

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
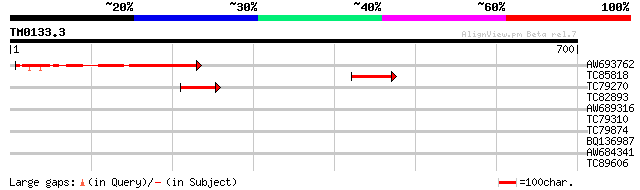
Query= TM0133.3
(700 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW693762 202 3e-52
TC85818 homologue to GP|21261784|emb|CAD32500. protein kinase Ck... 109 4e-24
TC79270 weakly similar to GP|17473774|gb|AAL38323.1 unknown prot... 43 4e-04
TC82893 weakly similar to GP|11762112|gb|AAG40334.1 AT3g11890 {A... 40 0.004
AW689316 similar to GP|20259452|g unknown protein {Arabidopsis t... 32 0.63
TC79310 similar to GP|19920098|gb|AAM08530.1 Putative calmodulin... 32 1.1
TC79874 similar to GP|17065608|gb|AAL33784.1 unknown protein {Ar... 31 1.4
BQ136987 31 1.8
AW684341 homologue to GP|1730449|emb| POU domain protein {Danio ... 30 3.1
TC89606 similar to GP|7239157|gb|AAF43095.1| homeodomain protein... 28 9.2
>AW693762
Length = 655
Score = 202 bits (515), Expect = 3e-52
Identities = 128/248 (51%), Positives = 152/248 (60%), Gaps = 19/248 (7%)
Frame = +1
Query: 8 TATTMSLDMDLALFDD---DEFEIPLTQTATI------------HRPLKKHKL-ETTAFG 51
T TTM+ D D FD+ D+FE+PLTQT + RP KK K +TT+F
Sbjct: 10 TPTTMA-DSDHTQFDNNDNDDFEVPLTQTQSPILKPLSQSNNQNQRPSKKPKTNKTTSF- 183
Query: 52 GSGKENVPPNASSSTIHDDGYEIENCSLDFIPSTIDSVGTALTDSSSSSSSASVSAFVIS 111
GKEN+PP +GY+I N SLDF+PST+DS T V S
Sbjct: 184 -PGKENIPPF--------EGYDIGNSSLDFVPSTLDSDST-----------------VFS 285
Query: 112 SPVKVKMKGTA--YFGNSIESKLVVSRANAINNAEAGSELNLSDELDGSVRCPLCEVDIS 169
SPV K YF NS+ES+LV SR NA + E+NL DE SV CPLC VDIS
Sbjct: 286 SPVSELKKSNKRDYFSNSLESRLVASRKNAFD-----LEVNLCDEFGSSVDCPLCGVDIS 450
Query: 170 NLTEEQRHLHTNDCLDR-GDDVAVRDDNVDAQFAPKVSRVVDWLRDLGLGKYEDVFVREE 228
NLTEEQR+LHTNDCLD+ G+DVA +D+V AQF PK S VV+W+R LGL KYE+VFVREE
Sbjct: 451 NLTEEQRNLHTNDCLDKSGEDVAPPNDDVGAQFGPKNSPVVEWIRGLGLAKYEEVFVREE 630
Query: 229 VDWDTLQW 236
VDWDTLQW
Sbjct: 631 VDWDTLQW 654
>TC85818 homologue to GP|21261784|emb|CAD32500. protein kinase Ck2
regulatory subunit 2 {Nicotiana tabacum}, partial (44%)
Length = 827
Score = 109 bits (272), Expect = 4e-24
Identities = 50/56 (89%), Positives = 53/56 (94%)
Frame = +2
Query: 422 ARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVL 477
ARLVNMNIGIPYDKLHILPLNQK+EIAGI VTCLDANHCPGSI+ILF+PPNGK L
Sbjct: 290 ARLVNMNIGIPYDKLHILPLNQKIEIAGISVTCLDANHCPGSILILFEPPNGKRSL 457
>TC79270 weakly similar to GP|17473774|gb|AAL38323.1 unknown protein
{Arabidopsis thaliana}, partial (18%)
Length = 1288
Score = 43.1 bits (100), Expect = 4e-04
Identities = 23/49 (46%), Positives = 33/49 (66%)
Frame = +3
Query: 212 LRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHAL 260
L+ LGL KY+ +F EEVD L+ + E DL +G+ +GPRKK++ AL
Sbjct: 1047 LQALGLQKYQILFKAEEVDMTALRQMGENDLKELGI-PMGPRKKLLLAL 1190
>TC82893 weakly similar to GP|11762112|gb|AAG40334.1 AT3g11890 {Arabidopsis
thaliana}, partial (9%)
Length = 967
Score = 39.7 bits (91), Expect = 0.004
Identities = 22/72 (30%), Positives = 40/72 (55%)
Frame = +2
Query: 189 DVAVRDDNVDAQFAPKVSRVVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVT 248
D ++R N++ + + + WL LGL ++ +F + + L LT + L MG
Sbjct: 563 DRSLRASNLERKLE-RAGGLNKWLASLGLEQFVRMFQGKIISKYQLVNLTMKKLKDMGAN 739
Query: 249 ALGPRKKIVHAL 260
A+GPR+K++HA+
Sbjct: 740 AVGPRRKLIHAM 775
>AW689316 similar to GP|20259452|g unknown protein {Arabidopsis thaliana},
partial (13%)
Length = 648
Score = 32.3 bits (72), Expect = 0.63
Identities = 26/67 (38%), Positives = 34/67 (49%)
Frame = +2
Query: 92 ALTDSSSSSSSASVSAFVISSPVKVKMKGTAYFGNSIESKLVVSRANAINNAEAGSELNL 151
A+ SSS SS SVS+ V+ S K KGT S+ES L S+ LN+
Sbjct: 329 AMDRSSSLSSGTSVSSGVLLSQAKSLGKGTE---RSLESVLHASKQKVTAIESMLRGLNM 499
Query: 152 SDELDGS 158
SD+ +GS
Sbjct: 500 SDKHNGS 520
>TC79310 similar to GP|19920098|gb|AAM08530.1 Putative calmodulin-binding
protein similar to ER66 {Oryza sativa}, partial (13%)
Length = 733
Score = 31.6 bits (70), Expect = 1.1
Identities = 19/50 (38%), Positives = 28/50 (56%)
Frame = +1
Query: 53 SGKENVPPNASSSTIHDDGYEIENCSLDFIPSTIDSVGTALTDSSSSSSS 102
S K N+ NA S+ + D ++ + S IP+T SV + TDS S +SS
Sbjct: 502 SNKSNIGGNADSNEVISDSQKVNSPSSG-IPATYSSVPSLSTDSMSPTSS 648
>TC79874 similar to GP|17065608|gb|AAL33784.1 unknown protein {Arabidopsis
thaliana}, partial (88%)
Length = 1154
Score = 31.2 bits (69), Expect = 1.4
Identities = 26/79 (32%), Positives = 37/79 (45%)
Frame = -2
Query: 95 DSSSSSSSASVSAFVISSPVKVKMKGTAYFGNSIESKLVVSRANAINNAEAGSELNLSDE 154
DSS ++ A V F SP + G SIE +L+ A+AI+ A GSE+ +S
Sbjct: 577 DSSPPANIARVPFFAPRSPPET--------GASIE*QLLTDAASAISTAREGSEVVMSTR 422
Query: 155 LDGSVRCPLCEVDISNLTE 173
+ +R V S TE
Sbjct: 421 MPPDLRPARAPVFGSRRTE 365
>BQ136987
Length = 681
Score = 30.8 bits (68), Expect = 1.8
Identities = 37/134 (27%), Positives = 59/134 (43%), Gaps = 6/134 (4%)
Frame = +2
Query: 13 SLDMDLALFDDDEFEIPLTQTATIHRPLKKHKLETTAFGGSGKENVPPNASSSTIH---D 69
+L D+ F D + + P QT + K+ G G + + P+ + +H D
Sbjct: 185 NLHFDMVHFGDTKKDQPFYQTPLAPVDILKNY-------GKGFKRLLPDDEGNILHPVND 343
Query: 70 DGYEIENCSLDFIPSTIDSV---GTALTDSSSSSSSASVSAFVISSPVKVKMKGTAYFGN 126
G N + + STI+ + GT SSSSS SA S +++ P G ++ G
Sbjct: 344 FGLVTNNENEKKLLSTIEIMKMAGTRFIQSSSSSKSA--SGLILNHPF-----GFSFSGL 502
Query: 127 SIESKLVVSRANAI 140
S E K VS A ++
Sbjct: 503 SDEEKENVSLAESL 544
>AW684341 homologue to GP|1730449|emb| POU domain protein {Danio rerio},
partial (2%)
Length = 308
Score = 30.0 bits (66), Expect = 3.1
Identities = 16/39 (41%), Positives = 22/39 (56%)
Frame = +2
Query: 418 FECFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLD 456
F CFA L + + I + I+ L +KVE+ IDV C D
Sbjct: 38 FACFAHLNDSQLKIIIIIIIIINLKRKVEVNWIDVKCRD 154
>TC89606 similar to GP|7239157|gb|AAF43095.1| homeodomain protein {Malus x
domestica}, partial (12%)
Length = 1018
Score = 28.5 bits (62), Expect = 9.2
Identities = 14/76 (18%), Positives = 42/76 (54%)
Frame = +2
Query: 262 EFRKGIAASNEKPEDALAEPRRNRNQNVKMQPHRPERKVDGTSKPAANKKITEYFPGFAT 321
+F G+A+ P + + + ++++ H+ +R +D ++ + ++ + GF++
Sbjct: 197 DFNMGVASPVPFPSSS------SYSSIIQLKTHQNQRTLDCSTSNSMSQDYQQAIFGFSS 358
Query: 322 NGKKVSTAPEEQQEMK 337
NG + S++ ++QQ+ +
Sbjct: 359 NGFERSSSSQQQQQQQ 406
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.134 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,672,569
Number of Sequences: 36976
Number of extensions: 320328
Number of successful extensions: 2132
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1866
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2013
length of query: 700
length of database: 9,014,727
effective HSP length: 103
effective length of query: 597
effective length of database: 5,206,199
effective search space: 3108100803
effective search space used: 3108100803
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0133.3


