- 382
P++ TF + AC+ + G +L L+K+ ++ + L+++ S++G
Sbjct: 200 VKPDDITFIAVFTACSYSGMAYEGLML---LDKMCSVYNIVPKSEHYGCLVDLLSRAGLF 370
Query: 383 --------DIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEA 418
I +S+N +T+ W + + +HG Q A
Sbjct: 371 EEAMVMIRKITNSWNGSEETL-----AWRAFLSACCNHGETQLA 487
Score = 32.0 bits (71), Expect = 0.84
Identities = 19/88 (21%), Positives = 46/88 (51%)
Frame = +2
Query: 165 VKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMG 224
+ ++L +MYA+C ++ LA ++ ++ + D+ +N++++ + G + A+ + M
Sbjct: 20 LSTSLLDMYAKCGNLELAKRLFDSM---NMRDVVCWNAMISGMAMHGDGKGALKLFYDME 190
Query: 225 NECMVWDNATYVGVMGLCAQIGDLQLGL 252
+ D+ T++ V C+ G GL
Sbjct: 191 KVGVKPDDITFIAVFTACSYSGMAYEGL 274
>AW774626 similar to PIR|T47909|T47 hypothetical protein T20K12.70 -
Arabidopsis thaliana, partial (12%)
Length = 703
Score = 123 bits (308), Expect = 3e-28
Identities = 73/239 (30%), Positives = 127/239 (52%), Gaps = 6/239 (2%)
Frame = +3
Query: 273 LIDMYGKCCHVLNARKVFDGLQ--NRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTL 330
L+DMY KC V A +F GL+ +N V+WT++++ Y QN + ++ M +
Sbjct: 3 LVDMYAKCKCVSEAEFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQGVE 182
Query: 331 PNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNV 390
N+ TF +L AC+ + G+ +H + K GF + V V +AL++MY+K GD+ ++ N+
Sbjct: 183 CNQYTFPTILTACSSVLARCFGEQVHGFIVKSGFGSNVYVQSALVDMYAKCGDLKNAKNM 362
Query: 391 FSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALV 450
DV +WNS++ G+ HGL +EALR+F++M ++ TF VL+ C ++
Sbjct: 363 LETMEDDDVVSWNSLMVGFVRHGLEEEALRLFKNMHGRNMKIDDYTFPSVLNCCVVGSIN 542
Query: 451 DKGLDYLYNEMKNHKIEPGLEHY----TCMVVLYCRAGLLEKAENFMKTTKVKWDIVAW 505
K + L I+ G E+Y +V +Y + G ++ A + K D+++W
Sbjct: 543 PKSVHGLI-------IKTGFENYKLVSNALVDMYAKTGDMDCAYTVFEKMLEK-DVISW 695
Score = 115 bits (288), Expect = 6e-26
Identities = 68/236 (28%), Positives = 129/236 (53%)
Frame = +3
Query: 169 LFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECM 228
L +MYA+C V+ A + L D + L+ +++ ++G +A+ R M + +
Sbjct: 3 LVDMYAKCKCVSEA-EFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQGV 179
Query: 229 VWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARK 288
+ T+ ++ C+ + G +VH +++ G + +V S L+DMY KC + NA+
Sbjct: 180 ECNQYTFPTILTACSSVLARCFGEQVHGFIVKSGFGSNVYVQSALVDMYAKCGDLKNAKN 359
Query: 289 VFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIAL 348
+ + +++ +VV W SLM ++++ EE L L M + ++ TF +L C ++
Sbjct: 360 MLETMEDDDVVSWNSLMVGFVRHGLEEEALRLFKNMHGRNMKIDDYTFPSVLNCCVVGSI 539
Query: 349 LKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNS 404
+ +H + K GF+NY +V+NAL++MY+K+GD+D +Y VF + +DV +W S
Sbjct: 540 --NPKSVHGLIIKTGFENYKLVSNALVDMYAKTGDMDCAYTVFEKMLEKDVISWTS 701
Score = 103 bits (257), Expect = 2e-22
Identities = 70/242 (28%), Positives = 124/242 (50%), Gaps = 2/242 (0%)
Frame = +3
Query: 64 LIHLYVKCGRLRLARSMFDKMPL--RDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTT 121
L+ +Y KC + A +F + ++ W ++TGY +G+ + + F+ + +
Sbjct: 3 LVDMYAKCKCVSEAEFLFKGLEFDRKNHVLWTAMVTGYAQNGDGYKAVEFFRYMHAQGVE 182
Query: 122 QHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNL 181
C N+Y F T+L++CS G Q HG++ K G S +V+SAL +MYA+C +
Sbjct: 183 ---C-NQYTFPTILTACSSVLARCFGEQVHGFIVKSGFGSNVYVQSALVDMYAKCGDLKN 350
Query: 182 ALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGL 241
A +L T +++D+ +NSL+ V G EEA+ + + M M D+ T+ V+
Sbjct: 351 AKNMLETM---EDDDVVSWNSLMVGFVRHGLEEEALRLFKNMHGRNMKIDDYTFPSVLNC 521
Query: 242 CAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVW 301
C +G + VH +++ G + V + L+DMY K + A VF+ + ++V+ W
Sbjct: 522 CV-VGSIN-PKSVHGLIIKTGFENYKLVSNALVDMYAKTGDMDCAYTVFEKMLEKDVISW 695
Query: 302 TS 303
TS
Sbjct: 696 TS 701
Score = 73.9 bits (180), Expect = 2e-13
Identities = 49/153 (32%), Positives = 83/153 (54%)
Frame = +3
Query: 33 AKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSW 92
A+C G+ +H ++ ++ S+V ++L+ +Y KCG L+ A++M + M DV SW
Sbjct: 234 ARCF--GEQVHGFIV---KSGFGSNVYVQSALVDMYAKCGDLKNAKNMLETMEDDDVVSW 398
Query: 93 NDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHG 152
N LM G++ G E L LFKN+ ++ ++Y F +VL+ C G + HG
Sbjct: 399 NSLMVGFVRHGLEEEALRLFKNMHG----RNMKIDDYTFPSVLNCCV-VGSINP-KSVHG 560
Query: 153 YLFKYGLVSYQFVKSALFNMYARCLHVNLALQV 185
+ K G +Y+ V +AL +MYA+ ++ A V
Sbjct: 561 LIIKTGFENYKLVSNALVDMYAKTGDMDCAYTV 659
Score = 32.3 bits (72), Expect = 0.64
Identities = 13/31 (41%), Positives = 21/31 (66%)
Frame = +3
Query: 62 NSLIHLYVKCGRLRLARSMFDKMPLRDVFSW 92
N+L+ +Y K G + A ++F+KM +DV SW
Sbjct: 603 NALVDMYAKTGDMDCAYTVFEKMLEKDVISW 695
>BI308000 similar to GP|7363286|dbj| Similar to Arabidopsis thaliana DNA
chromosome 4 BAC clone F4I10; vegetative storage
protein., partial (12%)
Length = 771
Score = 115 bits (288), Expect = 6e-26
Identities = 55/111 (49%), Positives = 75/111 (67%)
Frame = +3
Query: 595 SNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPA 654
S EI ++ L I+ GY+P+ +++ HDVE++ KE L+ HSE+LAIA+GL+
Sbjct: 6 SEEIYAFLEALGDKIRDAGYIPDTNSI-HDVEEKVKEQLLSSHSERLAIAFGLLNTNHGT 182
Query: 655 PIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
PI + KNLR+C DCH K IS VT R IIVRD RFHHF++G C+C D+W
Sbjct: 183 PIHVRKNLRVCGDCHDVTKYISLVTGREIIVRDLRRFHHFKNGICSCGDYW 335
>BE202782 weakly similar to GP|22093801|dbj hypothetical protein~predicted by
GlimmerM~similar to Arabidopsis thaliana chromosome3
At3g23330, partial (13%)
Length = 548
Score = 112 bits (281), Expect = 4e-25
Identities = 49/93 (52%), Positives = 66/93 (70%)
Frame = +2
Query: 613 GYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAV 672
G +P +VL DVE++ KE L HSEKLA+ GL+ P+++IKNLR+CDDCH +
Sbjct: 2 GCLPMTKSVLQDVEEQDKEQILCGHSEKLAVVLGLINTSPGQPLQVIKNLRICDDCHAVI 181
Query: 673 KLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
K+IS++ R I VRD NRFHHF++G C+CAD W
Sbjct: 182 KVISRLEGREIFVRDTNRFHHFKEGVCSCADFW 280
>BG645370 weakly similar to PIR|T00405|T004 hypothetical protein At2g44880
[imported] - Arabidopsis thaliana, partial (11%)
Length = 784
Score = 100 bits (248), Expect(2) = 2e-24
Identities = 42/138 (30%), Positives = 81/138 (58%)
Frame = -2
Query: 437 FVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTT 496
F VL+AC H +DKG +Y + N+ + P ++ Y C++ LY R G L KA + M+
Sbjct: 783 FTAVLTACNHAGFIDKGEEYFNKMITNYGLSPDIDIYACLIDLYARNGNLRKARDLMEEM 604
Query: 497 KVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVV 556
+ + W + L++C+++ + +LG+ A +++M+P + Y L++++ T +W+
Sbjct: 603 PYDPNCIIWSSFLSACKIYGDVELGREAAIQLIKMEPCNAAPYLTLAHIYTTKGLWNEAS 424
Query: 557 EIRKMMKERNIKKEPGAS 574
E+R +M++R +K PG S
Sbjct: 423 EVRSLMQQRVKRKPPGWS 370
Score = 30.8 bits (68), Expect(2) = 2e-24
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = -3
Query: 574 SWLEIRNVNHVFVSEGCKHPESNEI 598
SW+E+ NHVF + H +SNEI
Sbjct: 365 SWVEVDKHNHVFAVDDVTHQQSNEI 291
>BF642497 weakly similar to PIR|H96713|H96 hypothetical protein T6L1.11
[imported] - Arabidopsis thaliana, partial (20%)
Length = 461
Score = 108 bits (271), Expect = 5e-24
Identities = 50/150 (33%), Positives = 90/150 (59%)
Frame = +3
Query: 299 VVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHAR 358
+ WTS+++ + QN + +D+ M E+ ++ TF +L AC G+ L+ G +HA
Sbjct: 6 ISWTSMITGFTQNGLDRDAIDIFREMKLENLQMDQYTFGSVLTACGGVMALQEGKQVHAY 185
Query: 359 LEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEA 418
+ + +K+ + V +AL++MY K +I S+ VF C++V +W +M+ GY +G +EA
Sbjct: 186 IIRTDYKDNIFVASALVDMYCKCKNIKSAEAVFKKMTCKNVVSWTAMLVGYGQNGYSEEA 365
Query: 419 LRVFQDMKSAGESPNNVTFVGVLSACAHLA 448
++ F DM+ G P++ T V+S+CA+LA
Sbjct: 366 VKTFSDMQKYGIEPDDFTLGSVISSCANLA 455
Score = 87.4 bits (215), Expect = 2e-17
Identities = 48/150 (32%), Positives = 81/150 (54%)
Frame = +3
Query: 200 YNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLL 259
+ S++ ++G +AI + R M E + D T+ V+ C + LQ G +VHA ++
Sbjct: 12 WTSMITGFTQNGLDRDAIDIFREMKLENLQMDQYTFGSVLTACGGVMALQEGKQVHAYII 191
Query: 260 RGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLD 319
R + FV S L+DMY KC ++ +A VF + +NVV WT+++ Y QN Y EE +
Sbjct: 192 RTDYKDNIFVASALVDMYCKCKNIKSAEAVFKKMTCKNVVSWTAMLVGYGQNGYSEEAVK 371
Query: 320 LLTCMDQEDTLPNECTFKVLLGACAGIALL 349
+ M + P++ T ++ +CA +A L
Sbjct: 372 TFSDMQKYGIEPDDFTLGSVISSCANLASL 461
Score = 58.2 bits (139), Expect = 1e-08
Identities = 38/158 (24%), Positives = 74/158 (46%)
Frame = +3
Query: 91 SWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQC 150
SW ++TG+ +G + + +F+ + + ++Y F +VL++C ++EG Q
Sbjct: 9 SWTSMITGFTQNGLDRDAIDIFREMKLENLQM----DQYTFGSVLTACGGVMALQEGKQV 176
Query: 151 HGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVES 210
H Y+ + FV SAL +MY +C ++ A V + ++ + ++L ++
Sbjct: 177 HAYIIRTDYKDNIFVASALVDMYCKCKNIKSAEAVFKKMTC---KNVVSWTAMLVGYGQN 347
Query: 211 GRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDL 248
G EEA+ M + D+ T V+ CA + L
Sbjct: 348 GYSEEAVKTFSDMQKYGIEPDDFTLGSVISSCANLASL 461
Score = 54.7 bits (130), Expect = 1e-07
Identities = 27/105 (25%), Positives = 56/105 (52%)
Frame = +3
Query: 36 LTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDL 95
L GK +HA ++ D + ++ ++L+ +Y KC ++ A ++F KM ++V SW +
Sbjct: 156 LQEGKQVHAYIIRTDY---KDNIFVASALVDMYCKCKNIKSAEAVFKKMTCKNVVSWTAM 326
Query: 96 MTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSH 140
+ GY +G E + F ++ P+++ +V+SSC++
Sbjct: 327 LVGYGQNGYSEEAVKTFSDMQKYGIE----PDDFTLGSVISSCAN 449
>BE202961 weakly similar to GP|6016735|gb|A hypothetical protein {Arabidopsis
thaliana}, partial (14%)
Length = 613
Score = 108 bits (269), Expect = 9e-24
Identities = 57/187 (30%), Positives = 95/187 (50%)
Frame = +2
Query: 234 TYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGL 293
T V V+ CA I +L+ G+++H + G + V + L+DMY KC A +F+ +
Sbjct: 53 TVVSVLRACACISNLEEGMKIHELAVNYGFEMETTVSTALMDMYMKCFSPEKAVDIFNRM 232
Query: 294 QNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGD 353
++V+ W L S Y N E++ + M T P+ +L + + +L+
Sbjct: 233 PKKDVIAWAVLFSGYADNGMVHESMWVFRNMLSSGTRPDAIALVKILTTVSELGILQQAV 412
Query: 354 LLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHG 413
HA + K GF+N + +LI +Y+K I+ + VF +DV TW+S+I Y HG
Sbjct: 413 CFHAFVIKNGFENNQFIGASLIEVYAKCSSIEDANKVFKGMTYKDVVTWSSIIAAYGFHG 592
Query: 414 LGQEALR 420
G+EAL+
Sbjct: 593 QGEEALK 613
Score = 97.4 bits (241), Expect = 2e-20
Identities = 57/205 (27%), Positives = 103/205 (49%)
Frame = +2
Query: 114 NLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMY 173
+L S + PN +VL +C+ + EGM+ H YG V +AL +MY
Sbjct: 5 DLSSEMLDKRIKPNWVTVVSVLRACACISNLEEGMKIHELAVNYGFEMETTVSTALMDMY 184
Query: 174 ARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNA 233
+C A+ + N + D+ + L + ++G E++ V R M + D
Sbjct: 185 MKCFSPEKAVDIFNRMP---KKDVIAWAVLFSGYADNGMVHESMWVFRNMLSSGTRPDAI 355
Query: 234 TYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGL 293
V ++ +++G LQ + HA +++ G ++F+G+ LI++Y KC + +A KVF G+
Sbjct: 356 ALVKILTTVSELGILQQAVCFHAFVIKNGFENNQFIGASLIEVYAKCSSIEDANKVFKGM 535
Query: 294 QNRNVVVWTSLMSAYLQNRYFEETL 318
++VV W+S+++AY + EE L
Sbjct: 536 TYKDVVTWSSIIAAYGFHGQGEEAL 610
Score = 80.9 bits (198), Expect = 2e-15
Identities = 53/202 (26%), Positives = 100/202 (49%), Gaps = 4/202 (1%)
Frame = +2
Query: 318 LDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINM 377
LDL + M + PN T +L ACA I+ L+ G +H GF+ V+ AL++M
Sbjct: 2 LDLSSEMLDKRIKPNWVTVVSVLRACACISNLEEGMKIHELAVNYGFEMETTVSTALMDM 181
Query: 378 YSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTF 437
Y K + + ++F+ +DV W + GY+ +G+ E++ VF++M S+G P+ +
Sbjct: 182 YMKCFSPEKAVDIFNRMPKKDVIAWAVLFSGYADNGMVHESMWVFRNMLSSGTRPDAIAL 361
Query: 438 VGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHY----TCMVVLYCRAGLLEKAENFM 493
V +L+ + L ++ + + + +KN G E+ ++ +Y + +E A
Sbjct: 362 VKILTTVSELGILQQAVCFHAFVIKN-----GFENNQFIGASLIEVYAKCSSIEDANKVF 526
Query: 494 KTTKVKWDIVAWRTLLNSCRVH 515
K K D+V W +++ + H
Sbjct: 527 KGMTYK-DVVTWSSIIAAYGFH 589
Score = 80.9 bits (198), Expect = 2e-15
Identities = 55/207 (26%), Positives = 104/207 (49%)
Frame = +2
Query: 11 VVNSRYLPPLEYIVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVK 70
+++ R P +V +L+ A L G IH +L V + V +L+ +Y+K
Sbjct: 20 MLDKRIKPNWVTVVSVLRACACISNLEEGMKIH-ELAVNYGFEMETTVS--TALMDMYMK 190
Query: 71 CGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYV 130
C A +F++MP +DV +W L +GY +G E + +F+N++SS T P+
Sbjct: 191 CFSPEKAVDIFNRMPKKDVIAWAVLFSGYADNGMVHESMWVFRNMLSSGTR----PDAIA 358
Query: 131 FTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTES 190
+L++ S G +++ + H ++ K G + QF+ ++L +YA+C + A +V
Sbjct: 359 LVKILTTVSELGILQQAVCFHAFVIKNGFENNQFIGASLIEVYAKCSSIEDANKVF---K 529
Query: 191 GGDENDIFLYNSLLNVLVESGRWEEAI 217
G D+ ++S++ G+ EEA+
Sbjct: 530 GMTYKDVVTWSSIIAAYGFHGQGEEAL 610
>BQ122819 similar to PIR|C84748|C84 hypothetical protein At2g33680 [imported]
- Arabidopsis thaliana, partial (15%)
Length = 771
Score = 104 bits (259), Expect = 1e-22
Identities = 49/146 (33%), Positives = 85/146 (57%)
Frame = +2
Query: 482 RAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTL 541
RAG L +A+ F+++ V + WR LL +C+ HRNY+LG E ++++ + Y L
Sbjct: 26 RAGKLNEAKEFIESATVDHGLCLWRILLGACKNHRNYELGVYAGEKLVELGSPESSAYVL 205
Query: 542 LSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKK 601
LS+++ + V +R++MK R + KEPG SW+E++ + HVFV +HP+ +EI +
Sbjct: 206 LSSIYTALGDRENVERVRRIMKARGVNKEPGCSWIELKGLVHVFVVGDNQHPQVDEIRLE 385
Query: 602 VQQLLAMIKPLGYVPNISAVLHDVED 627
++ L ++ GY P + + V D
Sbjct: 386 LELLTKLMIDEGYQPLLDRLPETVID 463
>BF640739 weakly similar to GP|7363286|dbj Similar to Arabidopsis thaliana
DNA chromosome 4 BAC clone F4I10; vegetative storage
protein., partial (14%)
Length = 444
Score = 100 bits (249), Expect = 2e-21
Identities = 52/142 (36%), Positives = 82/142 (57%), Gaps = 1/142 (0%)
Frame = +1
Query: 316 ETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALI 375
E L+L M + P+ TF ++ A A +++ + +H + V V AL+
Sbjct: 7 EALNLFCTMQSQGIKPDSFTFVSVITALADLSVTRQAKWIHGLAIRTNMDTNVFVATALV 186
Query: 376 NMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKS-AGESPNN 434
+MY+K G I+++ +F R V TWN+MI GY HGLG+ AL +F DM++ A PN+
Sbjct: 187 DMYAKCGAIETARELFDMMQERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLKPND 366
Query: 435 VTFVGVLSACAHLALVDKGLDY 456
+TF+ V+SAC+H V++GL Y
Sbjct: 367 ITFLSVISACSHSGFVEEGLYY 432
Score = 72.8 bits (177), Expect = 4e-13
Identities = 40/139 (28%), Positives = 74/139 (52%), Gaps = 1/139 (0%)
Frame = +1
Query: 215 EAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLI 274
EA+ + M ++ + D+ T+V V+ A + + +H +R + + FV + L+
Sbjct: 7 EALNLFCTMQSQGIKPDSFTFVSVITALADLSVTRQAKWIHGLAIRTNMDTNVFVATALV 186
Query: 275 DMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTL-PNE 333
DMY KC + AR++FD +Q R+V+ W +++ Y + + LDL M E +L PN+
Sbjct: 187 DMYAKCGAIETARELFDMMQERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLKPND 366
Query: 334 CTFKVLLGACAGIALLKHG 352
TF ++ AC+ ++ G
Sbjct: 367 ITFLSVISACSHSGFVEEG 423
Score = 72.4 bits (176), Expect = 6e-13
Identities = 43/133 (32%), Positives = 69/133 (51%)
Frame = +1
Query: 24 VKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDK 83
V ++ AD K IH + + + ++V +L+ +Y KCG + AR +FD
Sbjct: 70 VSVITALADLSVTRQAKWIHGLAI---RTNMDTNVFVATALVDMYAKCGAIETARELFDM 240
Query: 84 MPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGR 143
M R V +WN ++ GY G L LF ++ + ++ + PN+ F +V+S+CSHSG
Sbjct: 241 MQERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLK---PNDITFLSVISACSHSGF 411
Query: 144 VREGMQCHGYLFK 156
V EG+ Y FK
Sbjct: 412 VEEGL----YYFK 438
Score = 50.8 bits (120), Expect = 2e-06
Identities = 32/140 (22%), Positives = 68/140 (47%), Gaps = 1/140 (0%)
Frame = +1
Query: 114 NLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMY 173
NL + +Q P+ + F +V+++ + R+ HG + + + FV +AL +MY
Sbjct: 16 NLFCTMQSQGIKPDSFTFVSVITALADLSVTRQAKWIHGLAIRTNMDTNVFVATALVDMY 195
Query: 174 ARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDN- 232
A+C + A ++ + E + +N++++ G + A+ + M NE + N
Sbjct: 196 AKCGAIETARELFDMM---QERHVITWNAMIDGYGTHGLGKAALDLFDDMQNEASLKPND 366
Query: 233 ATYVGVMGLCAQIGDLQLGL 252
T++ V+ C+ G ++ GL
Sbjct: 367 ITFLSVISACSHSGFVEEGL 426
>BG646586 weakly similar to GP|10177683|dbj selenium-binding protein-like
{Arabidopsis thaliana}, partial (9%)
Length = 687
Score = 99.4 bits (246), Expect = 4e-21
Identities = 47/112 (41%), Positives = 70/112 (61%)
Frame = -3
Query: 466 IEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIA 525
IEP EHYT MV L RAG LEKA + +K D W +LL +CRV+ N +LG+ A
Sbjct: 664 IEPLPEHYTGMVDLLGRAGQLEKAYSLIKENSTSADATTWGSLLAACRVYGNVELGEIAA 485
Query: 526 ESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLE 577
+ ++DP D G Y LL+N +A+ + W+ E++K+M ++ +KK G SW++
Sbjct: 484 RHLFEIDPTDSGNYVLLANTYASNDKWERAEEVKKLMSKKGMKKPSGYSWIQ 329
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.137 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,538,493
Number of Sequences: 36976
Number of extensions: 366901
Number of successful extensions: 1882
Number of sequences better than 10.0: 119
Number of HSP's better than 10.0 without gapping: 1664
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1817
length of query: 705
length of database: 9,014,727
effective HSP length: 103
effective length of query: 602
effective length of database: 5,206,199
effective search space: 3134131798
effective search space used: 3134131798
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0141.1


