

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0140.12
(414 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
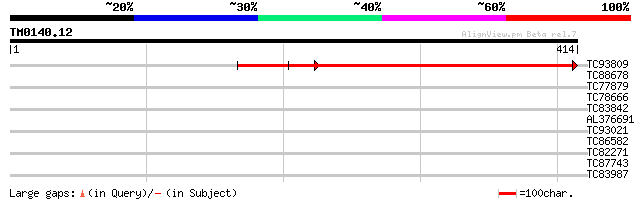
Score E
Sequences producing significant alignments: (bits) Value
TC93809 similar to PIR|H86428|H86428 hypothetical protein AAF197... 397 e-111
TC88678 similar to PIR|S51171|S51171 amino acid transport protei... 35 0.053
TC77879 similar to PIR|D86442|D86442 probable amino acid permeas... 35 0.069
TC78666 homologue to GP|6850938|emb|CAB71135.1 putative imbibiti... 32 0.59
TC83842 similar to PIR|S51171|S51171 amino acid transport protei... 30 1.3
AL376691 28 5.0
TC93021 homologue to GP|3413719|gb|AAC31242.1| unknown protein {... 28 6.5
TC86582 similar to GP|21593999|gb|AAM65917.1 ATP phosphoribosyl ... 28 6.5
TC82271 similar to GP|18491137|gb|AAL69537.1 AT5g45420/MFC19_9 {... 28 6.5
TC87743 similar to GP|15810581|gb|AAL07178.1 putative 2-oxogluta... 28 8.5
TC83987 PIR|G85579|G85579 probable membrane pump protein ybhI [i... 23 9.3
>TC93809 similar to PIR|H86428|H86428 hypothetical protein AAF19744.1
[imported] - Arabidopsis thaliana, partial (25%)
Length = 868
Score = 397 bits (1021), Expect = e-111
Identities = 192/211 (90%), Positives = 206/211 (96%)
Frame = +2
Query: 204 GIPEPDGSVSWNFNALVGLFFPAVTGIMAGSNRSSSLKDTQRSIPLGTLAATLVTTFMYL 263
GIPEPDGSV+WNFN+LVGLFFPAVTGIMAGSNRSSSL+DTQRSIP+GTL+ATL T+FMYL
Sbjct: 113 GIPEPDGSVTWNFNSLVGLFFPAVTGIMAGSNRSSSLRDTQRSIPVGTLSATLSTSFMYL 292
Query: 264 VSVIMFGALATREKLLTDRLLTATVAWPFPSLIKIGIILSTMGAALQSLTGAPRLLAAIA 323
+SVI+FGA+ATR+KLLTDRLLTAT+AWP PSLIKIGIILSTMGAALQSLTGAPRLLAAIA
Sbjct: 293 ISVILFGAVATRDKLLTDRLLTATIAWPLPSLIKIGIILSTMGAALQSLTGAPRLLAAIA 472
Query: 324 NDDILPILKYFKVADGSEPHVATLFTAFLCSGCVVIGNLDLITPTVTMFFLLCYAGVNLS 383
NDDILPIL YFKVADGSEPH+ATLFTA LC GCVVIGNLDLITPTVTMFFLLCY+GVNLS
Sbjct: 473 NDDILPILNYFKVADGSEPHIATLFTALLCIGCVVIGNLDLITPTVTMFFLLCYSGVNLS 652
Query: 384 CFLLDLLDAPSWRPRWKFHHWSLSLVGALLC 414
CFLLDLLDAPSWRPRWKFHHWSLSL+GALLC
Sbjct: 653 CFLLDLLDAPSWRPRWKFHHWSLSLLGALLC 745
Score = 68.2 bits (165), Expect = 6e-12
Identities = 32/60 (53%), Positives = 40/60 (66%)
Frame = +1
Query: 167 LGILLAREDHPAEGITGLSLETLKDNWGSEYQKTNDAGIPEPDGSVSWNFNALVGLFFPA 226
+G+L A++DHP EGITGLS ETLK+NW S+YQKTNDA + F G FFP+
Sbjct: 1 VGVLFAKKDHPTEGITGLSFETLKENWSSDYQKTNDAWNSRT*WFCNLEFQFFGGPFFPS 180
>TC88678 similar to PIR|S51171|S51171 amino acid transport protein AAT1 -
Arabidopsis thaliana, partial (72%)
Length = 1520
Score = 35.0 bits (79), Expect = 0.053
Identities = 38/169 (22%), Positives = 70/169 (40%), Gaps = 7/169 (4%)
Frame = +3
Query: 222 LFFPAVTGIMAGSNRSSSLKDTQRSIPLGTLAATLVTTFMYLVSVIMFGALATREKLLTD 281
LFF A G A S + K+ R IP+G + + +TT +Y + + ++L TD
Sbjct: 858 LFF-AYIGFDAVSTMAEETKNPGRDIPIGLVGSMTITTAIYCLLGATLCLMQNYKELDTD 1034
Query: 282 RLLTATVA-----WPFPSLIKIGIILSTMGAALQSLTGAPRLLAAIANDDILPILKYFKV 336
+ + W ++ +G + L S G R L IA ++P +F +
Sbjct: 1035 APFSVAFSAVGMDWA-KYIVSLGALKGMTTVLLVSAVGQARYLTHIARTHMMP--PWFAL 1205
Query: 337 AD--GSEPHVATLFTAFLCSGCVVIGNLDLITPTVTMFFLLCYAGVNLS 383
D P AT+ + NL +++ +++ L ++ V L+
Sbjct: 1206 VDERTGTPMNATISMLIATAIVAFFTNLSILSSLLSISTLFIFSLVALA 1352
>TC77879 similar to PIR|D86442|D86442 probable amino acid permease
[imported] - Arabidopsis thaliana, partial (81%)
Length = 2037
Score = 34.7 bits (78), Expect = 0.069
Identities = 66/328 (20%), Positives = 124/328 (37%), Gaps = 7/328 (2%)
Frame = +2
Query: 56 IGRALGPEVGVSIGLCFFLGNAVAGALYVLGAVETFLKAVPAAGIFRETITQVNGTTIAQ 115
+ ALGP G G +L + ALY + ++ AVPA G + G
Sbjct: 536 VSSALGPFWGFQQGWMKWLSGVIDNALYPVLFLDYLKSAVPAVGGGLPRVFATWG----- 700
Query: 116 PIESPSSHDLQIYGIVVTIVLCFIVFGGVKMINRVAPAFLIPVLFSLI-CIYLGILLARE 174
+TIVL ++ + G+ ++ VA I FSL+ +++G L +
Sbjct: 701 ----------------LTIVLTYLNYRGLTIVGLVAVCLGI---FSLLPFVFMGFLSIPD 823
Query: 175 DHPAEGITGLSLETLKDNWGSEYQKTNDAGIPEPDGSVSWNFNALVGLFFPAVTGIMAGS 234
P + W E ND ++ WN N ++ ++ S
Sbjct: 824 MKP-------------ERWFVE-TNLNDVDWNLYLNTLFWNLN-----YWDSI------S 928
Query: 235 NRSSSLKDTQRSIPLGTLAATLVTTFMYLVSVIM-FGALATREKLLTDRLLTATV----- 288
+ +++ ++++P G A ++ Y +++ GA+ + +L TD +
Sbjct: 929 TLAGEVENPKKNLPKGLFYALILVVVAYFFPLLIGTGAVPVQRELWTDGYFSEIAMIIGG 1108
Query: 289 AWPFPSLIKIGIILSTMGAALQSLTGAPRLLAAIANDDILPILKYFKVADGSEPHVATLF 348
W ++ +S MG + ++ L +A +LP + K + P + LF
Sbjct: 1109VW-LRWWLQAAAAMSNMGMFVAEMSSDSYQLLGMAERGMLPEF-FTKRSRHGTPLIGILF 1282
Query: 349 TAFLCSGCVVIGNLDLITPTVTMFFLLC 376
+A SG +++ L FL C
Sbjct: 1283SA---SGVILLSWLSFQEIVAAENFLYC 1357
>TC78666 homologue to GP|6850938|emb|CAB71135.1 putative imbibition protein
{Cicer arietinum}, partial (96%)
Length = 1507
Score = 31.6 bits (70), Expect = 0.59
Identities = 28/80 (35%), Positives = 37/80 (46%), Gaps = 5/80 (6%)
Frame = +3
Query: 301 ILSTMGAALQSLTGAPRLLAAIANDDILPILKYFKVADGSEPHVATLFTAFLCSGCVVIG 360
I+ T+GA L LLAAI LP+L F D S V TL + S ++ G
Sbjct: 21 IIETLGAGL---VAESHLLAAIIMHLRLPLLVTFPTMDASRVCVITLMDFTVLSRLLLCG 191
Query: 361 NLDLIT-----PTVTMFFLL 375
L + T PT ++F LL
Sbjct: 192 PLMIFTRVIPLPTPSIFRLL 251
>TC83842 similar to PIR|S51171|S51171 amino acid transport protein AAT1 -
Arabidopsis thaliana, partial (55%)
Length = 1295
Score = 30.4 bits (67), Expect = 1.3
Identities = 17/47 (36%), Positives = 25/47 (53%)
Frame = +3
Query: 216 FNALVGLFFPAVTGIMAGSNRSSSLKDTQRSIPLGTLAATLVTTFMY 262
F A LFF A G A S + K+ R IP+G + + ++ TF+Y
Sbjct: 1020 FKASAVLFF-AYIGFDAVSTMAEETKNPGRDIPIGLVGSMVIITFIY 1157
>AL376691
Length = 267
Score = 28.5 bits (62), Expect = 5.0
Identities = 14/47 (29%), Positives = 25/47 (52%)
Frame = -2
Query: 326 DILPILKYFKVADGSEPHVATLFTAFLCSGCVVIGNLDLITPTVTMF 372
+++ I+K +K S PHV +L CS ++I N + P + +F
Sbjct: 143 EVISIIKRYKT---SNPHVKSLTKILKCSNIIIIHNGEAQKPKI*LF 12
>TC93021 homologue to GP|3413719|gb|AAC31242.1| unknown protein {Arabidopsis
thaliana}, partial (2%)
Length = 660
Score = 28.1 bits (61), Expect = 6.5
Identities = 18/47 (38%), Positives = 24/47 (50%), Gaps = 6/47 (12%)
Frame = +2
Query: 21 LLLVALCGTCTFLTAISLSAIATNGAMK----GGG--PYYLIGRALG 61
LL V L TCTFLT I ++ ++ GGG Y L+G +G
Sbjct: 287 LLYVILLSTCTFLTGIESQEKVSSSQLRSLCLGGGF*SYVLLGFKIG 427
>TC86582 similar to GP|21593999|gb|AAM65917.1 ATP phosphoribosyl transferase
{Arabidopsis thaliana}, partial (45%)
Length = 814
Score = 28.1 bits (61), Expect = 6.5
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = -2
Query: 194 GSEYQKTNDAGIPEPDGSVSW 214
G YQKT+D + PDGS+ W
Sbjct: 180 GRVYQKTSDLYLVNPDGSLVW 118
>TC82271 similar to GP|18491137|gb|AAL69537.1 AT5g45420/MFC19_9 {Arabidopsis
thaliana}, partial (43%)
Length = 1116
Score = 28.1 bits (61), Expect = 6.5
Identities = 9/11 (81%), Positives = 9/11 (81%)
Frame = -2
Query: 393 PSWRPRWKFHH 403
PSW PRW FHH
Sbjct: 545 PSWFPRWIFHH 513
>TC87743 similar to GP|15810581|gb|AAL07178.1 putative 2-oxoglutarate/malate
translocator protein {Arabidopsis thaliana}, partial
(50%)
Length = 1359
Score = 27.7 bits (60), Expect = 8.5
Identities = 23/85 (27%), Positives = 38/85 (44%), Gaps = 2/85 (2%)
Frame = +3
Query: 261 MYLVSVIMFGALATREKLLTDRLLTATVAWPFPSLIKIGIILSTMGAALQSLTGAPRL-- 318
++L+ + F A + L DR+ T V W S + + + G + AP +
Sbjct: 756 IWLIVISFFFARGFVKTGLGDRIATYFVKWMGKSTLGL-----SYGLTFSEVLIAPAMPS 920
Query: 319 LAAIANDDILPILKYFKVADGSEPH 343
A A LPI+K ++ GSEP+
Sbjct: 921 TTARAGGVFLPIIKSLSLSAGSEPN 995
>TC83987 PIR|G85579|G85579 probable membrane pump protein ybhI [imported] -
Escherichia coli (strain O157:H7), partial (40%)
Length = 664
Score = 23.5 bits (49), Expect(2) = 9.3
Identities = 17/59 (28%), Positives = 27/59 (44%)
Frame = +3
Query: 321 AIANDDILPILKYFKVADGSEPHVATLFTAFLCSGCVVIGNLDLITPTVTMFFLLCYAG 379
A A +LPI+ VA GSEP + G ++ ++ ++T T + F AG
Sbjct: 360 ARAGGIVLPIINSVAVALGSEPEKSPRRV-----GHYLMMSIYMVTKTTSYMFFTAMAG 521
Score = 22.3 bits (46), Expect(2) = 9.3
Identities = 7/11 (63%), Positives = 7/11 (63%)
Frame = +1
Query: 395 WRPRWKFHHWS 405
W PRW HH S
Sbjct: 622 WSPRW*LHHVS 654
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.142 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,789,435
Number of Sequences: 36976
Number of extensions: 220924
Number of successful extensions: 1599
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1580
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1598
length of query: 414
length of database: 9,014,727
effective HSP length: 99
effective length of query: 315
effective length of database: 5,354,103
effective search space: 1686542445
effective search space used: 1686542445
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0140.12


