

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0134.4
(311 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
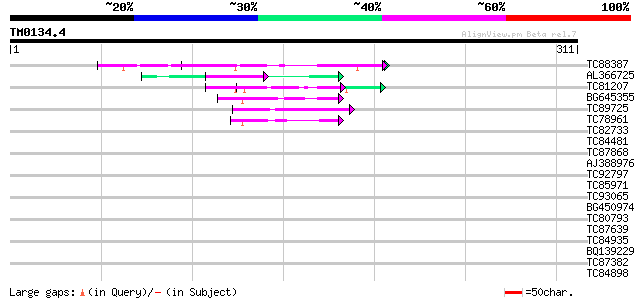
Score E
Sequences producing significant alignments: (bits) Value
TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 60 1e-09
AL366725 52 3e-07
TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-... 49 2e-06
BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.... 43 1e-04
TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.... 43 1e-04
TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 43 2e-04
TC82733 similar to GP|10177404|dbj|BAB10535. gene_id:K24M7.12~pi... 39 0.003
TC84481 similar to GP|11875626|gb|AAG40731.1 XRN4 {Arabidopsis t... 33 0.14
TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 33 0.14
AJ388976 similar to PIR|E84638|E84 probable RSZp22 splicing fact... 31 0.53
TC92797 weakly similar to GP|10178202|dbj|BAB11626. gene_id:K9D7... 30 0.91
TC85971 similar to GP|19424073|gb|AAL87350.1 unknown protein {Ar... 30 0.91
TC93065 30 0.91
BG450974 similar to PIR|T05112|T05 splicing factor 9G8-like SR p... 30 1.2
TC80793 similar to SP|Q9LUA9|COLE_ARATH Zinc finger protein cons... 30 1.6
TC87639 GP|9663153|emb|CAC01132.1 transport-secretion protein 2.... 30 1.6
TC84935 similar to PIR|G96631|G96631 probable RNA-binding protei... 29 2.0
BQ139229 weakly similar to GP|20453199|gb AT3g18290/MIE15_8 {Ara... 29 2.0
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 29 2.0
TC84898 similar to GP|20259267|gb|AAM14369.1 unknown protein {Ar... 29 2.7
>TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (96%)
Length = 1286
Score = 60.1 bits (144), Expect = 1e-09
Identities = 45/169 (26%), Positives = 69/169 (40%), Gaps = 10/169 (5%)
Frame = +1
Query: 49 QSRLDRAGTGGPV----RSGSQEYRQLGPYQHPKGRGFTPRSFKSSTTATPGARSQSAHP 104
+S +DR+ + PV RS +R+ PY+ RGF+ + PG +
Sbjct: 298 RSPMDRSRSRSPVDRRIRSERFSHRE-APYRRDSRRGFSQDNL-CKNCKRPGHYVRECPN 471
Query: 105 EVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGT 164
C C GH A+ C + C+NC + GH+A +C P++
Sbjct: 472 VAVCHNCSLPGHIASECSTKS-LCWNCKEPGHMASSC------PNEG------------- 591
Query: 165 EKQVSCYRCGEIGHFASRCSATGPR------CFNCHRVGHLAVVCKKPK 207
C+ CG+ GH A C+ C NC++ GH+AV C K
Sbjct: 592 ----ICHTCGKAGHRARECTVPQKPPGDLRLCNNCYKQGHIAVECTNEK 726
Score = 52.4 bits (124), Expect = 2e-07
Identities = 37/120 (30%), Positives = 44/120 (35%), Gaps = 6/120 (5%)
Frame = +1
Query: 95 PGARSQSAHPEVTCFRCGKNGHYANACL------GQGPRCFNCNQIGHLAVNCKTFQAGP 148
PG + S E C CGK GH A C G C NC + GH+AV C +A
Sbjct: 556 PGHMASSCPNEGICHTCGKAGHRARECTVPQKPPGDLRLCNNCYKQGHIAVECTNEKA-- 729
Query: 149 SDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKV 208
C C + GH A C P C C+ GH+A C K V
Sbjct: 730 ---------------------CNNCRKTGHLARDC-PNDPICNLCNISGHVARQCPKSNV 843
Score = 32.0 bits (71), Expect = 0.31
Identities = 16/39 (41%), Positives = 20/39 (51%), Gaps = 2/39 (5%)
Frame = +1
Query: 105 EVTCFRCGKNGHYANACLGQGPR--CFNCNQIGHLAVNC 141
+V C C + GH + C+G GP C NC GH A C
Sbjct: 904 DVVCRSCQQFGHMSRDCMG-GPLMICQNCGGRGHQAYEC 1017
>AL366725
Length = 485
Score = 52.0 bits (123), Expect = 3e-07
Identities = 27/111 (24%), Positives = 42/111 (37%)
Frame = +2
Query: 73 PYQHPKGRGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCN 132
PY P +G K + + A E+ CF G+ GH +N C + +C C+
Sbjct: 221 PYSAPADKG------KQRMVDDRRPKKKDAPAEIVCFNYGEKGHKSNVCPKEIKKCVRCD 382
Query: 133 QIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC 183
+ GH+ +CK + C+ C E GH S+C
Sbjct: 383 KKGHIVADCK----------------------RNDIVCFNCNEEGHIGSQC 469
Score = 41.6 bits (96), Expect = 4e-04
Identities = 16/35 (45%), Positives = 18/35 (50%)
Frame = +2
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCK 142
C RC K GH C CFNCN+ GH+ CK
Sbjct: 368 CVRCDKKGHIVADCKRNDIVCFNCNEEGHIGSQCK 472
Score = 35.0 bits (79), Expect = 0.037
Identities = 12/39 (30%), Positives = 21/39 (53%)
Frame = +2
Query: 167 QVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKK 205
++ C+ GE GH ++ C +C C + GH+ CK+
Sbjct: 299 EIVCFNYGEKGHKSNVCPKEIKKCVRCDKKGHIVADCKR 415
>TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-1 {Triticum
aestivum}, partial (39%)
Length = 630
Score = 48.9 bits (115), Expect = 2e-06
Identities = 26/96 (27%), Positives = 37/96 (38%), Gaps = 14/96 (14%)
Frame = +3
Query: 125 GPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC- 183
G C+ C GH+A +C +++ G G ++ +CY CG HFA C
Sbjct: 342 GGGCYTCGDTGHIARDCDRSDRNDRNDRSGG-----GGGGDRDRACYTCGSFEHFARDCM 506
Query: 184 -------------SATGPRCFNCHRVGHLAVVCKKP 206
G C+ C VGH+A C P
Sbjct: 507 RGGGNNNNGGGGYGGGGTSCYRCGGVGHIARDCATP 614
Score = 45.1 bits (105), Expect = 4e-05
Identities = 30/94 (31%), Positives = 39/94 (40%), Gaps = 17/94 (18%)
Frame = +3
Query: 108 CFRCGKNGHYANACL--------------GQGPR---CFNCNQIGHLAVNCKTFQAGPSD 150
C+ CG GH A C G G R C+ C H A +C + G ++
Sbjct: 351 CYTCGDTGHIARDCDRSDRNDRNDRSGGGGGGDRDRACYTCGSFEHFARDC--MRGGGNN 524
Query: 151 NKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCS 184
N G + GT SCYRCG +GH A C+
Sbjct: 525 NNGGGGYGG--GGT----SCYRCGGVGHIARDCA 608
Score = 37.0 bits (84), Expect = 0.010
Identities = 16/54 (29%), Positives = 24/54 (43%), Gaps = 14/54 (25%)
Frame = +3
Query: 108 CFRCGKNGHYANACL--------------GQGPRCFNCNQIGHLAVNCKTFQAG 147
C+ CG H+A C+ G G C+ C +GH+A +C T +G
Sbjct: 462 CYTCGSFEHFARDCMRGGGNNNNGGGGYGGGGTSCYRCGGVGHIARDCATPSSG 623
>BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.5 [imported]
- Arabidopsis thaliana, partial (5%)
Length = 627
Score = 43.1 bits (100), Expect = 1e-04
Identities = 23/72 (31%), Positives = 34/72 (46%), Gaps = 3/72 (4%)
Frame = -1
Query: 115 GHYANACLGQGP---RCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCY 171
G Y N G G +C+ C Q GH A NC + A N++ G + +CY
Sbjct: 549 GAYVNTVSGSGGASGKCYKCQQPGHWASNCPSMSAA---NRVSGGSGGASG------NCY 397
Query: 172 RCGEIGHFASRC 183
+C + GH+A+ C
Sbjct: 396 KCNQPGHWANNC 361
Score = 40.0 bits (92), Expect = 0.001
Identities = 18/57 (31%), Positives = 25/57 (43%), Gaps = 13/57 (22%)
Frame = -1
Query: 108 CFRCGKNGHYANACL-------------GQGPRCFNCNQIGHLAVNCKTFQAGPSDN 151
C++C + GH+A+ C G C+ CNQ GH A NC A P +
Sbjct: 501 CYKCQQPGHWASNCPSMSAANRVSGGSGGASGNCYKCNQPGHWANNCPNMSAAPQSH 331
>TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.150 -
Arabidopsis thaliana, partial (17%)
Length = 378
Score = 43.1 bits (100), Expect = 1e-04
Identities = 24/67 (35%), Positives = 28/67 (40%)
Frame = +1
Query: 123 GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASR 182
G G C+NC + GH+A C + G G G SCY CGE GHFA
Sbjct: 1 GVGGGCYNCGESGHMARECTS--GGGGGGGRYGGGGGGGGGGGGGGSCYSCGESGHFARD 174
Query: 183 CSATGPR 189
C R
Sbjct: 175 CPTGSAR 195
Score = 35.8 bits (81), Expect = 0.022
Identities = 17/59 (28%), Positives = 24/59 (39%), Gaps = 20/59 (33%)
Frame = +1
Query: 108 CFRCGKNGHYANACL--------------------GQGPRCFNCNQIGHLAVNCKTFQA 146
C+ CG++GH A C G G C++C + GH A +C T A
Sbjct: 16 CYNCGESGHMARECTSGGGGGGGRYGGGGGGGGGGGGGGSCYSCGESGHFARDCPTGSA 192
Score = 32.0 bits (71), Expect = 0.31
Identities = 16/54 (29%), Positives = 20/54 (36%), Gaps = 20/54 (37%)
Frame = +1
Query: 170 CYRCGEIGHFASRCSATGP--------------------RCFNCHRVGHLAVVC 203
CY CGE GH A C++ G C++C GH A C
Sbjct: 16 CYNCGESGHMARECTSGGGGGGGRYGGGGGGGGGGGGGGSCYSCGESGHFARDC 177
>TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (71%)
Length = 974
Score = 42.7 bits (99), Expect = 2e-04
Identities = 27/67 (40%), Positives = 29/67 (42%), Gaps = 5/67 (7%)
Frame = +2
Query: 122 LGQGP-----RCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEI 176
LG+GP RCFNC GH A +CK AG NK CYRCGE
Sbjct: 326 LGRGPPPGSGRCFNCGIDGHWARDCK---AGDWKNK-----------------CYRCGER 445
Query: 177 GHFASRC 183
GH C
Sbjct: 446 GHIEKNC 466
Score = 39.3 bits (90), Expect = 0.002
Identities = 15/37 (40%), Positives = 21/37 (56%), Gaps = 2/37 (5%)
Frame = +2
Query: 108 CFRCGKNGHYANACLGQG--PRCFNCNQIGHLAVNCK 142
CF CG +GH+A C +C+ C + GH+ NCK
Sbjct: 359 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCK 469
Score = 33.1 bits (74), Expect = 0.14
Identities = 14/37 (37%), Positives = 19/37 (50%), Gaps = 2/37 (5%)
Frame = +2
Query: 170 CYRCGEIGHFASRCSATG--PRCFNCHRVGHLAVVCK 204
C+ CG GH+A C A +C+ C GH+ CK
Sbjct: 359 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCK 469
>TC82733 similar to GP|10177404|dbj|BAB10535.
gene_id:K24M7.12~pir||S42136~similar to unknown protein
{Arabidopsis thaliana}, partial (57%)
Length = 710
Score = 38.9 bits (89), Expect = 0.003
Identities = 35/134 (26%), Positives = 42/134 (31%), Gaps = 12/134 (8%)
Frame = +3
Query: 84 PRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLGQGPR-----CFNCNQIGHLA 138
P S T R P +CF C H A C + C C + GH A
Sbjct: 210 PDSNSKPRTGKRPLRVPGMKPGDSCFICKGLDHIAKFCTQKAEWEKNKICLRCRRRGHRA 389
Query: 139 VNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRC-------SATGPRCF 191
NC G S K Y CG+ GH + C +CF
Sbjct: 390 QNCPD---GGSKEDFK--------------Y*YNCGDNGHSLANCPHPLQEGGTMFAQCF 518
Query: 192 NCHRVGHLAVVCKK 205
C GHL+ C K
Sbjct: 519 VCKEQGHLSKNCPK 560
>TC84481 similar to GP|11875626|gb|AAG40731.1 XRN4 {Arabidopsis thaliana},
partial (23%)
Length = 727
Score = 33.1 bits (74), Expect = 0.14
Identities = 13/31 (41%), Positives = 18/31 (57%)
Frame = +2
Query: 123 GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKM 153
GQ +C+ C Q+GHLA C+ P DN +
Sbjct: 194 GQQEKCYMCGQVGHLAAECR---GKPGDNAL 277
Score = 30.4 bits (67), Expect = 0.91
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = +2
Query: 166 KQVSCYRCGEIGHFASRC 183
+Q CY CG++GH A+ C
Sbjct: 197 QQEKCYMCGQVGHLAAEC 250
>TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (91%)
Length = 860
Score = 33.1 bits (74), Expect = 0.14
Identities = 22/83 (26%), Positives = 31/83 (36%), Gaps = 5/83 (6%)
Frame = +3
Query: 163 GTEKQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVEPSVNTARGKHP-- 220
G + CY CGE GHFA C G R + P + RG+ P
Sbjct: 315 GGGSDLKCYECGEPGHFARECRNRGGGGAGRRRSRSPPRFRRSPSYGRRSYSPRGRSPRR 494
Query: 221 ---AARGKVYTMDGEEAEGADEL 240
+ RG+ Y+ G +E+
Sbjct: 495 RSVSPRGRSYSRSPPPYRGREEV 563
Score = 28.5 bits (62), Expect = 3.5
Identities = 8/21 (38%), Positives = 14/21 (66%)
Frame = +3
Query: 105 EVTCFRCGKNGHYANACLGQG 125
++ C+ CG+ GH+A C +G
Sbjct: 327 DLKCYECGEPGHFARECRNRG 389
>AJ388976 similar to PIR|E84638|E84 probable RSZp22 splicing factor
[imported] - Arabidopsis thaliana, partial (62%)
Length = 508
Score = 31.2 bits (69), Expect = 0.53
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = +2
Query: 163 GTEKQVSCYRCGEIGHFASRCSAT 186
G + CY CGE GHFA C+++
Sbjct: 323 GGGSDLKCYXCGEPGHFARXCNSS 394
Score = 28.1 bits (61), Expect = 4.5
Identities = 9/24 (37%), Positives = 14/24 (57%)
Frame = +2
Query: 98 RSQSAHPEVTCFRCGKNGHYANAC 121
RS ++ C+ CG+ GH+A C
Sbjct: 314 RSGGGGSDLKCYXCGEPGHFARXC 385
>TC92797 weakly similar to GP|10178202|dbj|BAB11626. gene_id:K9D7.13~unknown
protein {Arabidopsis thaliana}, partial (4%)
Length = 660
Score = 30.4 bits (67), Expect = 0.91
Identities = 27/105 (25%), Positives = 42/105 (39%), Gaps = 25/105 (23%)
Frame = +2
Query: 170 CYRCGEIGHFASRCSAT----------------GPR-----CFNCHRVGHLAVVCKKPKV 208
C CG+ GH S CSA GP C C ++ H AV C
Sbjct: 65 CLFCGKRGHQLSDCSAVAESELEDLQENVNSYEGPENFPLMCIKCFQLNHWAVSC----- 229
Query: 209 EPSVNTARGKHPAARGKVYTMDGE----EAEGADELVKGERKNDG 249
S + ++ KH ++ K + +G + + AD ++ G +DG
Sbjct: 230 --SSSISKRKH-ESKVKTFLHEGSARPAQIDEADRILSGGAIHDG 355
Score = 28.9 bits (63), Expect = 2.7
Identities = 16/57 (28%), Positives = 21/57 (36%), Gaps = 21/57 (36%)
Frame = +2
Query: 108 CFRCGKNGHYANACLG----------------QGPR-----CFNCNQIGHLAVNCKT 143
C CGK GH + C +GP C C Q+ H AV+C +
Sbjct: 65 CLFCGKRGHQLSDCSAVAESELEDLQENVNSYEGPENFPLMCIKCFQLNHWAVSCSS 235
>TC85971 similar to GP|19424073|gb|AAL87350.1 unknown protein {Arabidopsis
thaliana}, partial (77%)
Length = 1540
Score = 30.4 bits (67), Expect = 0.91
Identities = 22/82 (26%), Positives = 29/82 (34%), Gaps = 6/82 (7%)
Frame = +3
Query: 107 TCFRCGKNGHYANACLGQGPRCFNCNQIGH------LAVNCKTFQAGPSDNKMKGKFPAI 160
TC + GHY + Q P C+ I H L V C KGK P
Sbjct: 690 TCNADEEPGHYKENQVTQCPSCYGRGLIAHKDGSDTLCVKCSG----------KGKLPCA 839
Query: 161 ARGTEKQVSCYRCGEIGHFASR 182
G+ ++C C G +R
Sbjct: 840 TCGSRGLLNCETCNGSGSLLAR 905
>TC93065
Length = 783
Score = 30.4 bits (67), Expect = 0.91
Identities = 14/36 (38%), Positives = 19/36 (51%), Gaps = 1/36 (2%)
Frame = +2
Query: 108 CFRCGKNGHYANACLGQ-GPRCFNCNQIGHLAVNCK 142
C C K+ H C + G +C CNQ+GH+ CK
Sbjct: 572 CPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCK 679
Score = 28.5 bits (62), Expect = 3.5
Identities = 14/43 (32%), Positives = 21/43 (48%), Gaps = 1/43 (2%)
Frame = +2
Query: 163 GTEKQVSCYRCGEIGHFASRC-SATGPRCFNCHRVGHLAVVCK 204
G +K C C + H C G +C C+++GH+ VCK
Sbjct: 551 GKKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCK 679
>BG450974 similar to PIR|T05112|T05 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (54%)
Length = 364
Score = 30.0 bits (66), Expect = 1.2
Identities = 10/16 (62%), Positives = 11/16 (68%)
Frame = +1
Query: 168 VSCYRCGEIGHFASRC 183
+ CY CGE GHFA C
Sbjct: 313 LKCYECGEPGHFAREC 360
Score = 28.5 bits (62), Expect = 3.5
Identities = 9/26 (34%), Positives = 14/26 (53%)
Frame = +1
Query: 96 GARSQSAHPEVTCFRCGKNGHYANAC 121
G R ++ C+ CG+ GH+A C
Sbjct: 283 GGRGGRGGDDLKCYECGEPGHFAREC 360
>TC80793 similar to SP|Q9LUA9|COLE_ARATH Zinc finger protein constans-like
14. [Mouse-ear cress] {Arabidopsis thaliana}, partial
(27%)
Length = 937
Score = 29.6 bits (65), Expect = 1.6
Identities = 22/88 (25%), Positives = 32/88 (36%), Gaps = 1/88 (1%)
Frame = +1
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
C C +N H AN + R C + PA+ R +E++
Sbjct: 310 CLSCDRNVHSANTLARRHSRTLLCERCSSQ--------------------PALVRCSEEK 429
Query: 168 VS-CYRCGEIGHFASRCSATGPRCFNCH 194
VS C C +GH S S + NC+
Sbjct: 430 VSLCQNCDWLGHGNSTSSNHKRQTINCY 513
>TC87639 GP|9663153|emb|CAC01132.1 transport-secretion protein 2.2 (TTS-2.2)
{Homo sapiens}, partial (2%)
Length = 1522
Score = 29.6 bits (65), Expect = 1.6
Identities = 21/70 (30%), Positives = 30/70 (42%), Gaps = 5/70 (7%)
Frame = -1
Query: 172 RCGEIGHFASRCSATGPR-----CFNCHRVGHLAVVCKKPKVEPSVNTARGKHPAARGKV 226
R G +S A PR C++CH GH+A C NT+ K KV
Sbjct: 385 RRGSSSSVSSAEGAPPPRRDMRECYSCHERGHIARNC--------TNTSAAKKKKKGPKV 230
Query: 227 YTMDGEEAEG 236
++ ++AEG
Sbjct: 229 AKVEPQKAEG 200
Score = 29.3 bits (64), Expect = 2.0
Identities = 13/35 (37%), Positives = 20/35 (57%)
Frame = -1
Query: 128 CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIAR 162
C++C++ GH+A NC A K K K P +A+
Sbjct: 316 CYSCHERGHIARNCTNTSAA----KKKKKGPKVAK 224
>TC84935 similar to PIR|G96631|G96631 probable RNA-binding protein F8A5.17
[imported] - Arabidopsis thaliana, partial (41%)
Length = 552
Score = 29.3 bits (64), Expect = 2.0
Identities = 14/45 (31%), Positives = 22/45 (48%), Gaps = 6/45 (13%)
Frame = +2
Query: 149 SDNKMKGKFPAIARGTEK------QVSCYRCGEIGHFASRCSATG 187
+D + +G F + RG+ Q C++CG GH+A C G
Sbjct: 392 ADQRYRGGFSSGGRGSYGAGDRVGQDDCFKCGRPGHWARDCPLAG 526
Score = 27.3 bits (59), Expect = 7.7
Identities = 8/14 (57%), Positives = 11/14 (78%)
Frame = +2
Query: 108 CFRCGKNGHYANAC 121
CF+CG+ GH+A C
Sbjct: 473 CFKCGRPGHWARDC 514
>BQ139229 weakly similar to GP|20453199|gb AT3g18290/MIE15_8 {Arabidopsis
thaliana}, partial (4%)
Length = 648
Score = 29.3 bits (64), Expect = 2.0
Identities = 20/54 (37%), Positives = 27/54 (49%), Gaps = 5/54 (9%)
Frame = -3
Query: 182 RCSATGPRCFNCHRVGHLAVVCKKPKVEPSV---NTARGK--HPAARGKVYTMD 230
R S+T PRC NC R VC +P SV +ARG H + G+ ++D
Sbjct: 241 RSSSTPPRCHNCGR*QKPFAVCHRPCRACSVLLCGSARGPRFHRGSAGRCTSVD 80
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 29.3 bits (64), Expect = 2.0
Identities = 12/26 (46%), Positives = 15/26 (57%)
Frame = +2
Query: 188 PRCFNCHRVGHLAVVCKKPKVEPSVN 213
P+CF C GH+A+ C KV VN
Sbjct: 1040 PKCFICQGYGHIALDCVNQKVFTIVN 1117
>TC84898 similar to GP|20259267|gb|AAM14369.1 unknown protein {Arabidopsis
thaliana}, partial (68%)
Length = 792
Score = 28.9 bits (63), Expect = 2.7
Identities = 27/94 (28%), Positives = 35/94 (36%), Gaps = 17/94 (18%)
Frame = +1
Query: 118 ANACLGQGPRCFNCNQIGHLAVNCKTFQAGPS---DNKMKG-----KFPAIARGTEKQVS 169
A +G RC +C+ G AV C T AG D+ M+ K + G +
Sbjct: 361 AEKAIGNNTRCNDCHAKG--AVLCATC-AGSGLYVDSIMESQGIIVKVRCLGCGGTGNIM 531
Query: 170 CYRCGEIGHFASR---------CSATGPRCFNCH 194
C CG GH S+ C T CF H
Sbjct: 532 CTECGGRGHLGSK*SKSDFLEFCCTTNM*CFGWH 633
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.134 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,454,291
Number of Sequences: 36976
Number of extensions: 163592
Number of successful extensions: 955
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 840
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 917
length of query: 311
length of database: 9,014,727
effective HSP length: 96
effective length of query: 215
effective length of database: 5,465,031
effective search space: 1174981665
effective search space used: 1174981665
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0134.4


