

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0137.5
(214 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
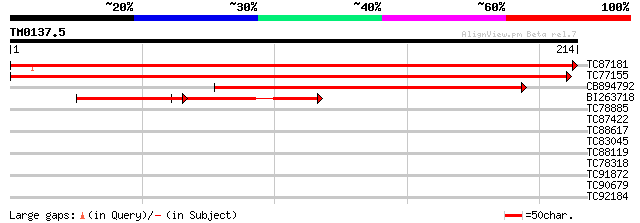
Score E
Sequences producing significant alignments: (bits) Value
TC87181 GP|19309913|emb|CAC41010. Mob1-like protein {Medicago sa... 432 e-122
TC77155 homologue to GP|15028015|gb|AAK76538.1 unknown protein {... 399 e-112
CB894792 weakly similar to GP|19309913|emb Mob1-like protein {Me... 131 2e-31
BI263718 homologue to GP|19309913|emb Mob1-like protein {Medicag... 73 9e-14
TC78885 similar to GP|20160912|dbj|BAB89849. putative transcript... 32 0.24
TC87422 similar to PIR|T00840|T00840 probable senescence-related... 30 0.91
TC88617 similar to GP|10177610|dbj|BAB10957. UDP-glucuronyltrans... 29 1.5
TC83045 28 2.6
TC88119 similar to GP|10178284|emb|CAC08342. rev interacting pro... 27 4.5
TC78318 similar to PIR|T45852|T45852 hypothetical protein F3A4.7... 27 5.9
TC91872 weakly similar to GP|10764862|gb|AAF24552.2 F1K23.23 {Ar... 27 7.7
TC90679 similar to PIR|T10521|T10521 beta-glucosidase (EC 3.2.1.... 27 7.7
TC92184 similar to GP|20466394|gb|AAM20514.1 unknown protein {Ar... 27 7.7
>TC87181 GP|19309913|emb|CAC41010. Mob1-like protein {Medicago sativa subsp.
falcata}, complete
Length = 1570
Score = 432 bits (1112), Expect = e-122
Identities = 209/215 (97%), Positives = 214/215 (99%), Gaps = 1/215 (0%)
Frame = +2
Query: 1 MSLFGLG-RNQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAV 59
MSLFGLG RNQKTFRPKKSAP+GSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAV
Sbjct: 329 MSLFGLGSRNQKTFRPKKSAPTGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAV 508
Query: 60 NTVDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLM 119
NTVDFFNQVNI+FGTLTEFCTPSNCP+MTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLM
Sbjct: 509 NTVDFFNQVNIMFGTLTEFCTPSNCPTMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLM 688
Query: 120 DWIESQLDDETIFPQKLGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHL 179
DWIESQLDDETIFPQ+LGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHL
Sbjct: 689 DWIESQLDDETIFPQRLGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHL 868
Query: 180 NTCFKHFVLFTWEFRLIDKAELAPLEDLVDSIIQL 214
NTCFKHFVLFTWEFRLI+KAELAPLEDLVDSIIQL
Sbjct: 869 NTCFKHFVLFTWEFRLIEKAELAPLEDLVDSIIQL 973
>TC77155 homologue to GP|15028015|gb|AAK76538.1 unknown protein {Arabidopsis
thaliana}, partial (98%)
Length = 1099
Score = 399 bits (1025), Expect = e-112
Identities = 187/212 (88%), Positives = 204/212 (96%)
Frame = +1
Query: 1 MSLFGLGRNQKTFRPKKSAPSGSKGAQLQKHIDATLGSGNLREAVKLPPGEDINEWLAVN 60
MSLFG+GRNQ+TFRPKKS PSGSKGAQL+KHIDATLGSGNLREAVKLPPGED+NEWLAVN
Sbjct: 91 MSLFGIGRNQRTFRPKKSTPSGSKGAQLRKHIDATLGSGNLREAVKLPPGEDLNEWLAVN 270
Query: 61 TVDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMD 120
TVDFFNQVN+L+GTLTEFCTP NC +M+AGPKYEYRWADGV IKKPIEVSAPKYVEYLMD
Sbjct: 271 TVDFFNQVNLLYGTLTEFCTPENCRTMSAGPKYEYRWADGVQIKKPIEVSAPKYVEYLMD 450
Query: 121 WIESQLDDETIFPQKLGAPFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLN 180
WIESQLDDE+IFPQKLG+PFP NF+DVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLN
Sbjct: 451 WIESQLDDESIFPQKLGSPFPTNFKDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLN 630
Query: 181 TCFKHFVLFTWEFRLIDKAELAPLEDLVDSII 212
TCFKHF+LFT EF LIDK ELAPL++L+++II
Sbjct: 631 TCFKHFILFTCEFGLIDKKELAPLQELIETII 726
>CB894792 weakly similar to GP|19309913|emb Mob1-like protein {Medicago
sativa subsp. falcata}, partial (13%)
Length = 358
Score = 131 bits (329), Expect = 2e-31
Identities = 60/118 (50%), Positives = 80/118 (66%)
Frame = -3
Query: 78 FCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDWIESQLDDETIFPQKLG 137
FC P +CPS++AGP + +RWA GV + P+ VSAP++V +LM W++S L FPQ LG
Sbjct: 356 FCPPGDCPSLSAGPGFGFRWAAGVAFRDPVGVSAPRFVGFLMGWVDSPLGVGPFFPQGLG 177
Query: 138 APFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLNTCFKHFVLFTWEFRL 195
APFPP+F VV+ F R FRV+A + S F+ +V L EA L CF+ FV F+W FRL
Sbjct: 176 APFPPHFGGVVRPFFWRFFRVFAPFFPSPFRGVVGLWGEAPLFPCFRPFVFFSWGFRL 3
>BI263718 homologue to GP|19309913|emb Mob1-like protein {Medicago sativa
subsp. falcata}, partial (31%)
Length = 662
Score = 72.8 bits (177), Expect = 9e-14
Identities = 33/57 (57%), Positives = 44/57 (76%)
Frame = +2
Query: 62 VDFFNQVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYL 118
VDFFNQVNI+FGTLTEFCTPSNCP+MTAGPKY G+ ++K + + + ++ Y+
Sbjct: 467 VDFFNQVNIMFGTLTEFCTPSNCPTMTAGPKY------GLILQKTLILVSQRFCIYI 619
Score = 71.6 bits (174), Expect = 2e-13
Identities = 35/42 (83%), Positives = 36/42 (85%)
Frame = +1
Query: 26 AQLQKHIDATLGSGNLREAVKLPPGEDINEWLAVNTVDFFNQ 67
AQLQKHIDATLGSGNLREAVKLPPGEDINEW + TV NQ
Sbjct: 7 AQLQKHIDATLGSGNLREAVKLPPGEDINEWPCL*TVRISNQ 132
>TC78885 similar to GP|20160912|dbj|BAB89849. putative transcription
initiation factor {Oryza sativa (japonica
cultivar-group)}, partial (67%)
Length = 1073
Score = 31.6 bits (70), Expect = 0.24
Identities = 18/63 (28%), Positives = 27/63 (42%), Gaps = 7/63 (11%)
Frame = -3
Query: 130 TIFPQKLGAPFPPNFRDVVKTIFKRLFRVYAHI-------YHSHFQKIVSLKEEAHLNTC 182
TIF + P P F ++ +F + HI +H + L+E+AH N
Sbjct: 489 TIFNHRFLCPLPAQFSIII-----HVFMTWLHIKLMLYFPFHCNLVTFACLREDAHRNEV 325
Query: 183 FKH 185
FKH
Sbjct: 324 FKH 316
>TC87422 similar to PIR|T00840|T00840 probable senescence-related protein
[imported] - Arabidopsis thaliana, partial (55%)
Length = 1660
Score = 29.6 bits (65), Expect = 0.91
Identities = 17/55 (30%), Positives = 26/55 (46%)
Frame = -1
Query: 67 QVNILFGTLTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPIEVSAPKYVEYLMDW 121
Q+ IL L CT + P + Y +D T+K+PI K + Y++DW
Sbjct: 1558 QLKILTYPLNFKCTMDSRPVKRTPLWFRYFASDKKTVKQPI*KQMVKSIFYILDW 1394
>TC88617 similar to GP|10177610|dbj|BAB10957. UDP-glucuronyltransferase-like
protein {Arabidopsis thaliana}, partial (75%)
Length = 1958
Score = 28.9 bits (63), Expect = 1.5
Identities = 18/56 (32%), Positives = 25/56 (44%), Gaps = 16/56 (28%)
Frame = -3
Query: 155 LFRVYAHIYHSHFQKIVSLKEEA---------------HLNTCFKHFVLF-TWEFR 194
LFR+ +Y +HF ++SL A H C +HF +F WEFR
Sbjct: 1626 LFRMCRDLYTNHFGVMLSLTSTASGSGCFGLCSVRVRMHFCACSEHFFVFIRWEFR 1459
>TC83045
Length = 709
Score = 28.1 bits (61), Expect = 2.6
Identities = 13/23 (56%), Positives = 16/23 (69%)
Frame = +1
Query: 118 LMDWIESQLDDETIFPQKLGAPF 140
++DW ES LDDET F LG+ F
Sbjct: 616 VLDWNESDLDDETRF*LWLGSGF 684
>TC88119 similar to GP|10178284|emb|CAC08342. rev interacting protein
mis3-like {Arabidopsis thaliana}, partial (79%)
Length = 1369
Score = 27.3 bits (59), Expect = 4.5
Identities = 14/35 (40%), Positives = 21/35 (60%)
Frame = +2
Query: 139 PFPPNFRDVVKTIFKRLFRVYAHIYHSHFQKIVSL 173
P PP+F ++VK IFK+L H+ + + VSL
Sbjct: 299 PSPPSFLNIVKNIFKKL----GHLLNPPSKNSVSL 391
>TC78318 similar to PIR|T45852|T45852 hypothetical protein F3A4.70 -
Arabidopsis thaliana, partial (48%)
Length = 1923
Score = 26.9 bits (58), Expect = 5.9
Identities = 10/28 (35%), Positives = 21/28 (74%)
Frame = -1
Query: 146 DVVKTIFKRLFRVYAHIYHSHFQKIVSL 173
++V+ +F FR++ H+++SH + I+SL
Sbjct: 513 NIVQNVF---FRLFIHLFNSHSRSIISL 439
>TC91872 weakly similar to GP|10764862|gb|AAF24552.2 F1K23.23 {Arabidopsis
thaliana}, partial (41%)
Length = 1088
Score = 26.6 bits (57), Expect = 7.7
Identities = 22/93 (23%), Positives = 36/93 (38%), Gaps = 6/93 (6%)
Frame = +3
Query: 21 SGSKGAQLQKHIDATLGSGNLREAVK------LPPGEDINEWLAVNTVDFFNQVNILFGT 74
S KG+ L K + +L SG A+ LP + + ++VN++
Sbjct: 321 SNRKGSSLSKILSVSLASGMFLVAISALGQFCLPRLSKERKHTVEHRSLLTSEVNVMHDF 500
Query: 75 LTEFCTPSNCPSMTAGPKYEYRWADGVTIKKPI 107
L C S A K Y W+D + ++ I
Sbjct: 501 LDTTKVEEFCVSAVAKLKNAYGWSDEIKVEDGI 599
>TC90679 similar to PIR|T10521|T10521 beta-glucosidase (EC 3.2.1.21) -
common nasturtium, partial (43%)
Length = 1017
Score = 26.6 bits (57), Expect = 7.7
Identities = 14/36 (38%), Positives = 19/36 (51%), Gaps = 10/36 (27%)
Frame = -2
Query: 63 DFFNQVNILFGTLTEFCT----------PSNCPSMT 88
+F ++N LF T+ + CT PSNCPS T
Sbjct: 443 NFSCRINCLFYTIKDGCTCCKITTTKSLPSNCPSTT 336
>TC92184 similar to GP|20466394|gb|AAM20514.1 unknown protein {Arabidopsis
thaliana}, partial (26%)
Length = 1024
Score = 26.6 bits (57), Expect = 7.7
Identities = 14/41 (34%), Positives = 21/41 (51%)
Frame = -1
Query: 144 FRDVVKTIFKRLFRVYAHIYHSHFQKIVSLKEEAHLNTCFK 184
FRD + IF + +YAHIY + + S E+ + C K
Sbjct: 169 FRDKIM*IFTGPYWIYAHIYANRYSSTKS-NEQKSVCCCMK 50
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.139 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,994,556
Number of Sequences: 36976
Number of extensions: 99196
Number of successful extensions: 536
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 536
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 536
length of query: 214
length of database: 9,014,727
effective HSP length: 92
effective length of query: 122
effective length of database: 5,612,935
effective search space: 684778070
effective search space used: 684778070
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0137.5


