

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
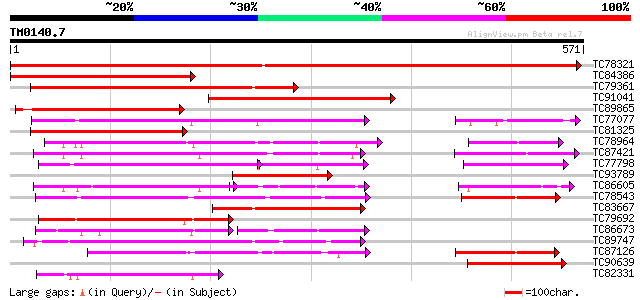
Query= TM0140.7
(571 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 953 0.0
TC84386 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 335 2e-92
TC79361 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 305 4e-83
TC91041 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 280 1e-75
TC89865 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 212 3e-55
TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter p... 201 8e-52
TC81325 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 196 2e-50
TC78964 weakly similar to PIR|T00450|T00450 probable monosacchar... 172 3e-43
TC87421 sugar transporter 169 3e-42
TC77798 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_14... 154 7e-38
TC93789 similar to GP|21689841|gb|AAM67564.1 unknown protein {Ar... 153 2e-37
TC86605 similar to PIR|T01853|T01853 probable hexose transport p... 98 3e-37
TC78543 similar to PIR|T14545|T14545 probable sugar transporter ... 151 7e-37
TC83667 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 150 1e-36
TC79692 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_14... 149 3e-36
TC86673 weakly similar to PIR|S25015|S25015 monosaccharide trans... 108 8e-36
TC89747 similar to PIR|F96696|F96696 protein F1N21.12 [imported]... 147 1e-35
TC87126 similar to GP|15724240|gb|AAL06513.1 At1g75220/F22H5_6 {... 139 3e-33
TC90639 similar to GP|17380890|gb|AAL36257.1 putative membrane t... 124 1e-28
TC82331 weakly similar to EGAD|120604|128852 hypothetical protei... 115 5e-26
>TC78321 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (80%)
Length = 2093
Score = 953 bits (2464), Expect = 0.0
Identities = 471/571 (82%), Positives = 519/571 (90%), Gaps = 1/571 (0%)
Frame = +2
Query: 1 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKA 60
MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRD+F A
Sbjct: 146 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDEFPA 325
Query: 61 VDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPA 120
V++KTWLQEAIVSTAIAGAIIGAA+GGWINDRFGR+V+II+ADTLFL+GS+I+AAAPNPA
Sbjct: 326 VEKKTWLQEAIVSTAIAGAIIGAAIGGWINDRFGRKVSIIVADTLFLLGSIILAAAPNPA 505
Query: 121 ILLFGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN 180
L+ GRVFVG GVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFT
Sbjct: 506 TLIVGRVFVGLGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTK 685
Query: 181 APGTWRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIE 240
APGTWRWMLGVAAAPA+IQIVLMLSLPESPRWLYRKG+EEE+K ILKKIY ED D EI+
Sbjct: 686 APGTWRWMLGVAAAPAVIQIVLMLSLPESPRWLYRKGKEEEAKVILKKIYEVEDYDNEIQ 865
Query: 241 ALKESVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQ 300
ALKESVE E++E T KIS+M L+KTT+VRRGLYAG+GL FFQQF GINTVMYYSP+IVQ
Sbjct: 866 ALKESVEMELKE--TEKISIMQLVKTTSVRRGLYAGVGLAFFQQFTGINTVMYYSPSIVQ 1039
Query: 301 LAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAFR 360
LAGFAS RTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISL GVVLSL LLTV FR
Sbjct: 1040LAGFASKRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLTGVVLSLTLLTVTFR 1219
Query: 361 ESELHAPTVSAIETSHYNNTCPDFR-AAVNPGRWTCMTCLKASPSCGFCAASPNKLLPGA 419
ESE+HAP VS +E+SHYNNTCP+F+ AA N W CM C++A +CGFCAA + PGA
Sbjct: 1220ESEIHAPMVSLVESSHYNNTCPEFKTAATNNHNWNCMKCIRAESTCGFCAAPGDMTKPGA 1399
Query: 420 CLISDDTTKDMCGNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVPWVINSEIYP 479
CLIS+ T+D+CG DHRAWYT+GCPS FGW A++ LALYII FSPGMGTVPWV+NSEIYP
Sbjct: 1400CLISNKNTEDICGVDHRAWYTKGCPSNFGWIAILALALYIIFFSPGMGTVPWVVNSEIYP 1579
Query: 480 LRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPET 539
LRYRG+CGGIASTTVW+SNL+VSQSFLSLT IG AWTFM+F I+A+VAIFFVI+FVPET
Sbjct: 1580LRYRGICGGIASTTVWVSNLVVSQSFLSLTVAIGPAWTFMIFAIIAIVAIFFVIIFVPET 1759
Query: 540 KGVPMEEVEKMLEQRSLQFKFWQKRDSGSEK 570
KGVPMEEVE MLE+R +Q KFW+KRDS SEK
Sbjct: 1760KGVPMEEVESMLEKRFVQIKFWKKRDSPSEK 1852
>TC84386 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (30%)
Length = 664
Score = 335 bits (860), Expect = 2e-92
Identities = 168/185 (90%), Positives = 176/185 (94%)
Frame = +2
Query: 1 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKA 60
MEGGVPEAD+SAFRECLSLSWKNPYVLRLAFSAGIGG LFGYDTGVISGALLYIRDDFKA
Sbjct: 110 MEGGVPEADVSAFRECLSLSWKNPYVLRLAFSAGIGGFLFGYDTGVISGALLYIRDDFKA 289
Query: 61 VDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPA 120
VDR+TWLQEAIVSTA+AGAIIGA+VGGWINDRFGR+ AII+AD LF IGSVIMAAA NPA
Sbjct: 290 VDRQTWLQEAIVSTALAGAIIGASVGGWINDRFGRKKAIILADALFFIGSVIMAAAINPA 469
Query: 121 ILLFGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN 180
IL+ GRVFVG GVGMASMASPLYISEASPTRVRGALVSLN FLITGGQ LSY+INLAFTN
Sbjct: 470 ILIVGRVFVGLGVGMASMASPLYISEASPTRVRGALVSLNGFLITGGQXLSYVINLAFTN 649
Query: 181 APGTW 185
APGTW
Sbjct: 650 APGTW 664
>TC79361 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (55%)
Length = 949
Score = 305 bits (780), Expect = 4e-83
Identities = 147/267 (55%), Positives = 196/267 (73%)
Frame = +1
Query: 21 WKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAI 80
+KNPY+L LA AGIGGLLFGYDTGVISGALLYI+DDF V +LQE IVS AIAGAI
Sbjct: 154 FKNPYILGLAAVAGIGGLLFGYDTGVISGALLYIKDDFPQVRNSNFLQETIVSMAIAGAI 333
Query: 81 IGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMAS 140
+GAA GGW+ND +GR+ A ++AD +F++G+++MAAAP+P +L+ GR+ VG GVG+AS+ +
Sbjct: 334 VGAAFGGWLNDAYGRKKATLLADVIFILGAILMAAAPDPYVLIAGRLLVGLGVGIASVTA 513
Query: 141 PLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAAPAIIQI 200
P+YI+E +P+ +RG+LVS N +ITGGQF+SYL+NL FT PGTWRWMLGV+ PA+IQ
Sbjct: 514 PVYIAEVAPSEIRGSLVSTNVLMITGGQFVSYLVNLVFTQVPGTWRWMLGVSGVPALIQF 693
Query: 201 VLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKISM 260
+ ML LPESPRWL+ K R+ E+ ++ KIY ++ EI+ L + +SE E + S I
Sbjct: 694 ICMLFLPESPRWLFIKNRKNEAVDVISKIYDLSRLEDEIDFL--TAQSEQERQRRSTIKF 867
Query: 261 MTLLKTTTVRRGLYAGMGLQFFQQFVG 287
+ ++ R G GL FQQF G
Sbjct: 868 WHVFRSKETRLAFLDGGGLLAFQQFTG 948
>TC91041 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (31%)
Length = 564
Score = 280 bits (716), Expect = 1e-75
Identities = 139/187 (74%), Positives = 164/187 (87%), Gaps = 1/187 (0%)
Frame = +3
Query: 199 QIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTS-K 257
Q LM+ LPESPRWL+RKG+EEE+K++L++IY+PE+ +AEI LKESVE EI+ES++S K
Sbjct: 3 QFTLMIMLPESPRWLFRKGKEEEAKALLRRIYSPEEAEAEINTLKESVELEIKESESSDK 182
Query: 258 ISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLIT 317
S++ +LKT TVRRGLYAGMGLQ FQQFVGINTVMYYSP I+QLAGFASN+TALLLSL+T
Sbjct: 183 ASIIKMLKTKTVRRGLYAGMGLQIFQQFVGINTVMYYSPAIIQLAGFASNQTALLLSLVT 362
Query: 318 SGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAFRESELHAPTVSAIETSHY 377
SGLNAFGSILSIYFIDKTGRKKL L SL GVVLSL +LTV F ES H+P +SAIETSH+
Sbjct: 363 SGLNAFGSILSIYFIDKTGRKKLLLFSLSGVVLSLVVLTVVFHESSTHSPMISAIETSHF 542
Query: 378 NNTCPDF 384
NNTC D+
Sbjct: 543 NNTCTDY 563
>TC89865 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (33%)
Length = 617
Score = 212 bits (540), Expect = 3e-55
Identities = 104/169 (61%), Positives = 133/169 (78%)
Frame = +1
Query: 6 PEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKT 65
PE +S F KNPY+L L A IGGLLFGYDTGVISGALLYI+DDF+AV
Sbjct: 55 PERKVSVF--------KNPYILCLTSVASIGGLLFGYDTGVISGALLYIKDDFQAVRYSH 210
Query: 66 WLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFG 125
+LQE IVS A+AGAI+GAAVGGW+NDR+GR+ A IIAD +F++G+++MAAAP+P IL+ G
Sbjct: 211 FLQETIVSMAVAGAIVGAAVGGWMNDRYGRKKATIIADVIFILGAIVMAAAPDPYILILG 390
Query: 126 RVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLI 174
RV VG GVG+AS+ +P+YI+E SP+ +RG LV+ N +ITGGQF+SYL+
Sbjct: 391 RVLVGLGVGIASVTAPVYIAELSPSEIRGGLVATNVLMITGGQFISYLV 537
>TC77077 similar to GP|9293909|dbj|BAB01812.1 sugar transporter protein
{Arabidopsis thaliana}, partial (85%)
Length = 1752
Score = 201 bits (510), Expect = 8e-52
Identities = 117/351 (33%), Positives = 188/351 (53%), Gaps = 14/351 (3%)
Frame = +1
Query: 22 KNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAII 81
+N Y A A + +L GYD GV+SGA +YI+ D K D K E ++ ++I
Sbjct: 97 RNKYAFACAMLASMTSILLGYDIGVMSGAAIYIKRDLKVSDGKI---EVLLGIINIYSLI 267
Query: 82 GAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASP 141
G+ + G +D GRR I+ A +F +G+++M +PN L+FGR G G+G A M +P
Sbjct: 268 GSCLAGRTSDWIGRRYTIVFAGAIFFVGALLMGFSPNYNFLMFGRFVAGVGIGYALMIAP 447
Query: 142 LYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFT--NAPGTWRWMLGVAAAPAIIQ 199
+Y +E SP RG L S I G L Y+ N AF+ + WR MLGV A P++I
Sbjct: 448 VYTAEVSPASSRGFLTSFPEVFINSGILLGYISNYAFSKLSLKLGWRMMLGVGAIPSVIL 627
Query: 200 IVLMLSLPESPRWLYRKGREEESKSILKKIY-APEDVDAEIEALKES-----------VE 247
V +L++PESPRWL +GR ++ +L K + E+ + +K++ VE
Sbjct: 628 AVGVLAMPESPRWLVMRGRLGDAIKVLNKTSDSKEEAQLRLAEIKQAAGIPEDCNDDVVE 807
Query: 248 SEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASN 307
+++ + + L T VR + A +G+ FFQQ G++ V+ YSPTI + AG +
Sbjct: 808 VKVKNTGEGVWKELFLYPTPAVRHIVIAALGIHFFQQASGVDAVVLYSPTIFKKAGINGD 987
Query: 308 RTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVA 358
L+ ++ + +++ + +D+ GR+ L L S+ G+V+SL L V+
Sbjct: 988 THLLIATIAVGFVKTLFILVATFMLDRYGRRPLLLTSVGGMVVSLLTLAVS 1140
Score = 73.9 bits (180), Expect = 2e-13
Identities = 43/131 (32%), Positives = 70/131 (52%), Gaps = 6/131 (4%)
Frame = +1
Query: 445 SKFGWAALVGLAL---YIITFSPGMGTVPWVINSEIYPLRYR---GVCGGIASTTVWISN 498
+K WA + +A Y+ TFS G G + WV +SEI+PLR R CG + + +++
Sbjct: 1165 TKLNWAIGLSIATVLSYVATFSIGAGPITWVYSSEIFPLRLRAQGAACGVVVNR---VTS 1335
Query: 499 LIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQF 558
++S +FLSL++ I F +FG +A + F + +PET+G +EE+E
Sbjct: 1336 GVISMTFLSLSKGITIGGAFFLFGGIATIGWIFFYIMLPETQGKTLEEMEASFG------ 1497
Query: 559 KFWQKRDSGSE 569
K W+K + E
Sbjct: 1498 KIWRKSKNTKE 1530
>TC81325 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (32%)
Length = 636
Score = 196 bits (499), Expect = 2e-50
Identities = 96/158 (60%), Positives = 125/158 (78%), Gaps = 1/158 (0%)
Frame = +3
Query: 21 WKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAI 80
+KNPY+L + +AGIGGLLFGYDTGVISGALLYI+DDF V ++LQE IVS A+ GAI
Sbjct: 153 FKNPYILGVTAAAGIGGLLFGYDTGVISGALLYIKDDFDDVRNSSFLQETIVSMALVGAI 332
Query: 81 IGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMAS 140
IGAA GGWIND FGR+ A + AD +F +GSV+MA+AP+ +L+ GR+ VG GVG+AS+ +
Sbjct: 333 IGAATGGWINDAFGRKKATLSADVVFTLGSVVMASAPDAYVLILGRLLVGIGVGVASVTA 512
Query: 141 PLYISEASPTRVRGALVSLNSFLITGGQ-FLSYLINLA 177
P+YI+E+SP+ +RG+LVS N +ITGGQ F +NLA
Sbjct: 513 PVYIAESSPSEIRGSLVSTNVLMITGGQVFFLTFVNLA 626
>TC78964 weakly similar to PIR|T00450|T00450 probable monosaccharide
transport protein T14N5.7 - Arabidopsis thaliana,
partial (98%)
Length = 1771
Score = 172 bits (436), Expect = 3e-43
Identities = 116/360 (32%), Positives = 190/360 (52%), Gaps = 23/360 (6%)
Frame = +1
Query: 35 IGGLLFGYDTGVISGALL---YIRDDFKAVDRK--TWLQEA------------IVSTAIA 77
+GG LFGYD GV G ++ F V RK L+E S+
Sbjct: 133 LGGSLFGYDLGVSGGVTSMDDFLEKFFPDVYRKKHAHLKETDYCKYDNQVLTLFTSSLYF 312
Query: 78 GAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMAS 137
A++ ++ GR+ II+ FLIG+++ AAA N L+ GRVF+G G+G +
Sbjct: 313 SALVMTFFASYLTRNKGRKATIIVGALSFLIGAILNAAAQNIPTLIIGRVFLGGGIGFGN 492
Query: 138 MASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNA--PGTWRWMLGVAAAP 195
A PLY+SE +P RGA+ L F G ++ L+N FT+ P WR LG+A P
Sbjct: 493 QAVPLYLSEMAPASSRGAVNQLFQFTTCAGILIANLVNY-FTDKIHPHGWRISLGLAGIP 669
Query: 196 AIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKT 255
A++ ++ + E+P L +GR +E++ +L+K+ ++VDAE E LK++ SE+ ++
Sbjct: 670 AVLMLLGGIFCAETPNSLVEQGRLDEARKVLEKVRGTKNVDAEFEDLKDA--SELAQAVK 843
Query: 256 SKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSL 315
S + LLK + + +G FQQ G N++++Y+P I Q +GF SN AL S
Sbjct: 844 SPFKV--LLKRKYRPQLIIGALGTPAFQQLTGNNSILFYAPVIFQSSGFGSN-AALFSSF 1014
Query: 316 ITSGLNAFGSILSIYFIDKTGRKKLALIS----LCGVVLSLALLTVAFRESELHAPTVSA 371
IT+G +++S++ +DK G +K L + +C ++++ +L V F + + +SA
Sbjct: 1015ITNGALLVATVISMFLVDKFGTRKFFLEAGFEMICCMIITAVVLAVEFGHGKELSKGISA 1194
Score = 56.2 bits (134), Expect = 3e-08
Identities = 32/95 (33%), Positives = 50/95 (51%)
Frame = +1
Query: 458 YIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWT 517
+++ + G + W++ SE++PL R IA I +V+Q FL L+
Sbjct: 1219 FVLAYGRSWGPLGWLVPSELFPLEIRSAAQSIAVCVNMIFTALVAQLFL-LSLCHLKYGI 1395
Query: 518 FMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLE 552
F++FG L VV FV +PETK VP+EE+ + E
Sbjct: 1396 FLLFGGLIVVMSVFVFFLLPETKQVPIEEIYLLFE 1500
>TC87421 sugar transporter
Length = 1753
Score = 169 bits (427), Expect = 3e-42
Identities = 111/355 (31%), Positives = 182/355 (51%), Gaps = 24/355 (6%)
Frame = +2
Query: 24 PYVLRLAFSAGIGGLLFGYDTGVISGALL---YIRDDFKAVDRKTWLQEA---------- 70
P+V A +GGL+FGYD G+ G +++ F AV RK ++
Sbjct: 89 PFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQYDSQ 268
Query: 71 ----IVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGR 126
S+ A++ + V I RFGR+++++ LFL+G++I A + +L+ GR
Sbjct: 269 TLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGR 448
Query: 127 VFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWR 186
+ +GFG+G A+ PLY+SE +P + RGAL IT G ++ ++N F G W
Sbjct: 449 ILLGFGIGFANQPVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWG 628
Query: 187 W--MLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKE 244
W LG A PA+I + L LP++P + +G + +K+ LK+I EDVD E L
Sbjct: 629 WRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRGIEDVDEEFNDLVA 808
Query: 245 SVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGF 304
+ E+ ++ + L R L + + FFQQF GIN +M+Y+P + GF
Sbjct: 809 ASEASMQVENPWR-----NLLQRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIGF 973
Query: 305 ASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKL-----ALISLCGVVLSLAL 354
+ +L+ ++IT +N + +SIY +DK GR+ L A + +C V ++ A+
Sbjct: 974 KDD-ASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAI 1135
Score = 60.1 bits (144), Expect = 2e-09
Identities = 30/125 (24%), Positives = 62/125 (49%)
Frame = +2
Query: 444 PSKFGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQ 503
P + ++ + +Y+ F+ G + W++ SEI+PL R + + + +V+Q
Sbjct: 1175 PEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQ 1354
Query: 504 SFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQK 563
FL + + F+ F +V +V +PETKG+P+EE++++ + +F +
Sbjct: 1355 VFLIMLCHMKFG-LFLFFAFFVLVMSIYVFFLLPETKGIPIEEMDRVWKSHPFWSRFVEH 1531
Query: 564 RDSGS 568
D G+
Sbjct: 1532 GDHGN 1546
>TC77798 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_140
{Arabidopsis thaliana}, partial (88%)
Length = 2467
Score = 154 bits (390), Expect = 7e-38
Identities = 85/230 (36%), Positives = 139/230 (59%), Gaps = 6/230 (2%)
Frame = +3
Query: 29 LAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIGAAVGGW 88
+A +A IG L G+D I+G++LYI+ D +T ++ +V+ ++ GA + G
Sbjct: 129 VAIAASIGNFLQGWDNATIAGSILYIKKDLAL---QTTMEGLVVAMSLIGATVITTCSGP 299
Query: 89 INDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPLYISEAS 148
I+D GRR +II+ L+ +GS++M +PN +L R+ GFG+G+A P+YISE +
Sbjct: 300 ISDWLGRRPMMIISSVLYFLGSLVMLWSPNVYVLCLARLLDGFGIGLAVTLVPVYISETA 479
Query: 149 PTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPG-TWRWMLGVAAAPAIIQIVL-MLSL 206
P+ +RG+L +L F +GG FLSY + + +P +WR MLGV + P++ +L + L
Sbjct: 480 PSDIRGSLNTLPQFSGSGGMFLSYCMVFVMSLSPSPSWRIMLGVLSIPSLFYFLLTVFFL 659
Query: 207 PESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESV----ESEIEE 252
PESPRWL KG+ E+K +L+++ +DV E+ L E + ++ IEE
Sbjct: 660 PESPRWLVSKGKMLEAKKVLQRLRGQDDVSGEMALLVEGLGIGGDASIEE 809
Score = 73.2 bits (178), Expect = 3e-13
Identities = 36/105 (34%), Positives = 60/105 (56%)
Frame = +3
Query: 453 VGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTI 512
V + +Y F G G +P ++ SEI+P R RG+C I + WI ++IV+ S + ++
Sbjct: 1965 VCVVVYFCFFVMGYGPIPNILCSEIFPTRVRGLCIAICALVFWIGDIIVTYSLPVMLSSL 2144
Query: 513 GTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQ 557
G A F ++ I+ ++ FV + VPETKG+P+E + + S Q
Sbjct: 2145 GLAGVFGVYAIVCCISWVFVYLKVPETKGMPLEVITEFFSVGSKQ 2279
Score = 62.8 bits (151), Expect = 3e-10
Identities = 41/118 (34%), Positives = 67/118 (56%), Gaps = 10/118 (8%)
Frame = +3
Query: 250 IEESKT-SKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFA--- 305
I SKT SK + L V+ L G+G+Q QQF GIN V+YY+P I++ AG A
Sbjct: 1563 IHPSKTASKGPIWEALLEPGVKHALIVGIGIQLLQQFSGINGVLYYTPQILEEAGVAVLL 1742
Query: 306 ------SNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTV 357
S ++ L+S +T+ L L++ +D TGR++L L+++ +++SL +L +
Sbjct: 1743 ADLGLSSTSSSFLISAVTTLLMLPSIGLAMRLMDVTGRRQLLLVTIPVLIVSLVILVL 1916
Score = 28.5 bits (62), Expect = 7.3
Identities = 26/110 (23%), Positives = 53/110 (47%), Gaps = 1/110 (0%)
Frame = +3
Query: 97 VAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPLYISEASPTRVRGAL 156
+ I++ ++ GSV+ AA ++++ F F +G + + L SE PTRVRG
Sbjct: 1899 LVILVLGSVIDFGSVVHAAISTVCVVVY---FCFFVMGYGPIPNIL-CSEIFPTRVRGLC 2066
Query: 157 VSLNSFLI-TGGQFLSYLINLAFTNAPGTWRWMLGVAAAPAIIQIVLMLS 205
+++ + + G ++Y + + ++ LG+A + IV +S
Sbjct: 2067 IAICALVFWIGDIIVTYSLPVMLSS--------LGLAGVFGVYAIVCCIS 2192
>TC93789 similar to GP|21689841|gb|AAM67564.1 unknown protein {Arabidopsis
thaliana}, partial (17%)
Length = 668
Score = 153 bits (386), Expect = 2e-37
Identities = 79/100 (79%), Positives = 94/100 (94%), Gaps = 1/100 (1%)
Frame = +3
Query: 223 KSILKKIYAPEDVDAEIEALKESVESEIEESKTS-KISMMTLLKTTTVRRGLYAGMGLQF 281
K+IL+KIY E+VDAEI+AL+ESV E++E+++S KISMMTLLKTT+VRRGLYAGMGLQ
Sbjct: 366 KAILRKIYPAEEVDAEIQALQESVAMELKEAESSEKISMMTLLKTTSVRRGLYAGMGLQI 545
Query: 282 FQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLN 321
FQQFVGINTVMY+SPTIVQLAGFASN+TA+LLSLIT+GLN
Sbjct: 546 FQQFVGINTVMYFSPTIVQLAGFASNQTAMLLSLITAGLN 665
>TC86605 similar to PIR|T01853|T01853 probable hexose transport protein
F9D12.17 - Arabidopsis thaliana, partial (91%)
Length = 3111
Score = 98.2 bits (243), Expect(2) = 3e-37
Identities = 69/226 (30%), Positives = 108/226 (47%), Gaps = 21/226 (9%)
Frame = +2
Query: 24 PYVLRLAFSAGIGGLLFGYDTGVISGALL---YIRDDFKAVDRKTW-------------- 66
P ++ A GGL+FGYD GV G +++ F V R+T
Sbjct: 1346 PIIIISCIMAATGGLMFGYDVGVSGGVASMPPFLKKFFPTVLRQTTESDGSESNYCKYDN 1525
Query: 67 --LQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLF 124
LQ S +AG + + GRR+ ++IA F+ G + A+A N +L+
Sbjct: 1526 QGLQLFTSSLYLAGLTV-TFFASYTTRVLGRRLTMLIAGFFFIAGVSLNASAQNLLMLIV 1702
Query: 125 GRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGT 184
GRV +G G+G A+ A P+++SE +P+R+RGAL L IT G + L+N A G
Sbjct: 1703 GRVLLGCGIGFANQAVPVFLSEIAPSRIRGALNILFQLDITLGILYANLVNYATNKIKGH 1882
Query: 185 WRW--MLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKK 228
W W LG+ PA++ + + ++P L +G + KS +K
Sbjct: 1883 WGWRISLGLGGIPALLLTLGAYLVVDTPNSLIERGHLGQRKSCSEK 2020
Score = 75.9 bits (185), Expect(2) = 3e-37
Identities = 48/139 (34%), Positives = 76/139 (54%)
Frame = +3
Query: 220 EESKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGL 279
++ K++L+KI ++++ E L E+ +K K LLK R L + L
Sbjct: 1995 DKGKAVLRKIRGTDNIEPEFLELVEASRV----AKEVKHPFRNLLKRNN-RPQLVISIAL 2159
Query: 280 QFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKK 339
FQQF GIN +M+Y+P + GF N AL ++IT +N +I+SIY +DK GR+K
Sbjct: 2160 MIFQQFTGINAIMFYAPVLFNTLGF-KNDAALYSAVITGAINVISTIVSIYSVDKLGRRK 2336
Query: 340 LALISLCGVVLSLALLTVA 358
L L + GV + L+ + +A
Sbjct: 2337 LLLEA--GVQMLLSQMVIA 2387
Score = 55.1 bits (131), Expect = 7e-08
Identities = 36/118 (30%), Positives = 61/118 (51%), Gaps = 2/118 (1%)
Frame = +3
Query: 448 GWAALVGL--ALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSF 505
G+AALV + +++ F+ G + W+I SEI+PL R + ++ +++Q+F
Sbjct: 2436 GYAALVVVMVCIFVSAFAWSWGPLAWLIPSEIFPLETRSAGQSVTVCVNFLFTAVIAQAF 2615
Query: 506 LSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQK 563
LS+ F G + ++ F V VPETK VP+EE M ++ Q FW++
Sbjct: 2616 LSMLCYFKFGIFFFFSGWILFMSTF-VFFLVPETKNVPIEE---MTQRVWKQHWFWKR 2777
>TC78543 similar to PIR|T14545|T14545 probable sugar transporter protein -
beet, partial (93%)
Length = 1653
Score = 151 bits (381), Expect = 7e-37
Identities = 103/335 (30%), Positives = 171/335 (50%), Gaps = 1/335 (0%)
Frame = +1
Query: 26 VLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIGAAV 85
V+ +G + FG+ G S I D + L S + GA++GA
Sbjct: 148 VIACVLIVALGPIQFGFTAGYTSPTQSAIITDLGLSVSEFSL---FGSLSNVGAMVGAIA 318
Query: 86 GGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPLYIS 145
G I + GR+ +++IA +IG ++++ A + + L GR+ GFGVG+ S P+YI+
Sbjct: 319 SGQIAEYMGRKGSLMIASIPNIIGWLMISFANDSSFLYMGRLLEGFGVGIISYTVPVYIA 498
Query: 146 EASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAAPAIIQIVLMLS 205
E SP +RG+LVS+N +T G L+YL+ L WR++ + P + I +
Sbjct: 499 EISPQNLRGSLVSVNQLSVTLGIMLAYLLGLFV-----EWRFLAILGIIPCTLLIPGLFF 663
Query: 206 LPESPRWLYRKGREEESKSILKKIYAPE-DVDAEIEALKESVESEIEESKTSKISMMTLL 264
+PESPRWL + G EE ++ L+ + E D+ E+ +K +V S + + L
Sbjct: 664 IPESPRWLAKMGMTEEFENSLQVLRGFETDISVEVNEIKTAVASANRRTTV----RFSEL 831
Query: 265 KTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFG 324
K L G+GL QQ GIN V++YS TI Q AG +S+ A + +
Sbjct: 832 KQRRYWLPLMIGIGLLVLQQLSGINGVLFYSSTIFQNAGISSSDVA---TFGVGAVQVLA 1002
Query: 325 SILSIYFIDKTGRKKLALISLCGVVLSLALLTVAF 359
+ L+++ DK+GR+ L ++S + LSL +++++F
Sbjct: 1003TTLTLWLADKSGRRLLLIVSSSAMTLSLLVVSISF 1107
Score = 70.9 bits (172), Expect = 1e-12
Identities = 31/99 (31%), Positives = 60/99 (60%)
Frame = +1
Query: 451 ALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQ 510
++ G+ + +I FS GMG +PW+I SEI P+ +G+ G A+ W + +++ + +L
Sbjct: 1168 SVAGVVVMVIAFSLGMGAMPWIIMSEILPINIKGLAGSFATLANWFFSWLITLT-ANLLL 1344
Query: 511 TIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEK 549
+ TF ++ ++ + FV ++VPETKG +EE+++
Sbjct: 1345 DWSSGGTFTIYTVVCAFTVGFVAIWVPETKGKTLEEIQQ 1461
>TC83667 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (26%)
Length = 454
Score = 150 bits (380), Expect = 1e-36
Identities = 82/152 (53%), Positives = 102/152 (66%)
Frame = +3
Query: 203 MLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKISMMT 262
ML LPESPRWLY RE E+ +L KIY + ++ E+ L + +SE + K + +
Sbjct: 3 MLFLPESPRWLYINNRENEAIIVLGKIYDFDRLEDEVALL--TAQSEQDRQKRADVRYRH 176
Query: 263 LLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNA 322
+ K+ +R AG GLQ FQQF GINTVMYYSPTIVQ+AGF SN AL LSLI +GLNA
Sbjct: 177 VFKSKEIRLAFLAGAGLQAFQQFTGINTVMYYSPTIVQMAGFHSNELALQLSLIVAGLNA 356
Query: 323 FGSILSIYFIDKTGRKKLALISLCGVVLSLAL 354
G++L IY ID GRKKLAL SL GV+ SL +
Sbjct: 357 AGTVLGIYLIDHAGRKKLALYSLGGVIASLII 452
>TC79692 similar to GP|20453189|gb|AAM19835.1 AT4g35300/F23E12_140
{Arabidopsis thaliana}, partial (49%)
Length = 1353
Score = 149 bits (376), Expect = 3e-36
Identities = 83/198 (41%), Positives = 124/198 (61%), Gaps = 3/198 (1%)
Frame = +2
Query: 29 LAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIGAAVGGW 88
+A +A IG LL G+D I+G++LYI+ +F+ T ++ IV+ ++ GA + G
Sbjct: 170 VAVAAAIGNLLQGWDNATIAGSILYIKREFQLQSEPT-VEGLIVAMSLIGATVVTTCSGA 346
Query: 89 INDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPLYISEAS 148
++D FGRR +II+ L+ + S++M +PN ILLF R+ G G+G+A PLYISE +
Sbjct: 347 LSDLFGRRPMLIISSLLYFLSSLVMFWSPNVYILLFARLLDGLGIGLAVTLVPLYISEIA 526
Query: 149 PTRVRGALVSLNSFLITGGQFLSY--LINLAFTNAPGTWRWMLGVAAAPAIIQIVL-MLS 205
P +RG+L +L F + G F SY + ++ T AP +WR MLGV + P++I L +L
Sbjct: 527 PPEIRGSLNTLPQFAGSAGMFFSYCMVFGMSLTKAP-SWRLMLGVLSIPSLIYFALTLLV 703
Query: 206 LPESPRWLYRKGREEESK 223
LPESPRWL KGR E+K
Sbjct: 704 LPESPRWLVSKGRMLEAK 757
>TC86673 weakly similar to PIR|S25015|S25015 monosaccharide transport
protein MST1 - common tobacco, partial (65%)
Length = 1162
Score = 108 bits (269), Expect(2) = 8e-36
Identities = 74/220 (33%), Positives = 111/220 (49%), Gaps = 22/220 (10%)
Frame = +2
Query: 26 VLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEA--------------- 70
V+ A GGL+FGYD GV SG + + K + QE+
Sbjct: 95 VIITCIMAATGGLIFGYDHGV-SGGVTSMDSFLKRFFPSVYEQESNLKPSANQYCKFNSQ 271
Query: 71 IVSTAIAGAIIGAAVGGW----INDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGR 126
I++ + I A V G I GRR +I+ F+ G+++ A N A+L+ GR
Sbjct: 272 ILTLFTSSLYISALVAGLGASSITRALGRRTTMILGGIFFVSGALLNGLAMNIAMLIAGR 451
Query: 127 VFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFT---NAPG 183
+ +GFG+G A+ A P+Y+SE +P + RGAL IT G F + + N F+ N G
Sbjct: 452 LLLGFGIGCANQAVPIYLSEMAPYKYRGALNMCFQLSITIGIFTANMFNYYFSKILNGEG 631
Query: 184 TWRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESK 223
WR LG+ A PA++ I+ + LP+SP L +GR EE++
Sbjct: 632 -WRLSLGLGAVPAVVFIIGSICLPDSPNSLVTRGRHEEAR 748
Score = 60.8 bits (146), Expect(2) = 8e-36
Identities = 38/131 (29%), Positives = 67/131 (51%)
Frame = +3
Query: 228 KIYAPEDVDAEIEALKESVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVG 287
KI +DVDAE + + E+ + K L+ R L + + FFQQF G
Sbjct: 762 KIRGTDDVDAEFRDIVAASEASAKVKHPWKT-----LQERKYRPQLVFAILIPFFQQFTG 926
Query: 288 INTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCG 347
+N + +Y+P + + GF S + +L+ + I +++SI+ +DK GR+ L L
Sbjct: 927 LNVITFYAPILFRTIGFGS-QASLMSAAIIGSFKPVSTLVSIFVVDKFGRRALFLEGGVQ 1103
Query: 348 VVLSLALLTVA 358
+++S ++TVA
Sbjct: 1104MLISXIIMTVA 1136
Score = 29.6 bits (65), Expect = 3.3
Identities = 16/51 (31%), Positives = 26/51 (50%), Gaps = 2/51 (3%)
Frame = +3
Query: 68 QEAIVSTAIAGAI--IGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAA 116
Q +++S AI G+ + V ++ D+FGRR + LI +IM A
Sbjct: 984 QASLMSAAIIGSFKPVSTLVSIFVVDKFGRRALFLEGGVQMLISXIIMTVA 1136
>TC89747 similar to PIR|F96696|F96696 protein F1N21.12 [imported] -
Arabidopsis thaliana, partial (64%)
Length = 1372
Score = 147 bits (370), Expect = 1e-35
Identities = 109/343 (31%), Positives = 164/343 (47%), Gaps = 2/343 (0%)
Frame = +3
Query: 14 RECLSLSWKN--PYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAI 71
+E + SWK P+VL A I LFGY GV++ L I D + T + +
Sbjct: 342 KETTNPSWKLSLPHVL----VATITSFLFGYHLGVVNEPLESISVDL-GFNGNTLAEGLV 506
Query: 72 VSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGF 131
VS + GA+ G + GWI D GRR A + +IG+ + AA N +L GR+FVG
Sbjct: 507 VSICLGGALFGCLLSGWIADAVGRRRAFQLCALPMIIGAAMSAATNNLFGMLVGRLFVGT 686
Query: 132 GVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGV 191
G+G+ + LY++E SP VRG +L G S I + G WR V
Sbjct: 687 GLGLGPPVAALYVTEVSPAFVRGTYGALIQIATCFGILGSLFIGIPVKEISGWWRVCFWV 866
Query: 192 AAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIE 251
+ PA I + M ESP WLY++GR E+++ +++ + + L S E
Sbjct: 867 STIPAAILALAMDFCAESPHWLYKQGRTAEAEAEFERLLGVSEAKFAMSQL--SKVDRGE 1040
Query: 252 ESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTAL 311
++ T K S + + V ++ G L QQ GIN V Y+S T+ + AG S+ +
Sbjct: 1041DTDTVKFSELLHGHHSKV---VFIGSTLFALQQLSGINAVFYFSSTVFKSAGVPSDFANV 1211
Query: 312 LLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLAL 354
+ + N GSI+S +DK GRK L S G+ +S+ +
Sbjct: 1212CIGV----ANLTGSIISTGLMDKLGRKVLLFWSFFGMAISMII 1328
>TC87126 similar to GP|15724240|gb|AAL06513.1 At1g75220/F22H5_6 {Arabidopsis
thaliana}, partial (79%)
Length = 1524
Score = 139 bits (350), Expect = 3e-33
Identities = 96/286 (33%), Positives = 155/286 (53%), Gaps = 4/286 (1%)
Frame = +3
Query: 78 GAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMAS 137
GA++GA G I + GR+ +++IA +IG + ++ A + + L GR+ GFGVG+ S
Sbjct: 6 GAMVGAIASGQIAEYVGRKGSLMIASIPNIIGWLAISFAKDSSFLFMGRLLEGFGVGIIS 185
Query: 138 MASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAAPAI 197
P+YI+E +P +RG+L S+N +T G L+YL+ L F N WR + + P
Sbjct: 186 YVVPVYIAEIAPENMRGSLGSVNQLSVTIGIMLAYLLGL-FAN----WRVLAILGILPCT 350
Query: 198 IQIVLMLSLPESPRWLYRKGREEESKSILKKIYA-PEDVDAEIEALKESVESEIEESKTS 256
+ I + +PESPRWL + G EE ++ L+ + D+ E+ +K++V S K +
Sbjct: 351 VLIPGLFFIPESPRWLAKMGMMEEFETSLQVLRGFDTDISVEVHEIKKAVAS---NGKRA 521
Query: 257 KISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLI 316
I L+ L G+GL QQ GIN V++YS +I AG +S+ A
Sbjct: 522 TIRFAD-LQRKRYWFPLSVGIGLLVLQQLSGINGVLFYSTSIFANAGISSSNAA------ 680
Query: 317 TSGLNAFGSI---LSIYFIDKTGRKKLALISLCGVVLSLALLTVAF 359
T GL A I ++ + +DK+GR+ L +IS + SL ++++AF
Sbjct: 681 TVGLGAIQVIATGVATWLVDKSGRRVLLIISSSLMTASLLVVSIAF 818
Score = 75.1 bits (183), Expect = 7e-14
Identities = 36/104 (34%), Positives = 64/104 (60%)
Frame = +3
Query: 445 SKFGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQS 504
S G ++VGL + +I FS G+G +PW+I SEI P+ +G+ G A+ W+ I++ +
Sbjct: 858 SILGIISVVGLVVMVIGFSLGLGPIPWLIMSEILPVNIKGLAGSTATMANWLVAWIITMT 1037
Query: 505 FLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVE 548
+L T + TF+++ ++A + F ++VPETKG +EE++
Sbjct: 1038-ANLLLTWSSGGTFLIYTVVAAFTVVFTSLWVPETKGRTLEEIQ 1166
>TC90639 similar to GP|17380890|gb|AAL36257.1 putative membrane transporter
protein {Arabidopsis thaliana}, partial (21%)
Length = 555
Score = 124 bits (311), Expect = 1e-28
Identities = 58/99 (58%), Positives = 77/99 (77%)
Frame = +2
Query: 457 LYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAW 516
LYI FSPGMG VP INSEIYP YRG+CGG+A+T WISNLIVS+SFLS+ IG A
Sbjct: 5 LYIGFFSPGMGPVPGTINSEIYPEEYRGICGGMAATVCWISNLIVSESFLSIADAIGIAS 184
Query: 517 TFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRS 555
TF++ ++AVVA FV+++VPET+G+ +EVE + ++R+
Sbjct: 185 TFLIIAVIAVVAFLFVLLYVPETQGLTFDEVELIWKERA 301
>TC82331 weakly similar to EGAD|120604|128852 hypothetical protein M01F1.5
{Caenorhabditis elegans}, partial (10%)
Length = 848
Score = 115 bits (288), Expect = 5e-26
Identities = 75/203 (36%), Positives = 110/203 (53%), Gaps = 16/203 (7%)
Frame = +3
Query: 27 LRLAFSAGIGGLLFGYDTGVISGALLYIRDDFK---------AVDRKTW---LQEAIVST 74
L L IGG LFGYDTG IS LL+ DF+ A KTW +Q +VS
Sbjct: 243 LVLGVVTSIGGFLFGYDTGQISSMLLFT--DFRQRFATGPEDANGVKTWVSIIQSLMVSL 416
Query: 75 AIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAI-LLFGRVFVGFGV 133
G +IGA G + D +GRR ++ +F+IG+++ A + + ++ GR G GV
Sbjct: 417 MSIGTLIGALSGAYTADWWGRRKSLSFGVVVFVIGNIVQITAMDTWVHMMMGRFIAGLGV 596
Query: 134 GMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN---APGTWRWMLG 190
G S+ P++ SE SP +RGA+V+ +IT G +S LIN N + +WR ++G
Sbjct: 597 GNLSVGVPMFQSECSPKEIRGAVVASYQLMITFGILISNLINFGVRNIQESDASWRIVIG 776
Query: 191 VAAAPAIIQIVLMLSLPESPRWL 213
+ A A+ + +L +PESPRWL
Sbjct: 777 LGIAFAMPLGLGILVVPESPRWL 845
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.138 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,997,343
Number of Sequences: 36976
Number of extensions: 261983
Number of successful extensions: 1515
Number of sequences better than 10.0: 116
Number of HSP's better than 10.0 without gapping: 1405
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1437
length of query: 571
length of database: 9,014,727
effective HSP length: 101
effective length of query: 470
effective length of database: 5,280,151
effective search space: 2481670970
effective search space used: 2481670970
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0140.7


