

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0134.14
(277 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
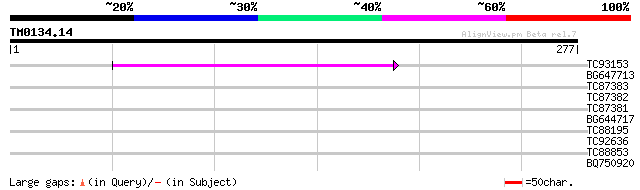
Score E
Sequences producing significant alignments: (bits) Value
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 95 3e-20
BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent vir... 38 0.005
TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA PO... 35 0.032
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 35 0.041
TC87381 31 0.46
BG644717 29 2.3
TC88195 similar to GP|19347889|gb|AAL86001.1 unknown protein {Ar... 28 3.0
TC92636 homologue to GP|15042313|gb|AAK82093.1 232R {Chilo iride... 27 6.6
TC88853 similar to SP|O04350|TBCA_ARATH Tubulin-specific chapero... 27 6.6
BQ750920 similar to GP|15811426|gb| glycine-rich protein {Coccid... 27 6.6
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 94.7 bits (234), Expect = 3e-20
Identities = 52/140 (37%), Positives = 72/140 (51%)
Frame = +2
Query: 51 DQNRGLNDFRRQDPPKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAE 110
D R L F R PP F G PD A ++EIE+IF V+Q E KV T++L +A+
Sbjct: 92 DGTRMLETFLRNHPPTFKGRYAPDGA*KWLKEIERIFRVMQCFETQKVQFGTHMLAEEAD 271
Query: 111 YWWKGTRGIMRANHEEVN*NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKL 170
WW ++ + V FR FL +YFP R ++E +FL L QG M+V EYA+K
Sbjct: 272 DWWISLLPVLEQDDAVVTWAMFRKEFLGRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKF 451
Query: 171 ESLAKHFQFFNNHVDERYMC 190
LA + ++ E C
Sbjct: 452 VELATFYPHYSAETAEFSKC 511
>BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent virus}, partial
(1%)
Length = 726
Score = 37.7 bits (86), Expect = 0.005
Identities = 28/115 (24%), Positives = 48/115 (41%)
Frame = +2
Query: 3 QNMVNSNQLAEMVATLVQAMTVQTNDNAHRRATEDTRELHRLQREVALDQNRGLNDFRRQ 62
Q VN+ Q L++A + +D+ + +R +R+ + + +
Sbjct: 200 QETVNAQQ------ALLEAQRRRNDDDGSGSDSSSSRSSRSHRRQTRMSKIK-------V 340
Query: 63 DPPKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKGTR 117
D P F G PD +Q IE++F+ + AE KV + L A WWK +
Sbjct: 341 DIPDF*GKLQPDEFVDWLQTIERVFKYKEVAEEQKVKIVAAKLKKHASIWWKNLK 505
>TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA POLYMERASE II
{Encephalitozoon cuniculi}, partial (0%)
Length = 1247
Score = 35.0 bits (79), Expect = 0.032
Identities = 43/185 (23%), Positives = 70/185 (37%), Gaps = 2/185 (1%)
Frame = -2
Query: 3 QNMVNSNQLAEMVATLVQAMTVQTNDNAHRRATEDTRELHRLQREVALDQNRGLNDFRRQ 62
Q +VN+ Q L++A + D+ + +R +RE + ND +
Sbjct: 871 QEIVNAQQ------ALLEAQQKRFKDHVSSSDSLSSRSSRSQRREFQM------NDIK*- 731
Query: 63 DPPKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKGT-RGIMR 121
D P F G PD+ +Q +E++F+ + E KV + L A WW+ R R
Sbjct: 730 DIPDFEGNLQPDDLLDWLQIMERLFKYKEVLEEQKVKIVAAKLKKLASIWWENVKRRRKR 551
Query: 122 ANHEEV-N*NSFRTAFLEKYFPTSARDEREAQFLTLGQGSMTVPEYASKLESLAKHFQFF 180
++ R KY P Q + T P+ + K S +HF
Sbjct: 550 EGKSKIKTWEKMRQKLTRKYLPPH-----------YYQDNYTQPQLSKK--SSYRHFSPT 410
Query: 181 NNHVD 185
N +D
Sbjct: 409 KNQID 395
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 34.7 bits (78), Expect = 0.041
Identities = 33/122 (27%), Positives = 48/122 (39%), Gaps = 11/122 (9%)
Frame = +2
Query: 7 NSNQLAEMVATLVQAMTVQTNDNAHRRATEDTRELHRLQREVALDQN---RGLNDFRRQ- 62
N L EM Q +Q NA + E E R + +V+ + R + RRQ
Sbjct: 428 NERSLQEMEDMRRQIQQLQEIINAQQALLE--AEQRRFEGDVSYSDSSSSRSSHSQRRQL 601
Query: 63 -------DPPKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKG 115
D P F G D+ +Q IE++FE + E KV + L A WW+
Sbjct: 602 QMNDIKVDIPDFEGNLQLDDFLDWLQTIERVFEYKEVPEEQKVKIVAAKLKKHALIWWEN 781
Query: 116 TR 117
+
Sbjct: 782 LK 787
>TC87381
Length = 814
Score = 31.2 bits (69), Expect = 0.46
Identities = 20/62 (32%), Positives = 28/62 (44%)
Frame = +3
Query: 56 LNDFRRQDPPKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLLLGDAEYWWKG 115
+ND + D P F G D *+Q IE +FE + E KV + L A WW+
Sbjct: 612 MNDIK-VDIPDFEGELQSDEFVD*LQAIECVFEYKEIPEDHKVKVVAV*LKKHALIWWEN 788
Query: 116 TR 117
+
Sbjct: 789 LK 794
>BG644717
Length = 267
Score = 28.9 bits (63), Expect = 2.3
Identities = 15/41 (36%), Positives = 23/41 (55%)
Frame = -2
Query: 65 PKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAKVSLATYLL 105
P+F G ++ + EI+KIFEV+ + V LA+Y L
Sbjct: 260 PEFLGSQINEDPQNFLDEIKKIFEVMHVSGNDLVELASYQL 138
>TC88195 similar to GP|19347889|gb|AAL86001.1 unknown protein {Arabidopsis
thaliana}, partial (55%)
Length = 1716
Score = 28.5 bits (62), Expect = 3.0
Identities = 21/74 (28%), Positives = 31/74 (41%), Gaps = 1/74 (1%)
Frame = +3
Query: 195 NGLRAD-IKDSVRPLGIIRFQALVEKATEVELMKNRRMNGAGTGGPMRSSSQDNLGKGRF 253
N L+A I+ + RP+ I + AT + R T P + G+G +
Sbjct: 1149 NALQASPIQLAGRPIYIEERRPSTSSATRGGRGRGRGRGSYPTDAPRGRFGGRSSGRGYY 1328
Query: 254 QMKKPYQRPMGEGY 267
Q Y RP G+GY
Sbjct: 1329 QDTSDYSRPRGDGY 1370
>TC92636 homologue to GP|15042313|gb|AAK82093.1 232R {Chilo iridescent
virus}, partial (1%)
Length = 772
Score = 27.3 bits (59), Expect = 6.6
Identities = 18/66 (27%), Positives = 28/66 (42%)
Frame = +3
Query: 32 RRATEDTRELHRLQREVALDQNRGLNDFRRQDPPKFTGGTDPDNADL*IQEIEKIFEVLQ 91
RR +D R + + L + + D P F G PD +Q IE++FE +
Sbjct: 339 RRVDDDGSSDSSSSRSSRSHRRKTLMNDIKVDIPDFEGELQPDEFVDWLQAIERVFEYKE 518
Query: 92 TAEGAK 97
GA+
Sbjct: 519 IPRGAQ 536
>TC88853 similar to SP|O04350|TBCA_ARATH Tubulin-specific chaperone A
(Tubulin-folding cofactor A) (CFA) (TCP1-chaperonin
cofactor A homolog), partial (73%)
Length = 689
Score = 27.3 bits (59), Expect = 6.6
Identities = 19/63 (30%), Positives = 33/63 (52%), Gaps = 5/63 (7%)
Frame = +3
Query: 43 RLQREVALDQNRG-----LNDFRRQDPPKFTGGTDPDNADL*IQEIEKIFEVLQTAEGAK 97
R + E AL+ +G LN+ +++ P+ D A I E+EK+FE ++ EG+
Sbjct: 384 RKRLEAALEDLKGILAELLNETDKKESPEI------DEARNTIVEVEKVFETIEA*EGSY 545
Query: 98 VSL 100
S+
Sbjct: 546 TSV 554
>BQ750920 similar to GP|15811426|gb| glycine-rich protein {Coccidioides
immitis}, partial (18%)
Length = 729
Score = 27.3 bits (59), Expect = 6.6
Identities = 14/39 (35%), Positives = 19/39 (47%)
Frame = +1
Query: 232 NGAGTGGPMRSSSQDNLGKGRFQMKKPYQRPMGEGYTLG 270
NG+G GG R S + G+GR + +G G LG
Sbjct: 421 NGSGGGGGRRGGSSRSGGRGRSSLGSGISGAVGLGLLLG 537
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.138 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,342,125
Number of Sequences: 36976
Number of extensions: 68732
Number of successful extensions: 386
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 384
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 386
length of query: 277
length of database: 9,014,727
effective HSP length: 95
effective length of query: 182
effective length of database: 5,502,007
effective search space: 1001365274
effective search space used: 1001365274
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0134.14


