

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0140.2
(123 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
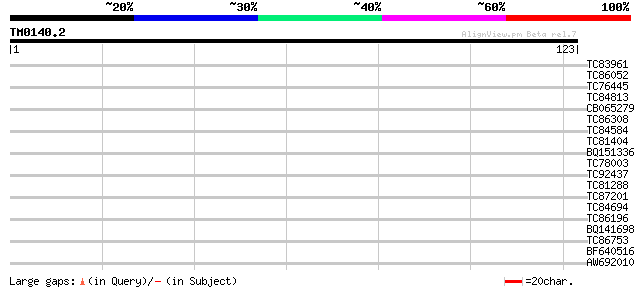
Score E
Sequences producing significant alignments: (bits) Value
TC83961 similar to GP|17979121|gb|AAL49818.1 unknown protein {Ar... 32 0.061
TC86052 similar to GP|3641865|emb|CAA09457.1 beta-galactosidase ... 29 0.52
TC76445 similar to PIR|T07623|T07623 extensin homolog HRGP2 - so... 28 0.89
TC84813 similar to PIR|T47503|T47503 hypothetical protein F9K21.... 28 1.2
CB065279 weakly similar to GP|17983777|gb| TRANSCRIPTIONAL REGUL... 27 1.5
TC86308 similar to SP|O48927|CP78_SOYBN Cytochrome P450 78A3 (EC... 27 1.5
TC84584 similar to PIR|C84547|C84547 hypothetical protein At2g17... 27 2.6
TC81404 similar to GP|16323167|gb|AAL15318.1 At1g30760/T5I8_22 {... 27 2.6
BQ151336 27 2.6
TC78003 similar to GP|20146093|dbj|BAB88935. glucosyltransferase... 27 2.6
TC92437 homologue to PIR|T06460|T06460 anthranilate phosphoribos... 26 3.4
TC81288 similar to GP|9945085|gb|AAG03122.1| F5A9.21 {Arabidopsi... 26 3.4
TC87201 similar to PIR|F86388|F86388 hypothetical protein AAG292... 26 3.4
TC84694 26 3.4
TC86196 similar to GP|22652125|gb|AAN03626.1 BEL1-related homeot... 26 3.4
BQ141698 similar to GP|7303636|gb|A CG13214-PA {Drosophila melan... 26 4.4
TC86753 homologue to PIR|C96633|C96633 probable Serine/Threonine... 26 4.4
BF640516 similar to GP|17065312|gb Unknown protein {Arabidopsis ... 26 4.4
AW692010 similar to GP|22335707|db nine-cis-epoxycarotenoid diox... 22 5.1
>TC83961 similar to GP|17979121|gb|AAL49818.1 unknown protein {Arabidopsis
thaliana}, partial (15%)
Length = 685
Score = 32.0 bits (71), Expect = 0.061
Identities = 19/63 (30%), Positives = 28/63 (44%), Gaps = 13/63 (20%)
Frame = +3
Query: 35 YQPMLVCH--LDNLMPRTPCVIMIA---------STPPSQHHHSCKPTVNRAP--GWFPE 81
+ P L H L +P PC + + S+PP +HHH P +R WF +
Sbjct: 99 FPPFL*THNPLPTPIPLNPCFLFFSPQ*HQRSSSSSPPQKHHHQHHPNGSRRE*LRWFQQ 278
Query: 82 KRG 84
K+G
Sbjct: 279 KKG 287
>TC86052 similar to GP|3641865|emb|CAA09457.1 beta-galactosidase {Cicer
arietinum}, partial (94%)
Length = 2539
Score = 28.9 bits (63), Expect = 0.52
Identities = 17/54 (31%), Positives = 24/54 (43%)
Frame = -2
Query: 46 LMPRTPCVIMIASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEPPKTALHNVTL 99
L P I++ TPP+ H C+PT R ++ + P T NVTL
Sbjct: 1791 LSPHLDPSILLGLTPPTFHVSKCRPTFGRPTATLSKEILLLPTFRFTLSLNVTL 1630
>TC76445 similar to PIR|T07623|T07623 extensin homolog HRGP2 - soybean
(fragment), partial (76%)
Length = 821
Score = 28.1 bits (61), Expect = 0.89
Identities = 14/37 (37%), Positives = 21/37 (55%)
Frame = +3
Query: 29 PHILHLYQPMLVCHLDNLMPRTPCVIMIASTPPSQHH 65
PH LHL+ L+ + L+P P +I+ +T P HH
Sbjct: 180 PH-LHLHLHHLLITIKVLLPHLPFLILPTTTNPLHHH 287
>TC84813 similar to PIR|T47503|T47503 hypothetical protein F9K21.210 -
Arabidopsis thaliana, partial (15%)
Length = 682
Score = 27.7 bits (60), Expect = 1.2
Identities = 16/48 (33%), Positives = 20/48 (41%), Gaps = 8/48 (16%)
Frame = -3
Query: 30 HILHLYQPMLVCHLDN--------LMPRTPCVIMIASTPPSQHHHSCK 69
HIL Q CH +N P PC + A+T H+H CK
Sbjct: 431 HILPPSQQSFPCHKENDTQGIAHPYHPSQPCPLCGANTIHRSHNHICK 288
>CB065279 weakly similar to GP|17983777|gb| TRANSCRIPTIONAL REGULATOR
{Brucella melitensis}, partial (18%)
Length = 617
Score = 27.3 bits (59), Expect = 1.5
Identities = 17/49 (34%), Positives = 21/49 (42%), Gaps = 2/49 (4%)
Frame = +1
Query: 47 MPRTPCVIMIASTPPSQHHHSCKPTVNR--APGWFPEKRGVFPEPPKTA 93
MP +PC + P H+C+PT N AP G P P TA
Sbjct: 292 MPASPCCSAMPMASP--WTHACQPTDNATPAPDCASAPAGTSPSPAPTA 432
>TC86308 similar to SP|O48927|CP78_SOYBN Cytochrome P450 78A3 (EC 1.14.-.-).
[Soybean] {Glycine max}, partial (88%)
Length = 2078
Score = 27.3 bits (59), Expect = 1.5
Identities = 28/77 (36%), Positives = 32/77 (41%), Gaps = 16/77 (20%)
Frame = -3
Query: 57 ASTPPSQHHHSCKPTV---NRAPGWFPEK---------RGVFPEPPKTA----LHNVTLL 100
AS PP HS PTV RA P+K RG F P +T L V
Sbjct: 1623 ASAPPLNQLHSLHPTVTIQTRARV*LPKKSR*LNLNSSRGNFSYPIRTGPT*DLIQVLKT 1444
Query: 101 QHLRNTTQSDIHRTGSS 117
Q LR+ T D + GSS
Sbjct: 1443 QLLRSQTSPD*IQVGSS 1393
>TC84584 similar to PIR|C84547|C84547 hypothetical protein At2g17030
[imported] - Arabidopsis thaliana, partial (4%)
Length = 812
Score = 26.6 bits (57), Expect = 2.6
Identities = 9/18 (50%), Positives = 12/18 (66%)
Frame = +3
Query: 59 TPPSQHHHSCKPTVNRAP 76
TPP+ HHH + T+N P
Sbjct: 270 TPPTNHHHHQQQTLNNNP 323
>TC81404 similar to GP|16323167|gb|AAL15318.1 At1g30760/T5I8_22 {Arabidopsis
thaliana}, partial (53%)
Length = 1205
Score = 26.6 bits (57), Expect = 2.6
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = -3
Query: 79 FPEKRGVFPEPPKTALHNVTLLQHLRNTTQ 108
FP K V E KT H++ +LQH ++ TQ
Sbjct: 1155 FPHKDNVKLEMFKTEKHSIIILQH*QHQTQ 1066
>BQ151336
Length = 1045
Score = 26.6 bits (57), Expect = 2.6
Identities = 21/67 (31%), Positives = 26/67 (38%)
Frame = +3
Query: 6 GSSSAIDITTYLASMSTNARTLSPHILHLYQPMLVCHLDNLMPRTPCVIMIASTPPSQHH 65
GSS A+ M T+ RT P LY C D+ P + S S H
Sbjct: 699 GSSVAVARRHVARIMETHTRTRDP*---LYSDAHPCSRDSPAP-------LFSHTESPHT 848
Query: 66 HSCKPTV 72
H C PT+
Sbjct: 849 HECNPTI 869
>TC78003 similar to GP|20146093|dbj|BAB88935. glucosyltransferase NTGT2
{Nicotiana tabacum}, partial (50%)
Length = 1740
Score = 26.6 bits (57), Expect = 2.6
Identities = 22/87 (25%), Positives = 31/87 (35%)
Frame = -2
Query: 19 SMSTNARTLSPHILHLYQPMLVCHLDNLMPRTPCVIMIASTPPSQHHHSCKPTVNRAPGW 78
S S N +P IL L C L+P S + HHH P P +
Sbjct: 1541 SSSNNPIQKTPQILF*RATFLHCFSSQLLPFLSTPSQFLSFLSTPHHHLQTP-----PNF 1377
Query: 79 FPEKRGVFPEPPKTALHNVTLLQHLRN 105
P +F +H+ + L H+ N
Sbjct: 1376 IPSHNSIFIH---FFIHSYSYLPHIFN 1305
>TC92437 homologue to PIR|T06460|T06460 anthranilate
phosphoribosyltransferase (EC 2.4.2.18) - garden pea
(fragment), partial (73%)
Length = 815
Score = 26.2 bits (56), Expect = 3.4
Identities = 18/73 (24%), Positives = 32/73 (43%)
Frame = +3
Query: 1 MLAFSGSSSAIDITTYLASMSTNARTLSPHILHLYQPMLVCHLDNLMPRTPCVIMIASTP 60
M FSG S + +L +ST ++ ++H+ MLVC + +MP + +
Sbjct: 510 MTVFSGFLS---VGRWLGEVSTWNHPMTTVLVHILFVMLVCFPELIMPTMFLYVFVIGMW 680
Query: 61 PSQHHHSCKPTVN 73
+ C P +N
Sbjct: 681 NWRFRPRCPPHMN 719
>TC81288 similar to GP|9945085|gb|AAG03122.1| F5A9.21 {Arabidopsis
thaliana}, partial (1%)
Length = 720
Score = 26.2 bits (56), Expect = 3.4
Identities = 10/15 (66%), Positives = 11/15 (72%)
Frame = +1
Query: 28 SPHILHLYQPMLVCH 42
S H LHL P+LVCH
Sbjct: 181 STHQLHLLSPVLVCH 225
>TC87201 similar to PIR|F86388|F86388 hypothetical protein AAG29221.1
[imported] - Arabidopsis thaliana, partial (41%)
Length = 991
Score = 26.2 bits (56), Expect = 3.4
Identities = 16/57 (28%), Positives = 26/57 (45%)
Frame = +2
Query: 20 MSTNARTLSPHILHLYQPMLVCHLDNLMPRTPCVIMIASTPPSQHHHSCKPTVNRAP 76
MST+ R H+LH++ L+ H + P ++ ++ HHH T N P
Sbjct: 386 MSTSHR----HLLHMFTNHLLLHNMSTNPLHHHLMFMSHLLHPHHHHLLHTTTNLYP 544
>TC84694
Length = 907
Score = 26.2 bits (56), Expect = 3.4
Identities = 18/53 (33%), Positives = 25/53 (46%), Gaps = 3/53 (5%)
Frame = -2
Query: 5 SGSSSAIDITTYLASMSTNARTL---SPHILHLYQPMLVCHLDNLMPRTPCVI 54
S + A D LA S N +TL +PH L+L + + H L TPC +
Sbjct: 324 SQMNEARDFEKLLAVHSQNDQTLQQSAPHHLYLQKYQVHRHFSYLQITTPCYL 166
>TC86196 similar to GP|22652125|gb|AAN03626.1 BEL1-related homeotic protein
29 {Solanum tuberosum}, partial (53%)
Length = 2227
Score = 26.2 bits (56), Expect = 3.4
Identities = 13/38 (34%), Positives = 20/38 (52%)
Frame = -1
Query: 30 HILHLYQPMLVCHLDNLMPRTPCVIMIASTPPSQHHHS 67
H+LH PM++ + +PC S+ P QHH+S
Sbjct: 895 HLLHYQLPMILQY-------SPCSFHHHSSYPIQHHYS 803
>BQ141698 similar to GP|7303636|gb|A CG13214-PA {Drosophila melanogaster},
partial (4%)
Length = 1180
Score = 25.8 bits (55), Expect = 4.4
Identities = 10/26 (38%), Positives = 14/26 (53%)
Frame = -1
Query: 50 TPCVIMIASTPPSQHHHSCKPTVNRA 75
T C + S+PPS +HH+ P A
Sbjct: 583 THCTLSPLSSPPSNNHHAPPPPTQSA 506
>TC86753 homologue to PIR|C96633|C96633 probable Serine/Threonine protein
kinase F8A5.29 [imported] - Arabidopsis thaliana,
complete
Length = 1624
Score = 25.8 bits (55), Expect = 4.4
Identities = 13/41 (31%), Positives = 21/41 (50%)
Frame = -1
Query: 24 ARTLSPHILHLYQPMLVCHLDNLMPRTPCVIMIASTPPSQH 64
+R+L P L+L ++ CHL + M +S+ P QH
Sbjct: 964 SRSLFPWSLYLVPLVIFCHLSQVYTLACHSNMFSSSVPQQH 842
>BF640516 similar to GP|17065312|gb Unknown protein {Arabidopsis thaliana},
partial (30%)
Length = 659
Score = 25.8 bits (55), Expect = 4.4
Identities = 17/60 (28%), Positives = 27/60 (44%)
Frame = +2
Query: 48 PRTPCVIMIASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEPPKTALHNVTLLQHLRNTT 107
P+TP + I T HHH P + R R + P P + +V LL++ ++ T
Sbjct: 92 PQTPPQLQILQTHHPPHHH---PFLRRPLRHHRRVRRLLPSPWSSLRVSVQLLRNSKSKT 262
>AW692010 similar to GP|22335707|db nine-cis-epoxycarotenoid dioxygenase4
{Pisum sativum}, partial (21%)
Length = 653
Score = 22.3 bits (46), Expect(2) = 5.1
Identities = 9/24 (37%), Positives = 15/24 (62%)
Frame = +1
Query: 14 TTYLASMSTNARTLSPHILHLYQP 37
TT + ++STN + H HL++P
Sbjct: 220 TTNIINISTNTQHTR*HYQHLHRP 291
Score = 21.6 bits (44), Expect(2) = 5.1
Identities = 8/16 (50%), Positives = 11/16 (68%)
Frame = +2
Query: 47 MPRTPCVIMIASTPPS 62
+P T CVI+ + PPS
Sbjct: 359 LPPTQCVIIKGTLPPS 406
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.131 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,835,768
Number of Sequences: 36976
Number of extensions: 85906
Number of successful extensions: 661
Number of sequences better than 10.0: 75
Number of HSP's better than 10.0 without gapping: 654
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 660
length of query: 123
length of database: 9,014,727
effective HSP length: 84
effective length of query: 39
effective length of database: 5,908,743
effective search space: 230440977
effective search space used: 230440977
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0140.2


