

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0136.7
(841 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
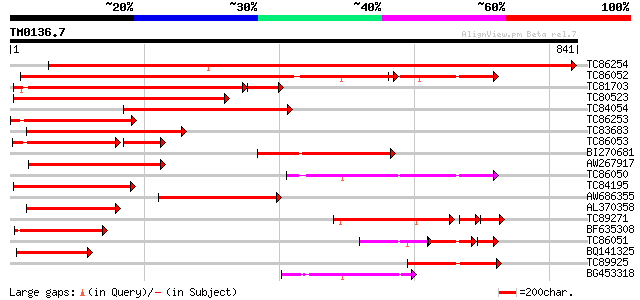
TC86254 similar to GP|14970841|emb|CAC44501. beta-galactosidase ... 1281 0.0
TC86052 similar to GP|3641865|emb|CAA09457.1 beta-galactosidase ... 670 0.0
TC81703 similar to GP|20259271|gb|AAM14371.1 putative beta-galac... 518 e-158
TC80523 homologue to GP|3860321|emb|CAA10128.1 beta-galactosidas... 462 e-130
TC84054 similar to GP|7939621|gb|AAF70823.1| beta-galactosidase ... 348 4e-96
TC86253 homologue to GP|14970841|emb|CAC44501. beta-galactosidas... 334 7e-92
TC83683 similar to GP|7939621|gb|AAF70823.1| beta-galactosidase ... 324 8e-89
TC86053 homologue to GP|3641863|emb|CAA06309.1 beta-galactosidas... 248 2e-86
BI270681 similar to GP|14970841|em beta-galactosidase {Fragaria ... 306 3e-83
AW267917 weakly similar to GP|14970841|em beta-galactosidase {Fr... 303 2e-82
TC86050 similar to GP|3641863|emb|CAA06309.1 beta-galactosidase ... 278 8e-75
TC84195 similar to GP|20259271|gb|AAM14371.1 putative beta-galac... 274 1e-73
AW686355 similar to GP|14970839|em beta-galactosidase {Fragaria ... 265 4e-71
AL370358 similar to GP|20260596|gb galactosidase putative {Arab... 234 1e-61
TC89271 weakly similar to GP|10177669|dbj|BAB11029. beta-galacto... 172 8e-56
BF635308 similar to GP|10177061|db beta-galactosidase {Arabidops... 184 1e-46
TC86051 homologue to GP|3641863|emb|CAA06309.1 beta-galactosidas... 95 2e-46
BQ141325 similar to GP|2300473|emb unnamed protein product {unid... 166 3e-41
TC89925 homologue to GP|3860321|emb|CAA10128.1 beta-galactosidas... 150 1e-36
BG453318 homologue to GP|3204134|emb beta-galactosidase {Cicer a... 147 2e-35
>TC86254 similar to GP|14970841|emb|CAC44501. beta-galactosidase {Fragaria x
ananassa}, partial (87%)
Length = 2581
Score = 1281 bits (3315), Expect = 0.0
Identities = 616/827 (74%), Positives = 683/827 (82%), Gaps = 44/827 (5%)
Frame = +3
Query: 58 MWPDLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIG 117
MWPDLIQK+KDGGLDVIETYVFWNLHEPV+GQY+F+GR DLVKFVK VA AGLYVHLRIG
Sbjct: 75 MWPDLIQKSKDGGLDVIETYVFWNLHEPVKGQYDFDGRKDLVKFVKAVAEAGLYVHLRIG 254
Query: 118 PYACAEWNYGGFPLWLHFIPGIQFRTDNEPFK--AEMQRFTAKIVDMMKQENLYASQGGP 175
PY CAEWNYGGFPLWLHFIPGI+FRTDNEPFK AEM+RFTAKIVD+MKQE LYASQGGP
Sbjct: 255 PYVCAEWNYGGFPLWLHFIPGIKFRTDNEPFKVEAEMKRFTAKIVDLMKQEKLYASQGGP 434
Query: 176 IILSQIENEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGF 235
IILSQIENEYG+++ AYG A YINWAA MATSLDTGVPWVMCQQE+APD IINTCNGF
Sbjct: 435 IILSQIENEYGDIDSAYGSAGKSYINWAAKMATSLDTGVPWVMCQQEDAPDSIINTCNGF 614
Query: 236 YCDQFTPNSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYM-- 293
YCDQFTPNS KPK WTE ++ W L FGG P+RPVEDLAFAVARFFQ GGT QNYYM
Sbjct: 615 YCDQFTPNSNTKPKMWTENWSAWYLLFGGGFPHRPVEDLAFAVARFFQRGGTFQNYYMVL 794
Query: 294 --------------------------------------YHGGTNFGRTSGGPFVATSYDY 315
YHGGTNF R++GGPF+ATSYD+
Sbjct: 795 QPEMFFTSSIYYMVLFLRPVFKRN*LILYHFLLPAQKQYHGGTNFDRSTGGPFIATSYDF 974
Query: 316 DAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGSNLEAAVYKTEAECVAF 375
DA IDEYG IRQPKWGHLKD+HKA+KLCEEALIAT+P ITSLG NLEAAVYKT + C AF
Sbjct: 975 DAPIDEYGIIRQPKWGHLKDLHKAVKLCEEALIATEPKITSLGPNLEAAVYKTGSVCAAF 1154
Query: 376 LANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFTTESLKE-V 434
LAN+D SD TV F+GNSY+LPAWSVSILPDCKNVVLNTAKINS S IS+F T+S KE +
Sbjct: 1155LANVDTKSDKTVNFSGNSYHLPAWSVSILPDCKNVVLNTAKINSASAISNFVTKSSKEDI 1334
Query: 435 GSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSDYLWYSLRIDLEDDAGAQT 494
SLE S WSWI+EP+GISK D FS+ GLLEQIN TADRSDYLWYSL +DL+DD G+QT
Sbjct: 1335SSLETSSSKWSWINEPVGISKDDIFSKTGLLEQINITADRSDYLWYSLSVDLKDDLGSQT 1514
Query: 495 VLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFG 554
VLHIESLGHALHAF+NGKLAGS GN+D K+NV+IPI ++ G N IDLLSLTVGLQ++G
Sbjct: 1515VLHIESLGHALHAFVNGKLAGSHTGNKDKPKLNVDIPIKVIYGNNQIDLLSLTVGLQNYG 1694
Query: 555 PFFDTWGAGITGPVILKGLKNGS-TLDFSSKQWTYQIGLKGEDLGLPSGTSGQWNSQSTL 613
FFD WGAGITGPVILKGLKNG+ TLD SS++WTYQIGLKGEDLGL SG+SG WNSQST
Sbjct: 1695AFFDRWGAGITGPVILKGLKNGNNTLDLSSRKWTYQIGLKGEDLGLSSGSSGGWNSQSTY 1874
Query: 614 PKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCN 673
PKNQPL WYK NF APSG++P+AIDFTGMGKGEAWVNGQSIGRYWPTY++ GCT SCN
Sbjct: 1875PKNQPLVWYKTNFDAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYVASNAGCTDSCN 2054
Query: 674 YRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPTQISIAIKLIGSSC 733
YRGPY SSKC KNCGKPSQTLYHVPRS+L+P+ NTLVLFEE+GGDPTQIS A K + S C
Sbjct: 2055YRGPYTSSKCRKNCGKPSQTLYHVPRSFLKPNGNTLVLFEENGGDPTQISFATKQLESVC 2234
Query: 734 SHVSESHPPPVDLWNSDTESDRSGGPVLSLECPYPNEVITTIKFASFGTPHGNCGNFSHG 793
SHVS+SHPP +DLWN DTES GP L L CP N+VI++IKFAS+GTP G CGNF G
Sbjct: 2235SHVSDSHPPQIDLWNQDTESGGKVGPALLLSCPNHNQVISSIKFASYGTPLGTCGNFYRG 2414
Query: 794 DCSSKQALSIVQKACIGSSSCSIGVSINTFGDPCGGVTKSLAVEASC 840
CSS +ALSIV+KACIGS SCS+GVS +TFGDPC GV KSLAVEA+C
Sbjct: 2415RCSSNKALSIVKKACIGSRSCSVGVSTDTFGDPCRGVPKSLAVEATC 2555
>TC86052 similar to GP|3641865|emb|CAA09457.1 beta-galactosidase {Cicer
arietinum}, partial (94%)
Length = 2539
Score = 670 bits (1729), Expect = 0.0
Identities = 334/567 (58%), Positives = 388/567 (67%), Gaps = 7/567 (1%)
Frame = +2
Query: 17 CVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIET 76
CV A+V+YDH+ALVIDG+RR+LISGSIHYPRSTPEMWPDL QKAKDGGLDVI+T
Sbjct: 110 CVCGFVVLLASVSYDHKALVIDGQRRILISGSIHYPRSTPEMWPDLFQKAKDGGLDVIQT 289
Query: 77 YVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFI 136
YVFWN HEP G Y FE R DLVKFVK AGLYVHLRIGPY CAEWNYGGFP+WL ++
Sbjct: 290 YVFWNGHEPSPGNYYFEDRFDLVKFVKLAQQAGLYVHLRIGPYVCAEWNYGGFPVWLKYV 469
Query: 137 PGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVEGAYGPAA 196
PG+ FRTDNEPFKA MQ+FT KIV MMK E+L+ +QGGPII+SQIENEYG VE G
Sbjct: 470 PGMAFRTDNEPFKAAMQKFTTKIVTMMKAESLFQTQGGPIIMSQIENEYGPVEWEIGAPG 649
Query: 197 VPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPNSKAKPKFWTEGYN 256
Y WAA MA LDTGVPW MC+QE+APDP+I+TCNG+YC+ FTPN KPK WTE ++
Sbjct: 650 KAYTKWAAQMAVGLDTGVPWDMCKQEDAPDPVIDTCNGYYCENFTPNENFKPKMWTENWS 829
Query: 257 GWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGRTSGGPFVATSYDYD 316
GW FGGA+ +RP EDLA++VARF Q G+ NYYMYHGGTNFGRTS G F+ATSYDYD
Sbjct: 830 GWYTDFGGAISHRPTEDLAYSVARFIQNRGSFVNYYMYHGGTNFGRTSSGLFIATSYDYD 1009
Query: 317 AAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDPTITSLGS-NLEAAVYKTEAE-CVA 374
A IDEYG +PKW HLK++HKAIK CE ALI+ DPT+T LG+ NLEA VY C A
Sbjct: 1010APIDEYGLPNEPKWSHLKNLHKAIKQCEPALISVDPTVTWLGNKNLEAHVYYVNTSICAA 1189
Query: 375 FLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFTTESLKEV 434
FLAN D S ATV F Y+LP WSVSILPDCK VV NTA +N S K +
Sbjct: 1190FLANYDTKSAATVTFGNGQYDLPPWSVSILPDCKTVVFNTATVNGHSF--------HKRM 1345
Query: 435 GSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSDYLWYSLRIDLEDDA---- 490
+E S+ EP S DS L EQIN T D SDYLWY +++
Sbjct: 1346TPVETTFDWQSYSEEPAYSSDDDSIIANALWEQINVTRDSSDYLWYLTDVNISPSESFIK 1525
Query: 491 -GAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLTVG 549
G L I S GH LH F+NG+L+G+ G DN KV + L G N I LLS+ VG
Sbjct: 1526NGQFPTLTINSAGHVLHVFVNGQLSGTVYGGLDNPKVTFSESVNLKVGNNKISLLSVAVG 1705
Query: 550 LQHFGPFFDTWGAGITGPVILKGLKNG 576
L + G F+TW G P ++G + G
Sbjct: 1706LPNVGLHFETWNVGGVRPSKIEGSR*G 1786
Score = 189 bits (480), Expect = 4e-48
Identities = 90/165 (54%), Positives = 110/165 (66%), Gaps = 3/165 (1%)
Frame = +3
Query: 563 GITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDLGLPSGTSGQ---WNSQSTLPKNQPL 619
G GPV LKGL G T D S ++W+Y++GLKGE L L + T W S+L K QPL
Sbjct: 1746 GXLGPVRLKGLDEG-TRDLSWQKWSYKVGLKGESLSLHTITGSSSIDWTQGSSLAKKQPL 1922
Query: 620 TWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYD 679
TWYK F APSGNDP+A+D + MGKGE W+N QSIGR+WP YI+ G CNY G +
Sbjct: 1923 TWYKTTFDAPSGNDPVALDMSSMGKGEIWINDQSIGRHWPAYIA--HGNCDECNYAGTFT 2096
Query: 680 SSKCLKNCGKPSQTLYHVPRSWLQPDSNTLVLFEESGGDPTQISI 724
+ KC NCG+P+Q YH+PRSWL N LV+ EE GGDPT IS+
Sbjct: 2097 NPKCRTNCGEPTQKWYHIPRSWLSSSGNVLVVLEEWGGDPTGISL 2231
>TC81703 similar to GP|20259271|gb|AAM14371.1 putative beta-galactosidase
{Arabidopsis thaliana}, partial (44%)
Length = 1310
Score = 518 bits (1333), Expect(2) = e-158
Identities = 235/350 (67%), Positives = 280/350 (79%), Gaps = 3/350 (0%)
Frame = +2
Query: 6 TQFLLLLLWL---LCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDL 62
++FL L + L L VY+ +VTYD +A++I+G+RR+L SGSIHYPRSTP+MW DL
Sbjct: 116 SKFLFLFVSLTLFLAVYS------DVTYDRKAIIINGQRRILFSGSIHYPRSTPDMWEDL 277
Query: 63 IQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACA 122
I KAK+GGLDVIETYVFWN+HEP G YNFEGR DLV+F++ V AGLY HLRIGPY CA
Sbjct: 278 IYKAKEGGLDVIETYVFWNVHEPSPGNYNFEGRNDLVRFIQTVHKAGLYAHLRIGPYVCA 457
Query: 123 EWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIE 182
EWN+GGFP+WL ++PGI FR DNEPFK MQ FT KIV MMK E LY SQGGPIILSQIE
Sbjct: 458 EWNFGGFPVWLKYVPGISFRQDNEPFKKAMQGFTEKIVGMMKSERLYESQGGPIILSQIE 637
Query: 183 NEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTP 242
NEYG GP Y++WAA MA + TGVPW+MC++++APDP+INTCNGFYCD+FTP
Sbjct: 638 NEYGAQSKMLGPVGYNYMSWAAKMAVEMGTGVPWIMCKEDDAPDPVINTCNGFYCDKFTP 817
Query: 243 NSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFGR 302
N KP WTE ++GW FGG + RPV+DLAFAVARF Q GG+ NYYMYHGGTNFGR
Sbjct: 818 NKPYKPTMWTEAWSGWFSEFGGPIHKRPVQDLAFAVARFIQKGGSFVNYYMYHGGTNFGR 997
Query: 303 TSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIATDP 352
T+GGPF+ TSYDYDA +DEYG IRQPK+GHLK++HKAIK+CE+ALI+TDP
Sbjct: 998 TAGGPFITTSYDYDAPLDEYGLIRQPKYGHLKELHKAIKMCEKALISTDP 1147
Score = 60.8 bits (146), Expect(2) = e-158
Identities = 31/53 (58%), Positives = 38/53 (71%), Gaps = 1/53 (1%)
Frame = +3
Query: 354 ITSLGSNLEAAVYKTEA-ECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILP 405
+TSLG+ +A VY TE+ +C AFL+N D+ S A V FN YNLP WSVSILP
Sbjct: 1152 VTSLGNFQQAYVYTTESGDCSAFLSNYDSKSSARVMFNNMHYNLPPWSVSILP 1310
>TC80523 homologue to GP|3860321|emb|CAA10128.1 beta-galactosidase {Cicer
arietinum}, partial (41%)
Length = 1109
Score = 462 bits (1190), Expect = e-130
Identities = 215/322 (66%), Positives = 248/322 (76%), Gaps = 1/322 (0%)
Frame = +1
Query: 6 TQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQK 65
++ LLLL + + VTYD +A++I+G+RR+LISGSIHYPRSTPEMW DLIQK
Sbjct: 142 SKLLLLLFFTIFFLGSEVIHCTVTYDRKAIIINGQRRILISGSIHYPRSTPEMWEDLIQK 321
Query: 66 AKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWN 125
AKDGGLDVI+TYVFWN+HEP G YNFEGR DLV+F+K V GLYVHLRIGPY CAEWN
Sbjct: 322 AKDGGLDVIDTYVFWNVHEPSPGNYNFEGRYDLVQFIKTVQKKGLYVHLRIGPYVCAEWN 501
Query: 126 YGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEY 185
+GGFP+WL ++PGI FRTDN PFKA MQ FT KIV MMK E L+ SQGGPIILSQIENEY
Sbjct: 502 FGGFPVWLKYVPGISFRTDNGPFKAAMQGFTQKIVQMMKNEKLFQSQGGPIILSQIENEY 681
Query: 186 GNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPNSK 245
G A G + Y NWAA MA L TGVPWVMC++++APDP+INTCNGFYCD F+PN
Sbjct: 682 GPQGRALGASGHAYSNWAAKMAVGLGTGVPWVMCKEDDAPDPVINTCNGFYCDDFSPNKP 861
Query: 246 AKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQNYYMYHGGTNFG-RTS 304
KPK WTE ++GW FGG+ P RPVEDLAFAVARF Q GG+ NYYMYHG F +
Sbjct: 862 YKPKLWTESWSGWFSEFGGSNPQRPVEDLAFAVARFIQKGGSFFNYYMYHGRN*FSDDRA 1041
Query: 305 GGPFVATSYDYDAAIDEYGFIR 326
GGPF+ TSYDYDA IDEYG +R
Sbjct: 1042GGPFITTSYDYDAPIDEYGLLR 1107
>TC84054 similar to GP|7939621|gb|AAF70823.1| beta-galactosidase
{Lycopersicon esculentum}, partial (29%)
Length = 917
Score = 348 bits (894), Expect = 4e-96
Identities = 162/251 (64%), Positives = 190/251 (75%), Gaps = 1/251 (0%)
Frame = +1
Query: 170 ASQGGPIILSQIENEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPII 229
ASQGGPIILSQIENEYG E Y Y WAA MA S +T VPW+MCQQ +APDP+I
Sbjct: 1 ASQGGPIILSQIENEYGYYENYYKEDGKKYALWAAKMAVSQNTSVPWIMCQQWDAPDPVI 180
Query: 230 NTCNGFYCDQFTPNSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVARFFQLGGTLQ 289
+TCN FYCDQFTP S +PK WTE + GW FGG P+RPVED+AF+VARFFQ GG+L
Sbjct: 181 DTCNSFYCDQFTPTSPKRPKMWTENWPGWFKTFGGRDPHRPVEDVAFSVARFFQKGGSLN 360
Query: 290 NYYMYHGGTNFGRTSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAIKLCEEALIA 349
NYYMYHGGTNFGRT+GGPF+ TSYDYDA IDEYG R PKWGHLK++HKAIKLCE L+
Sbjct: 361 NYYMYHGGTNFGRTAGGPFITTSYDYDAPIDEYGLPRLPKWGHLKELHKAIKLCEHVLLY 540
Query: 350 TDPTITSLGSNLEAAVY-KTEAECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCK 408
SLG ++EA +Y + C AF++N+D+ +D V F SY+LPAWSVSILPDCK
Sbjct: 541 GKSVNISLGPSVEADIYTDSSGACAAFISNVDDKNDKKVVFRNASYHLPAWSVSILPDCK 720
Query: 409 NVVLNTAKINS 419
NVV NTAK++S
Sbjct: 721 NVVFNTAKVSS 753
>TC86253 homologue to GP|14970841|emb|CAC44501. beta-galactosidase {Fragaria
x ananassa}, partial (19%)
Length = 748
Score = 334 bits (857), Expect = 7e-92
Identities = 157/188 (83%), Positives = 170/188 (89%)
Frame = +3
Query: 1 MAMRRTQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWP 60
+ MR + +L+LLW L P FC NV YDHRALVIDGKRRVLISGSIHYPRSTP+MWP
Sbjct: 195 ITMRAFEIVLVLLWFL----PKMFCTNVDYDHRALVIDGKRRVLISGSIHYPRSTPQMWP 362
Query: 61 DLIQKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYA 120
DLIQK+KDGGLDVIETYVFWNLHEPV+GQY+F+GR DLVKFVK VA AGLYVHLRIGPY
Sbjct: 363 DLIQKSKDGGLDVIETYVFWNLHEPVKGQYDFDGRKDLVKFVKAVAEAGLYVHLRIGPYV 542
Query: 121 CAEWNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQ 180
CAEWNYGGFPLWLHFIPGI+FRTDNEPFKAEM+RFTAKIVD+MKQE LYASQGGPIILSQ
Sbjct: 543 CAEWNYGGFPLWLHFIPGIKFRTDNEPFKAEMKRFTAKIVDLMKQEKLYASQGGPIILSQ 722
Query: 181 IENEYGNV 188
IENEYGN+
Sbjct: 723 IENEYGNI 746
>TC83683 similar to GP|7939621|gb|AAF70823.1| beta-galactosidase
{Lycopersicon esculentum}, partial (26%)
Length = 927
Score = 324 bits (831), Expect = 8e-89
Identities = 149/237 (62%), Positives = 179/237 (74%)
Frame = +2
Query: 26 ANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLHEP 85
+NV+YD R+L+IDG+R++LIS SIHYPRS P MWP LIQ AK+GG+DVIETYVFWN HE
Sbjct: 83 SNVSYDGRSLIIDGQRKLLISASIHYPRSVPAMWPALIQTAKEGGIDVIETYVFWNGHEL 262
Query: 86 VQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFIPGIQFRTDN 145
G Y F GR DLV+F K V AG+Y+ LRIGP+ AEWN+GG P WLH+IPG FRT N
Sbjct: 263 SPGNYYFGGRFDLVQFAKVVQDAGMYLILRIGPFVAAEWNFGGVPKWLHYIPGTVFRTYN 442
Query: 146 EPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNVEGAYGPAAVPYINWAAS 205
+PF M++FT IV++MK+E L+ASQGGPIILSQIENEYG E Y Y WAA
Sbjct: 443 QPFMHHMEKFTTYIVNLMKKEKLFASQGGPIILSQIENEYGYYENYYKEDGKKYALWAAK 622
Query: 206 MATSLDTGVPWVMCQQENAPDPIINTCNGFYCDQFTPNSKAKPKFWTEGYNGWLLYF 262
MA S +T VPW+MCQQ +APDP+I+TCN FYCDQFTP S +PK WTE + GW + F
Sbjct: 623 MAVSQNTSVPWIMCQQWDAPDPVIDTCNSFYCDQFTPTSPKRPKMWTENWPGWYVEF 793
>TC86053 homologue to GP|3641863|emb|CAA06309.1 beta-galactosidase {Cicer
arietinum}, partial (29%)
Length = 925
Score = 248 bits (633), Expect(2) = 2e-86
Identities = 114/161 (70%), Positives = 134/161 (82%)
Frame = +2
Query: 4 RRTQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLI 63
RR ++L L + +C A+VTYDH+A+VI+GKRR+LISGSIHYPRSTP+MWPDLI
Sbjct: 254 RRNCYILFLCFFVCYVT-----ASVTYDHKAIVINGKRRILISGSIHYPRSTPQMWPDLI 418
Query: 64 QKAKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAE 123
QKAKDGG+DVIETYVFWN HEP QG+Y FE R DLVKF+K V AGLYVHLRIGPY CAE
Sbjct: 419 QKAKDGGVDVIETYVFWNGHEPSQGKYYFEDRFDLVKFIKVVQQAGLYVHLRIGPYVCAE 598
Query: 124 WNYGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMK 164
WN+GGFP+WL ++PG+ FRTDNEPFKA MQ+FT KIV +MK
Sbjct: 599 WNFGGFPVWLKYVPGVAFRTDNEPFKAAMQKFTTKIVSIMK 721
Score = 90.9 bits (224), Expect(2) = 2e-86
Identities = 42/63 (66%), Positives = 47/63 (73%)
Frame = +1
Query: 169 YASQGGPIILSQIENEYGNVEGAYGPAAVPYINWAASMATSLDTGVPWVMCQQENAPDPI 228
+ SQGGPIILSQIENEYG VE G Y W + MA L+TGVPWVMC+QENAPDPI
Sbjct: 736 FQSQGGPIILSQIENEYGPVEWEIGAPGKSYTKWFSQMAVGLNTGVPWVMCKQENAPDPI 915
Query: 229 INT 231
I+T
Sbjct: 916 IDT 924
>BI270681 similar to GP|14970841|em beta-galactosidase {Fragaria x ananassa},
partial (20%)
Length = 610
Score = 306 bits (783), Expect = 3e-83
Identities = 153/205 (74%), Positives = 172/205 (83%)
Frame = +2
Query: 368 TEAECVAFLANIDNTSDATVKFNGNSYNLPAWSVSILPDCKNVVLNTAKINSESMISSFT 427
T + C AFLAN+D +D TV F+GNSY+LPAWSVSILPDCKNVVLNTAKINS S IS+F
Sbjct: 2 TGSVCAAFLANVDTKNDKTVNFSGNSYHLPAWSVSILPDCKNVVLNTAKINSASAISNFV 181
Query: 428 TESLKEVGSLEGPSPGWSWISEPIGISKADSFSRFGLLEQINTTADRSDYLWYSLRIDLE 487
TE ++ SLE S WSWI+EP+GISK D S+ GLLEQINTTADRSDYLWYSL +DL
Sbjct: 182 TE---DISSLETSSSKWSWINEPVGISKDDILSKTGLLEQINTTADRSDYLWYSLSLDLA 352
Query: 488 DDAGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAGKNTIDLLSLT 547
DD G+QTVLHIESLGHALHAFINGKLAG+ GN D SK+NV+IPI LV+GKN IDLLSLT
Sbjct: 353 DDPGSQTVLHIESLGHALHAFINGKLAGNQAGNSDKSKLNVDIPIALVSGKNKIDLLSLT 532
Query: 548 VGLQHFGPFFDTWGAGITGPVILKG 572
VGLQ++G FFDT GAGITGPVILKG
Sbjct: 533 VGLQNYGAFFDTVGAGITGPVILKG 607
>AW267917 weakly similar to GP|14970841|em beta-galactosidase {Fragaria x
ananassa}, partial (23%)
Length = 731
Score = 303 bits (776), Expect = 2e-82
Identities = 140/205 (68%), Positives = 165/205 (80%), Gaps = 1/205 (0%)
Frame = +1
Query: 28 VTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLHEPVQ 87
VTYD R+L+I+G+R +L SGSIHYPRSTP+MWP LI KAK GGLDVI+TYVFWNLHEP
Sbjct: 115 VTYDGRSLIINGQRNILFSGSIHYPRSTPQMWPGLIAKAKQGGLDVIQTYVFWNLHEPQP 294
Query: 88 GQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFIPGIQFRTDNEP 147
G+Y+F GR DLV F+K + GLYV LRIGP+ +EWNYGGFP WLH +PGI +RTDNEP
Sbjct: 295 GKYDFSGRNDLVGFIKEIHAQGLYVSLRIGPFIESEWNYGGFPFWLHDVPGIVYRTDNEP 474
Query: 148 FKAEMQRFTAKIVD-MMKQENLYASQGGPIILSQIENEYGNVEGAYGPAAVPYINWAASM 206
FK MQ FT KIV+ +K+E LYASQGGPIILSQIENEYGN++ A+G A Y+ WAA M
Sbjct: 475 FKFYMQNFTTKIVNHELKEEGLYASQGGPIILSQIENEYGNIQKAFGTAXSQYVGWAAKM 654
Query: 207 ATSLDTGVPWVMCQQENAPDPIINT 231
A L+T VPWVMC+Q +APDP INT
Sbjct: 655 AXGLNTSVPWVMCKQPDAPDPWINT 729
>TC86050 similar to GP|3641863|emb|CAA06309.1 beta-galactosidase {Cicer
arietinum}, partial (42%)
Length = 1539
Score = 278 bits (710), Expect = 8e-75
Identities = 147/323 (45%), Positives = 195/323 (59%), Gaps = 9/323 (2%)
Frame = +1
Query: 411 VLNTAKINSESMISSFTTESLKEVGSLEGPSPGW-SWISEPIGISKADSFSRFGLLEQIN 469
V NTAK+ + + S T + + W S+ +P ++ S++ GLLEQ++
Sbjct: 1 VFNTAKVRAPRVHRSMTPAN---------SAFNWQSYNEQPAFSGESGSWTANGLLEQLS 153
Query: 470 TTADRSDYLWYSLRIDLEDDAG-----AQTVLHIESLGHALHAFINGKLAGSGKGNRDNS 524
T D+SDYLWY +++ + G VL S GH LH FING+ G+ G+ DN
Sbjct: 154 QTWDKSDYLWYMTDVNISPNEGFIKNGQNPVLTAMSAGHVLHVFINGQFWGTAYGSLDNP 333
Query: 525 KVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSK 584
K+ + L G N I LLS+ GL + G ++ W G+ GPV LKGL G T + S +
Sbjct: 334 KLTFSNSVKLRVGNNKISLLSVAGGLSNVGVHYEKWNVGVLGPVTLKGLNEG-TRNLSKQ 510
Query: 585 QWTYQIGLKGEDLGL--PSGTSG-QWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTG 641
+W+Y+IGLKGE L L SG+S +W S L K QPLTWYK F+AP+GNDP+A+D +
Sbjct: 511 KWSYKIGLKGESLNLHTTSGSSSVKWTQGSFLSKKQPLTWYKTTFNAPAGNDPLALDMSS 690
Query: 642 MGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSW 701
MGKGE WVNGQSIGR+WP YI+ G SCNY G + KC NCG+P+Q YH+PRSW
Sbjct: 691 MGKGEIWVNGQSIGRHWPAYIA--RGNCGSCNYAGTFTDKKCRTNCGQPTQKWYHIPRSW 864
Query: 702 LQPDSNTLVLFEESGGDPTQISI 724
L P N LV+ EE GGDPT IS+
Sbjct: 865 LNPSGNVLVVSEEWGGDPTGISL 933
>TC84195 similar to GP|20259271|gb|AAM14371.1 putative beta-galactosidase
{Arabidopsis thaliana}, partial (19%)
Length = 673
Score = 274 bits (700), Expect = 1e-73
Identities = 125/181 (69%), Positives = 151/181 (83%)
Frame = +2
Query: 6 TQFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQK 65
++++ ++LC+ + ++VTYD +A++I+G+RR+L SGSIHYPRSTP+MW DLIQK
Sbjct: 74 SKWMFTFCFVLCLVSQFIH-SSVTYDRKAILINGQRRILFSGSIHYPRSTPDMWEDLIQK 250
Query: 66 AKDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWN 125
AK+GGLDVIETYVFWN+HEP G YNFEGR DLVKF+K V AGLY HLRIGPY CAEWN
Sbjct: 251 AKEGGLDVIETYVFWNVHEPSPGNYNFEGRYDLVKFIKTVQKAGLYAHLRIGPYVCAEWN 430
Query: 126 YGGFPLWLHFIPGIQFRTDNEPFKAEMQRFTAKIVDMMKQENLYASQGGPIILSQIENEY 185
+GGFP+WL ++PGI FRTDNEPFK EMQRFT KIV +MK E L+ SQGGPIILSQIENEY
Sbjct: 431 FGGFPVWLKYVPGISFRTDNEPFKREMQRFTEKIVGLMKSEQLFESQGGPIILSQIENEY 610
Query: 186 G 186
G
Sbjct: 611 G 613
>AW686355 similar to GP|14970839|em beta-galactosidase {Fragaria x ananassa},
partial (22%)
Length = 591
Score = 265 bits (678), Expect = 4e-71
Identities = 118/184 (64%), Positives = 145/184 (78%), Gaps = 1/184 (0%)
Frame = +2
Query: 221 QENAPDPIINTCNGFYCDQFTPNSKAKPKFWTEGYNGWLLYFGGAVPYRPVEDLAFAVAR 280
Q++APDP+INTCNGFYCD F+PN KPK WTE + GW FGG VP+RP ED+AF+VAR
Sbjct: 2 QDDAPDPVINTCNGFYCDYFSPNKDYKPKMWTEAWTGWFTEFGGPVPHRPAEDMAFSVAR 181
Query: 281 FFQLGGTLQNYYMYHGGTNFGRTSGGPFVATSYDYDAAIDEYGFIRQPKWGHLKDVHKAI 340
F Q GG+ NYYMYHGGTNFGRT+GGPF+ATSYDYDA +DEYG ++QPKWGHLKD+H+AI
Sbjct: 182 FIQKGGSFINYYMYHGGTNFGRTAGGPFIATSYDYDAPLDEYGLLQQPKWGHLKDLHRAI 361
Query: 341 KLCEEALIATDPTITSLGSNLEAAVYKTEA-ECVAFLANIDNTSDATVKFNGNSYNLPAW 399
KL E ALI+ DPT+T +G+ EA V+K+++ C AFL N + + ATV F YNLP W
Sbjct: 362 KLSEPALISGDPTVTRIGNYQEAHVFKSKSGACAAFLGNYNPKAFATVAFGNMHYNLPPW 541
Query: 400 SVSI 403
S+SI
Sbjct: 542 SISI 553
>AL370358 similar to GP|20260596|gb galactosidase putative {Arabidopsis
thaliana}, partial (17%)
Length = 456
Score = 234 bits (597), Expect = 1e-61
Identities = 106/139 (76%), Positives = 122/139 (87%)
Frame = +2
Query: 26 ANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDGGLDVIETYVFWNLHEP 85
A+V+YD +A+ I+G+ R+LISGSIHYPRSTPEMWPDLIQKAK+GGLDVI+TYVFWN HEP
Sbjct: 35 ASVSYDSKAITINGQSRILISGSIHYPRSTPEMWPDLIQKAKEGGLDVIQTYVFWNGHEP 214
Query: 86 VQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNYGGFPLWLHFIPGIQFRTDN 145
G+Y FEG DLVKF+K V AGLYVHLRIGPY CAEWN+GGFP+WL +IPGI FRTDN
Sbjct: 215 SPGKYYFEGNYDLVKFIKLVQQAGLYVHLRIGPYVCAEWNFGGFPVWLKYIPGISFRTDN 394
Query: 146 EPFKAEMQRFTAKIVDMMK 164
EPFK +MQ+FT KIVDMMK
Sbjct: 395 EPFKFQMQKFTEKIVDMMK 451
>TC89271 weakly similar to GP|10177669|dbj|BAB11029. beta-galactosidase
{Arabidopsis thaliana}, partial (30%)
Length = 1173
Score = 172 bits (437), Expect(3) = 8e-56
Identities = 93/185 (50%), Positives = 118/185 (63%), Gaps = 6/185 (3%)
Frame = +3
Query: 481 SLRIDLEDD---AGAQTVLHIESLGHALHAFINGKLAGSGKGNRDNSKVNVEIPITLVAG 537
S+ ID ++ G++ L IES GH LHAF+N K G+G GN +S + PI+L AG
Sbjct: 24 SILIDANEEFLKKGSKPALLIESKGHTLHAFVNQKYQGTGTGNGSHSAFTFKNPISLRAG 203
Query: 538 KNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGSTLDFSSKQWTYQIGLKGEDL 597
KN I +LSLTVGLQ GPF+D GAG+T I+ GL N T+D SS W Y+IG+ GE L
Sbjct: 204 KNEIAILSLTVGLQTAGPFYDFIGAGVTSVKII-GL-NNRTIDLSSNAWAYKIGVLGEHL 377
Query: 598 GLPSG---TSGQWNSQSTLPKNQPLTWYKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSI 654
+ G S +W S S PK Q LTWYK APSG++P+ +D MGKG AW+NG+ I
Sbjct: 378 SIYQGEGMNSVKWTSTSEPPKGQALTWYKAIVDAPSGDEPVGLDMLYMGKGLAWLNGEEI 557
Query: 655 GRYWP 659
GRYWP
Sbjct: 558 GRYWP 572
Score = 42.4 bits (98), Expect(3) = 8e-56
Identities = 17/36 (47%), Positives = 24/36 (66%)
Frame = +2
Query: 699 RSWLQPDSNTLVLFEESGGDPTQISIAIKLIGSSCS 734
RSW +P N LV+FEE GGDPT+I+ + + + S
Sbjct: 695 RSWFKPSGNVLVIFEEKGGDPTKITFVRRKVSGAPS 802
Score = 42.4 bits (98), Expect(3) = 8e-56
Identities = 15/31 (48%), Positives = 20/31 (64%)
Frame = +1
Query: 668 CTSSCNYRGPYDSSKCLKNCGKPSQTLYHVP 698
C C+YRG ++ KC CG+PSQ +HVP
Sbjct: 601 CVQECDYRGKFNPDKCDTGCGEPSQKWHHVP 693
Score = 32.7 bits (73), Expect = 0.60
Identities = 17/50 (34%), Positives = 28/50 (56%), Gaps = 1/50 (2%)
Frame = +3
Query: 793 GDCSSKQALSIVQKACIGSSSCSIGVSINTF-GDPCGGVTKSLAVEASCT 841
GDC + + +V+K C+ + I V + F + C G++ LAVEA C+
Sbjct: 816 GDCHNPYSSIVVEKVCVNKNDRVIKVIEDNFKTNLCHGLSMKLAVEAICS 965
>BF635308 similar to GP|10177061|db beta-galactosidase {Arabidopsis
thaliana}, partial (16%)
Length = 507
Score = 184 bits (467), Expect = 1e-46
Identities = 90/139 (64%), Positives = 103/139 (73%)
Frame = +2
Query: 7 QFLLLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKA 66
+FLL L +L V ANVTYD +LVI+G ++L SGSIHYPRSTP+MWPDLI KA
Sbjct: 92 RFLLHAL-ILTVSLCTVHGANVTYDRTSLVINGHHKILFSGSIHYPRSTPQMWPDLISKA 268
Query: 67 KDGGLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAEWNY 126
K+GGLDVI+TYVFWNLHEP QGQY F GR DL F+K + GLYV LRIGPY +E Y
Sbjct: 269 KEGGLDVIQTYVFWNLHEPQQGQYEFNGRFDLXGFIKEIQAQGLYVTLRIGPYIESECTY 448
Query: 127 GGFPLWLHFIPGIQFRTDN 145
GG PLWLH +PGI F TDN
Sbjct: 449 GGLPLWLHDVPGIVFXTDN 505
>TC86051 homologue to GP|3641863|emb|CAA06309.1 beta-galactosidase {Cicer
arietinum}, partial (29%)
Length = 1246
Score = 95.1 bits (235), Expect(3) = 2e-46
Identities = 41/71 (57%), Positives = 51/71 (71%)
Frame = +2
Query: 622 YKINFSAPSGNDPIAIDFTGMGKGEAWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSS 681
Y F+AP+GNDP+A+D + MGKGE WVNGQSIGR+WP YI+ G SCNY G +
Sbjct: 548 Y*TTFNAPAGNDPLALDMSSMGKGEIWVNGQSIGRHWPAYIA--RGNCGSCNYAGTFTDK 721
Query: 682 KCLKNCGKPSQ 692
KC NCG+P+Q
Sbjct: 722 KCRTNCGQPTQ 754
Score = 81.6 bits (200), Expect(3) = 2e-46
Identities = 54/155 (34%), Positives = 68/155 (43%), Gaps = 46/155 (29%)
Frame = +3
Query: 519 GNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGST 578
G+ DN K+ + L G N I LLS+ VGL + G ++ W G+ GPV LKGL G T
Sbjct: 12 GSLDNPKLTFSNSVKLRVGNNKISLLSVAVGLSNVGVHYEKWNVGVLGPVTLKGLNEG-T 188
Query: 579 LDFSSKQWTY-------------------------------------------QIGLKGE 595
D S ++W+Y QIGLKGE
Sbjct: 189 RDLSKQKWSYKVCYHLYIGVLRKHFNINHVHY*AYLCIYYHI*RHNSKVYVNWQIGLKGE 368
Query: 596 DLGL--PSGTSG-QWNSQSTLPKNQPLTWYKINFS 627
L L SG+S +W S L K QPLTWYK+ S
Sbjct: 369 SLNLHTTSGSSSVKWTQGSFLSKKQPLTWYKVRMS 473
Score = 49.3 bits (116), Expect(3) = 2e-46
Identities = 20/30 (66%), Positives = 23/30 (76%)
Frame = +1
Query: 695 YHVPRSWLQPDSNTLVLFEESGGDPTQISI 724
YH+PRSWL P N LV+ EE GGDPT IS+
Sbjct: 841 YHIPRSWLNPSGNVLVVLEEWGGDPTGISL 930
>BQ141325 similar to GP|2300473|emb unnamed protein product {unidentified},
partial (12%)
Length = 610
Score = 166 bits (421), Expect = 3e-41
Identities = 78/114 (68%), Positives = 95/114 (82%)
Frame = +1
Query: 10 LLLLWLLCVYAPACFCANVTYDHRALVIDGKRRVLISGSIHYPRSTPEMWPDLIQKAKDG 69
+LLL++L A+V+YDH+A+ I+G+RR+L+SGSIHYPRSTPEMWPDLIQKAK+G
Sbjct: 262 VLLLFILASSLIGYGEASVSYDHKAITINGQRRILLSGSIHYPRSTPEMWPDLIQKAKEG 441
Query: 70 GLDVIETYVFWNLHEPVQGQYNFEGRGDLVKFVKGVATAGLYVHLRIGPYACAE 123
GLDVI+TYVFWN HEP G+Y FEG DLVKF+K V AGLYVHLR+GPYACA+
Sbjct: 442 GLDVIQTYVFWNGHEPSPGKYYFEGNYDLVKFIKLVHQAGLYVHLRVGPYACAD 603
>TC89925 homologue to GP|3860321|emb|CAA10128.1 beta-galactosidase {Cicer
arietinum}, partial (20%)
Length = 765
Score = 150 bits (380), Expect = 1e-36
Identities = 72/143 (50%), Positives = 99/143 (68%), Gaps = 4/143 (2%)
Frame = +3
Query: 591 GLKGEDLGL--PSGTSG-QWNSQSTLPKNQP-LTWYKINFSAPSGNDPIAIDFTGMGKGE 646
GL GE + L P+G S W S+S +NQP L W+K +F+AP+G +P+A+D + MGKG+
Sbjct: 3 GLLGEAMNLVSPNGVSSVDWVSESLASQNQPQLKWHKAHFNAPNGVEPLALDMSSMGKGQ 182
Query: 647 AWVNGQSIGRYWPTYISPIDGCTSSCNYRGPYDSSKCLKNCGKPSQTLYHVPRSWLQPDS 706
W+NGQSIGRYW Y G +SCNY G Y +KC CG+P+Q YHVPRSWL+P +
Sbjct: 183 VWINGQSIGRYWMVYAK---GNCNSCNYAGTYRQAKCQVGCGQPTQRWYHVPRSWLKPKN 353
Query: 707 NTLVLFEESGGDPTQISIAIKLI 729
N +V+FEE GG+P +IS+ ++I
Sbjct: 354 NLMVVFEELGGNPWKISLVKRII 422
>BG453318 homologue to GP|3204134|emb beta-galactosidase {Cicer arietinum},
partial (29%)
Length = 641
Score = 147 bits (370), Expect = 2e-35
Identities = 85/206 (41%), Positives = 116/206 (56%), Gaps = 5/206 (2%)
Frame = +1
Query: 403 ILPDCKNVVLNTAKINSESMISSFTTESLKEVGSLEGPSPGWSWISEPIGISKADSFSRF 462
ILP+CK+ V NTA++ S+S + + G L W +E + SF+
Sbjct: 1 ILPNCKHTVYNTARVGSQSAQMKMSRVPIH--GGLS-----WQAFNEETTTTDDSSFTVT 159
Query: 463 GLLEQINTTADRSDYLWYSLRIDLEDDAG-----AQTVLHIESLGHALHAFINGKLAGSG 517
GLLEQ+N T D SDYLWYS + + + G VL + S GHALH FING+L+G+
Sbjct: 160 GLLEQVNATRDLSDYLWYSTDVVINSNEGFFRNGKNPVLTVLSAGHALHVFINGQLSGTA 339
Query: 518 KGNRDNSKVNVEIPITLVAGKNTIDLLSLTVGLQHFGPFFDTWGAGITGPVILKGLKNGS 577
G+ D K+ + L AG N I LLS+ VGL + GP F+TW AG+ GP+ L GL G
Sbjct: 340 YGSVDFPKLTFSESVKLRAGVNKISLLSVAVGLPNIGPHFETWNAGVLGPISLNGLNEGR 519
Query: 578 TLDFSSKQWTYQIGLKGEDLGLPSGT 603
D + ++W+Y+ GLKG L L S T
Sbjct: 520 R-DLTWQKWSYKXGLKGXALSLHSLT 594
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.136 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,687,382
Number of Sequences: 36976
Number of extensions: 495232
Number of successful extensions: 2259
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 2193
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2218
length of query: 841
length of database: 9,014,727
effective HSP length: 104
effective length of query: 737
effective length of database: 5,169,223
effective search space: 3809717351
effective search space used: 3809717351
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0136.7


